ISSN: 0973-7510
E-ISSN: 2581-690X
Ninety-one Bacterial samples were collected from Al-Sader Medical City in Al-Najaf province during a period of 11 weeks from 2-7-2016 to 15-9-2016. The isolates were identified according to cultural characteristics and biochemical activities. The results have revealed that 25 (27%) samples were E.coli, 22 (24%) samples were Staphylococcus spp, 27 (30%) samples were Proteus spp, and 17 (22%) samples were Enterococcus spp. The susceptibility of bacterial isolates to 22 antibiotics was tested using disc diffusion method. The results have revealed that in E.coli the most resistance was seen to Cotrimoxaz antibiotic with percentage of 67%. In Staphylococcus spp, most resistance was seen to Azactam antibiotic with percentage of 83% while susceptibility to Vancomycin was the most. In Enterococcus spp, the most resistance was seen to Vancomycin. Proteus spp, showed the most resistance against Ampicillin/cloxacillin. The distribution of bacterial infection among different genders and their effect on bacterial resistance were also tested. The results showed that infections with E.coli were 12 (48%) in males while in females 13 (52%). Most of Staphylococcus spp 13 (59%), has been isolated from males, while in females 9 (41%). Males infected with Proteus spp constitute 13 (48%), on the other hand, 14 (52%) were isolated from females. Infections with Enterococcus spp were distributed as 8 (47%) and 9 (53%) in males and females respectively. The different bacterial sites where also correlated and tested against the susceptibility results. All isolates displays different resistance to different antibiotics that varies with infection site and patient gender. In E. coli the resistance to antibiotics in UTI and intestinal disease was approximately the same. In Staphylococcus spp. and Proteus spp. wounds infections exhibit more resistant than other types of infections. In Enterococcus spp. UTI infections exhibit more resistant than wounds infections.
Bacterial Infection, Antibiotic Resistance, Al-Najaf
Antibiotics are produced naturally by some fungi and bacteria as a secondary metabolites to kill other microorganisms that live in the same habitat for the sake of nutritional competition especially in the environments of limited nutritional resources (Bennett and Feibelman, 2001). The antibiotics used to treat people nowadays are usually derived from these natural products (Clardy et al., 2009). Antibiotics are used to treat many illnesses caused by bacterial infections, from ear and skin infections to pneumonia, food poisoning, meningitis, and other life-threatening infections so that antibiotics are essential tools for physicians at any given time.
Between 25% and 40% of hospital patients are receiving antibiotics intravenously (Equi et al., 2002; Wilson, 2006). As antibiotic became widely used, resistant bacterial strains having the ability to inactivate the drug became prevalent, therefore, chemical and structural studies that deals with the antibiotic synthesis were undertaken to modify penicillin chemistry in order to prevent its enzymatic destruction by penicillinases (â-lactamases) which are produced by drug resistant bacteria (Gold and Moellering Jr, 1996; Kardos and Demain, 2011). Interestingly, the ability of bacteria to produce â-lactamases became possible by using the gene based molecular techniques (Al-Shamarti, 2010). This, however, is an appreciated prospect to provide resistance profile of pathogenic bacteria in order to determine the effective antibiotic to be used in treatment (D’costa et al., 2006). Antibiotic resistance could arise as a result of spontaneous or induced genetic alteration in bacteria in addition to the horizontal gene transfer by means of conjugation among different bacterial cells. Thus, antibiotic resistance genes which had evolved as a result of natural selection could be shared (Mazodier and Davies, 1991). According to the concept of natural selection, random exposure to antibiotic puts evolutionary stress on bacteria to develop antibiotic resistance traits. Many antibiotic resistance genes are found on plasmids, enabling their transfer (Bennett, 2008). When a bacterium has the ability to resist several types of different antibiotic as a result of having several resistance genes, it is formally termed as multidrug resistant (MDR) or, informally, super bacterium or superbug (D’Costa et al., 2011).
Aim of the study
The aim of this study is to correlate between the type of infection and antibiotic resistance. This aim is reached via surveying many infections caused by many bacteria, then performing antibiotic susceptibility test.
Preparation of Culture media
Preparation of culture media was according to the instruction manual supplied with each medium.
Collection and Identification of Bacterial Samples
The samples were collected from the hospital and bridged directly to the laboratory for identification. The identification of bacterial samples was carried out depending on the cultural characteristics and biochemical tests according to (Mac Faddin, 2000).
Procedure for Performing the Disc Diffusion Test
Inoculum Preparation
Direct Colony Suspension Method
As a convenient alternative to the growth method, the inoculums were prepared by making a direct broth or saline suspension of isolated colonies selected from a 18- to 24-hour agar plate (a nonselective medium, such as blood agar or nutrient agar, has been used). The suspension is adjusted to match the 0.5 McFarland turbidity standard, using saline and a vortex mixer.
Inoculation of Test Plates
Optimally, within 15 minutes after adjusting the turbidity of the inoculums suspensions, a sterile cotton swab is dipped into the adjusted suspension. The swab was rotated several times and pressed firmly on the inside wall of the tube above the fluid level. This is necessary to remove excess inoculum from the swab.
The dried surface of a Mueller-Hinton agar plates was inoculated by streaking the swab over the entire sterile agar surface. This procedure is repeated by streaking two more times, rotating the plate approximately 60° each time to ensure an equal distribution of inoculum. As a final step, the rim of the agar is swabbed. The plate lids were left ajar for 3 to 5 minutes, but no more than 15 minutes, to allow for any excess surface moisture to be absorbed before applying the drug impregnated disks.
Application of discs to inoculated agar plates
Antibiotics discs produced by bioanalysa company were used. The predetermined battery of antimicrobial discs is dispensed onto the surface of the inoculated agar plate. Each disc was pressed down to ensure complete contact with the agar surface. Then, the plates were inverted and placed in an incubator set to 37°C within 15 minutes after the discs were applied.
The correlation of infection with the type of bacteria
The 91 Bacterial isolates from different infections obtained from Al-Sader medical city in Al-Najaf province over the period from 2-7-2016 to 15-9-2016, were identified and correlated with the site of infection as shown below in table 1.
Table (1):
Distribution of bacteria and infections
Infection Bacteria |
Wound |
UTI |
Otitis media |
Intestinal Disease |
Total |
Percentage |
|---|---|---|---|---|---|---|
E.coli |
0 |
11 |
0 |
14 |
25 |
27% |
Staphylococcus |
10 |
7 |
5 |
0 |
22 |
24% |
Proteus |
9 |
10 |
8 |
0 |
27 |
30% |
Enterococcus |
8 |
9 |
0 |
0 |
17 |
19% |
total |
27 |
37 |
13 |
14 |
91 |
100% |
The identification of bacterial samples showed that most of the isolates (30%) were Proteus folled by E. coli (27%), Staphylococcus (24%) , and Enterococcus (19%).
The results showed that UTI is mainly caused by E. coli. This goes in agreement with (Nicolle, 2008) who showed that E. coli is the cause of 80–85% of urinary tract infections, with Staphylococcus saprophyticus as a causative agent in 5–10% of UTI. Most of wound infections has been shown to be caused by Staphylococcus in the present study. This agrees with (Toshkova et al., 2001; Flora, 2002; Baggett et al., 2004) in explaining that Staphylococcus aureus, is the common cause of wound infections.
Correlation between bacterial infections and gender
The type of infection has been correlated with the gender of the patient and with the type of bacteria as shown in table 2.
Table (2):
Correlation between bacterial infections and patient gender
| Bacteria Infection | E.coli | Staphylococci | Proteus | Enterococci | Total | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| male | female | male | female | male | female | male | female | male | female | ||
| Wound | – | – | 8 | 2 | 8 | 1 | 6 | 2 | 22 | 5 | |
| UTI | 4 | 7 | 2 | 5 | 3 | 7 | 2 | 7 | 11 | 26 | |
| Otitis media | – | – | 3 | 2 | 2 | 6 | – | – | 5 | 8 | |
| Intestinal Disease | 8 | 6 | – | – | – | – | – | – | 8 | 6 | |
| Total & percentage | 12 48% |
13 52% |
13 59% |
9 41% |
13 48% |
14 52% |
8 47% |
9 53% |
46 50.5% |
45 49.5% |
|
The results showed that the ratio of male to female patients is tend to be equal regardless the type of infections, but, when we consider the type of infection it is clear that males are significantly more affected by wound infections than females as 81.5% of wound infections appeared in mlase. This could be due to the outdoor working environment of the males which makes more exposed to have injuries than females and, consequently, having wound infections more frequently than females. On the other hand, the results showed that UTI is more frequent in females than in males as 70% of UTI appeared in urine samples taken from females. This is most likely due to the anatomy of the female urethra and genital area which makes female more vulnerable to have UTI than males. This agrees with the findings that has been shown previously that UTI is more common in females (Stamm, 1991; Foxman et al., 2000; Harrington and Hooton, 2000; Chittagong, 2011).
Correlation between the type of infection and antibiotic resistance in E.coli
E. coli was isolated only from urine and stool as a causative agent of UTI and intestinal infection. The antibiotic resistance against 13 different antibiotics was evaluated and correlated to the type of infection which are UTI and intestinal infections. The resistance percentage for each antibiotic was calculated, then the mean of resistance percentages was calculated for both types of the previously mentioned infections (Table 3). The results showed approximate similarity of antibiotic resistance in both UTI and intestinal infections (Figure 1).
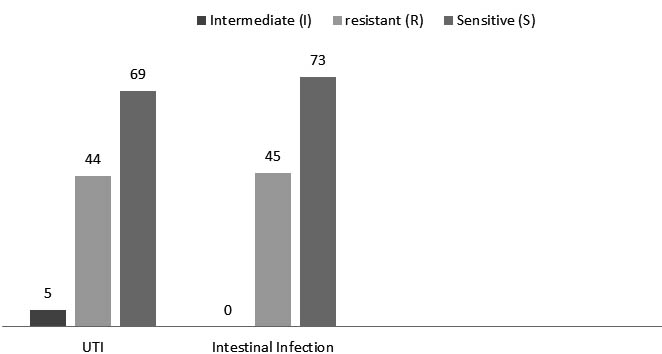
Fig. 1. The type of infection and antibiotic resistance in E. coli
Table (3):
Correlation between the type of infection and antibiotic resistance in E. coli
| Antibiotic Infection | Ampicillin 61% | Gentamycin 29% | Amikacin 33% | Cotrimoxazole 67% | Chloramphenic-ol 10% | Nitrofurantoin 36% | Ceftazidime 24% | Ceftriaxone 29% | Amoxyclav 31% | Nalidixic Acid 45% | Ciprofloxacin 45% | Cephalothin 29% | Ampicillin/cloxacillin 54% | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| UTI | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | |
| NO | 4 | – | 7 | 6 | – | 5 | 7 | – | 4 | 5 | – | 6 | 9 | – | 2 | 8 | – | 3 | 6 | 3 | 2 | 7 | – | 4 | 7 | 2 | 2 | 6 | – | 5 | 5 | – | 6 | 6 | – | 5 | 7 | – | 4 | |
| % | 36 % |
64 % |
54 % |
46 % |
64 % |
36 % |
46 % |
54 % |
88 % |
12 % |
72 % |
28 % |
55 % |
33 % |
12 % |
64 % |
36 % |
64 % |
24 % |
12 % |
54 % |
46 % |
46 % |
54 % |
54 % |
46 % |
64 % |
36 % |
||||||||||||
| Intestinal infection | NO | 6 | – | 8 | 12 | – | 2 | 10 | – | 4 | 3 | – | 11 | 13 | – | 1 | 8 | – | 6 | 9 | – | 5 | 11 | – | 3 | 7 | – | 7 | 8 | – | 6 | 9 | – | 5 | 12 | – | 2 | 4 | – | 10 |
| % | 43 % |
57 % |
88 % |
12 % |
71 % |
29 % |
21 % |
79 % |
93 % |
7 % |
57 % |
43 % |
64 % |
36 % |
79 % |
21 % |
50 % |
50 % |
57 % |
43 % |
64 % |
36 % |
88 % |
12 % |
29 % |
71 % |
||||||||||||||
The most resistance was seen to ampicillin with a percentage of 64% in UTI. This agrees with (Chittagong, 2011) who showed that the most resistance rates for E. coli detected from urine culture were Ampicillin, Doxycycline, Cephalexin, Cephradine, Cotrimoxazole, Cefixime, Ceftroxone and Ciprofloxacin except for Cefuraoxime to which E. coli was significantly sensitive. The antibiotic resistance pattern by E.coli has been shown to be 40% for Amoxicillin, 23% for Cephradine, 21% for Cotrimoxazole, 6% for Doxycycline, 4% for Cefixime, 2% for Cephalexin, 2% for Ceftroxone and 2% for Ciprofloxacin (Bhowmick and Rashid, 2004; Mohammadi et al., 2010; Manikandan et al., 2011). (Omoregie et al., 2011) showed that 60% of E. coli isolates were susceptible to Gentamycin.
Correlation between the type of infection and antibiotic resistance in Staphylococcus spp.
Staphylococcus isolated from wound infections has been shown to be more resistant against 12 antibiotics if compared with isolates from otitis media and UTI (figure 2). The most resistance was seen to Aztreonam antibiotic with a percentage of 83%, while the most effective antibiotic was Vancomycin with a resistance percentage of (8%) (table 4).
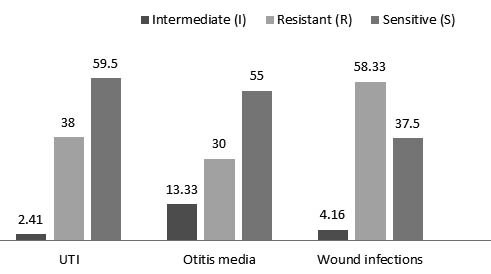
Fig. 2. The type of infection and antibiotic resistance in Staphylococcus spp.
Table (4):
Correlation between the type of infection and antibiotic resistance in Staphylococcus spp
| Antibiotic Infection | Amoxicillin+ Clavulanic acid 33% | Ceftriaxone 41% | Gentamycin 16% | Vancomycin 8% | Azactam 83% | Ciprobay 10% | Ceftazidime 52% | Azithromycin 53% | Ampicillin/cloxacillin 25% | Chloramphenicol 38% | Cephalothin 54% | Erythrocin 54% | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| UTI | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | |
| NO | 5 | 2 | – | 6 | – | 1 | 5 | 2 | 6 | – | 1 | – | – | 7 | 7 | – | – | 3 | – | 4 | 5 | – | 2 | 1 | – | 6 | 6 | – | 1 | 2 | – | 5 | 4 | – | 3 | ||
| % | 71
% |
29
% |
0
% |
86
% |
14
% |
71
% |
– | 29
% |
86
% |
14
% |
100
% |
100
% |
0
% |
43
% |
57
% |
71
% |
29
% |
14
% |
86
% |
86
% |
14
% |
29
% |
71
% |
57
% |
43
% |
||||||||||||
| Otitis media | NO | 4 | – | 1 | 3 | – | 2 | 4 | 1 | – | 4 | 1 | – | – | 2 | 3 | 4 | 1 | – | 3 | – | 2 | 1 | – | 4 | 4 | – | 1 | – | 2 | 3 | 4 | 1 | – | 2 | 1 | 2 |
| % | 80
% |
20
% |
60
% |
40
% |
80
% |
0
% |
80
% |
20
% |
0
% |
40
% |
60
% |
80
% |
20
% |
0
% |
60
% |
40
% |
20
% |
80
% |
80
% |
20
% |
40
% |
60
% |
80
% |
20
% |
0
% |
40
% |
20
% |
40
% |
|||||||||
| Wound | NO | 2 | – | 8 | 3 | – | 7 | 8 | 2 | 8 | 1 | 1 | – | 1 | 9 | 4 | – | 6 | 4 | – | 6 | 5 | – | 5 | 5 | – | 5 | 4 | 2 | 4 | 1 | – | 9 | 1 | 1 | 8 | |
| % | 20
% |
80
% |
30
% |
70
% |
80
% |
– | 20
% |
80
% |
10
% |
10
% |
10
% |
90
% |
40
% |
60
% |
40
% |
60
% |
50
% |
50
% |
50
% |
50
% |
40
% |
20
% |
40
% |
10
% |
90
% |
10
% |
10
% |
80
% |
|||||||||
(Diekema et al., 2001) who shwed that more than 80% of coagulase-negative staphylococcal isolates were resistant to methicillin and semisynthetic penicillins. (Liu et al., 2011) showed that Community-acquired MRSA (CA-MRSA) isolates often maintain susceptibility to tetracyclines (tetracycline, doxycycline, minocycline, tigecycline). The results by (Omoregie et al., 2011) revealed that Staphylococcus aureus have great suseptibility to Amoxicillin (96.2%). Schwalbe et al,. (1987) showed that Staphylococcus spp. are resistance to Vancomycin.
Correlation between the type of infection and antibiotic resistance in Enterococcus spp.
Enterococcus was isolated from UTI and wound infections. The results revealed that isolates of UTI were more resistant than those isolated from wounds when tested against 14 antibiotics (figure 3) (table 5). (Karmarkar et al., 2004) showed that 77.23% of enterococcal isolates were resistant to more than six drugs. (Herman and Gerding, 1991) studied serious enterococcal infections (e.g., bacteraemia and endocarditis) and showed that such infections require treatment with a bactericidal combination of antibiotics that should include a penicillin (ampicillin or penicillin G) to which the isolates were susceptible and an aminoglycoside (gentamicin or streptomycin) to which they were resistant. (Courvalin, 2006) showed that more than 55% of enterococcal isolates in ICUs of more than 300 hospitals were vancomycin-resistant.
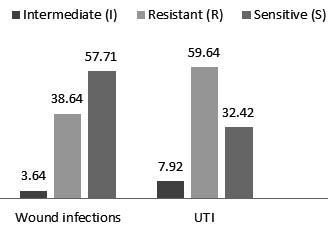
Fig. 3. The type of infection and antibiotic resistance in Enterococcus spp.
(Choudhury et al., 2015) revealed that there were rare isolates that lack resistance against gentamicin and streptomycin, while high level of resistance were seen against amikacin and kanamycin. And Results obtained by (Sapkota et al., 2007) revealed significant resistance by Enterococcus against erythromycin and tetracycline.
Table (5):
Correlation between the type of infection and antibiotic resistance in Enterococcus spp
| Antibiotic Infection | Azactam 51% | Ciprofloxacin 22% | Cephalothin 72% | Ampicillin 65% | Co-trimoxazole 57% | Gentamycin 11% | Ceftazidime 46% | Vancomycin 89% | Azithromycin 0% | Chloramphenicol 0% | Amoxyclav 71% | Nitrofurantoin 84% | Erythrocin 88% | Clarithromycin 69% | |||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Wound | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | |
| NO | 6 | 1 | 1 | 6 | 2 | – | 1 | – | 7 | 3 | – | 5 | 7 | – | 1 | 8 | – | – | 7 | – | 1 | – | – | 8 | 7 | 1 | – | 8 | – | – | 2 | – | 6 | 3 | – | 5 | 2 | – | 6 | 5 | – | 3 | |
| % | 74
% |
13
% |
13
% |
75
% |
25
% |
0
% |
12
% |
88
% |
37
% |
63
% |
87
% |
13
% |
100
% |
– | 0
% |
87
% |
– | 13
% |
– | – | 100
% |
87
% |
13
% |
0
% |
100
% |
0
% |
25
% |
– | 75
% |
37
% |
– | 63
% |
25
% |
– | 75
% |
62
% |
– | 38
% |
|||||
| UTI | NO | 1 | – | 8 | 5 | – | 4 | 4 | – | 5 | 3 | – | 6 | – | – | 9 | 4 | 3 | 2 | 1 | 1 | 7 | – | 2 | 7 | 9 | – | – | 9 | – | – | 1 | 2 | 6 | 4 | 2 | 3 | – | – | 9 | – | – | 9 |
| % | 11
% |
– | 89
% |
56
% |
– | 44
% |
44
% |
– | 56
% |
33
% |
– | 67
% |
– | – | 100
% |
44
% |
34
% |
22
% |
11
% |
11
% |
78
% |
– | 22
% |
78
% |
100
% |
– | 0
% |
100
% |
– | 0
% |
11
% |
22
% |
67
% |
44
% |
22
% |
34
% |
– | – | 100
% |
100
% |
|||
Table (6):
Correlation between the type of infection and antibiotic resistance in Proteus spp.
| Antibiotic Infection | Ceftriaxone 48% | Gentamycin 54% | Cephalothin 58% | Ceftazidime 41% | Amoxyclav 61% | Ciprofloxacin 3% | Amikacin 42% | Chloramphenicol 58% | Ampicillin/cloxacillin 66% | Cefixime 57% | Cefotaxime 50% | |||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Otitis Media | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | s | i | r | |
| NO | 6 | – | 2 | 7 | – | 1 | 7 | – | 1 | 6 | – | 2 | 5 | – | 3 | 8 | – | 0 | 5 | – | 3 | 7 | – | 1 | 4 | – | 4 | 6 | – | 2 | 6 | – | 2 | |
| % | 75 % |
25 % | 87
% |
13
% |
87
% |
13
% |
75
% |
25
% |
62
% |
38
% |
100
% |
0
% |
62
% |
38
% |
87
% |
13
% |
50
% |
50
% |
75
% |
25
% |
75
% |
25
% |
||||||||||||
| UTI | NO | 7 | – | 3 | 3 | – | 7 | 2 | – | 8 | 8 | – | 2 | 1 | – | 9 | 9 | – | 1 | 8 | – | 2 | 2 | – | 8 | 3 | – | 7 | 2 | – | 8 | 3 | – | 7 |
| % | 70
% |
30
% |
30
% |
70
% |
20
% |
80
% |
80
% |
20
% |
10
% |
90
% |
90
% |
10
% |
80
% |
20
% |
20
% |
80
% |
30
% |
70
% |
20
% |
80
% |
30
% |
70
% |
||||||||||||
| Wound | NO | 1 | – | 8 | 2 | – | 7 | 1 | – | 8 | 2 | – | 7 | 4 | – | 5 | 9 | – | – | 3 | – | 6 | 1 | – | 8 | 2 | – | 7 | 3 | – | 6 | 4 | – | 5 |
| % | 11
% |
89
% |
22
% |
78
% |
11
% |
89
% |
22
% |
78
% |
44
% |
56
% |
100
% |
0
% |
33
% |
67
% |
11
% |
89
% |
22
% |
78
% |
33
% |
67
% |
44
% |
56
% |
||||||||||||
Correlation between the type of infection and antibiotic resistance in Proteus spp.
The isolates of Proteus spp. were obtained from wounds, UTI and otitis media. The most resistant isolates against 11 antibiotics were those isolated from wound infections (figure 4) The most resistance where against Ampicillin/cloxacillin (table 6). (Feglo et al., 2010) showed that all proteus species were resistant to chloramphenicol, ampicillin and co-trimoxazole. However, 70 – 90 % of P. mirabilis and P. vulgaris isolates exhibited resistance to ampicillin, cotrimoxazole, tetracycline and chloramphenicol. (Newman et al., 2006) revealed that Proteus spp. isolates exhibited high antimicrobial resistance against tetracycline (85 %), chloramphenicol (82.5 %), co-trimoxazole (81 %) and ampicillin (77 %).
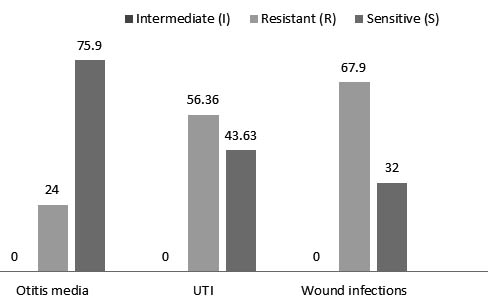
Fig. 4. The type of infection and antibiotic resistance in Proteus spp.
- E. coli was the main cause of intestinal diseases.
- E. coli was the most frequent bacteria casing UTI, Proteus spp. otitis media, and Staphylococcus spp. was the main cause of wound infections.
- In males the infections with Staphylococcus spp. were more than in females, while infections with E. coli, Proteus spp. and Enterococcus spp. were more in females than in males.
- In E. coli the resistance to antibiotics in UTI and intestinal disease was approximately the same.
- In Staphylococcus spp. and Proteus spp. wounds infections exhibit more resistant than other types of infections.
- In Enterococcus spp. UTI infections exhibit more resistant than wounds infections.
ACKNOWLEDGMENTS
None.
- Al-Shamarti, M.J., 2010. molecular evaluation of B-lactam resistance genes in klebsiella spp isolated from clinical cases in Al-Najaf province. Department of Biology. University of Kufa
- Baggett, H.C., Hennessy, T.W., Rudolph, K., Bruden, D., Reasonover, A., Parkinson, A., Sparks, R., Donlan, R.M., Martinez, P., Mongkolrattanothai, K., Community-onset methicillin-resistant Staphylococcus aureus associated with antibiotic use and the cytotoxin Panton-Valentine leukocidin during a furunculosis outbreak in rural Alaska. The Journal of infectious diseases 2004; 189: 1565-1573.
- Bennett, J., Feibelman, T., Fungal bacterial interactions. Fungal Associations. 2001; Springer, pp. 229-242.
- Bennett, P., Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria. British journal of pharmacology 2008; 153.
- Bhowmick, B., Rashid, H., Prevalence and antibiotic susceptibility of E. coli isolated from urinary tract infection (UTI) in Bangladesh. Pak. J. Biol. Sci 2004; 7: 717-720.
- Chittagong, B., A study of antibacterial susceptibility and resistance pattern of E. coli causing urinary tract infection in Chittagong, Bangladesh. Asian Journal of Biological Sciences 2011; 4: 548-555.
- Choudhury, B., Banik, A., Lyngdoh, V.W., Gurung, J., Khyriem, A.B., Rajkhowa, P., High Level Aminoglycoside Resistance among Clinical Enterococcal Isolates in a Tertiary Care Centre of North East India. International Journal of Health Sciences and Research (IJHSR) 2015; 5: 140-145.
- Clardy, J., Fischbach, M., Currie, C., The natural history of antibiotics. Current biology : CB, 2009; 19: R437-R441.
- Courvalin, P., Vancomycin resistance in gram-positive cocci. Clinical Infectious Diseases, 2006; 42: S25-S34.
- D’costa, V.M., McGrann, K.M., Hughes, D.W., Wright, G.D., Sampling the antibiotic resistome. Science, 2006; 311: 374-377.
- D’Costa, V.M., King, C.E., Kalan, L., Morar, M., Sung, W.W., Schwarz, C., Froese, D., Zazula, G., Calmels, F., Debruyne, R., Antibiotic resistance is ancient. Nature, 2011; 477: 457-461.
- Diekema, D., Pfaller, M., Schmitz, F., Smayevsky, J., Bell, J., Jones, R., Beach, M., Group, S.P., 2001.
- Survey of infections due to Staphylococcus species: frequency of occurrence and antimicrobial susceptibility of isolates collected in the United States, Canada, Latin America, Europe, and the Western Pacific region for the SENTRY Antimicrobial Surveillance Program, 1997–1999. Clinical Infectious Diseases 32: S114-S132.
- Equi, A., Balfour-Lynn, I., Bush, A., Rosenthal, M., Long term azithromycin in children with cystic fibrosis: a randomised, placebo-controlled crossover trial. The Lancet, 2002; 360: 978-984.
- Feglo, P.K., Gbedema, S.Y., Quay, S.N.A., Adu-Sarkodie, Y., Opoku-Okrah, C., Occurrence, species distribution and antibiotic resistance of Proteus isolates: A case study at the Komfo Anokye Teaching Hospital (KATH) in Ghana. Int J Pharm Sci Res, 2010; 1: 347-352.
- Flora, N.S., Evaluating and managing open skin wounds: colonization versus infection. AACN clinical issues, 2002; 13: 382-397.
- Foxman, B., Barlow, R., D’arcy, H., Gillespie, B., Sobel, J.D., Candida vaginitis: Self Reported Incidence and Associated Costs. Sexually transmitted diseases, 2000; 27: 230-235.
- Gold, H.S., Moellering Jr, R.C., Antimicrobial-drug resistance. New England Journal of Medicine, 1996; 335: 1445-1453.
- Harrington, R.D., Hooton, T.M., Urinary tract infection risk factors and gender. The journal of gender-specific medicine: JGSM: the official journal of the Partnership for Women’s Health at Columbia, 2000; 3: 27-34.
- Herman, D.J., Gerding, D.N., Screening and treatment of infections caused by resistant enterococci. Antimicrobial agents and chemotherapy, 1991; 35: 215.
- Kardos, N., Demain, A.L., Penicillin: the medicine with the greatest impact on therapeutic outcomes. Applied microbiology and biotechnology, 2011; 92: 677.
- Karmarkar, M., Gershom, E.S., Mehta, P., Enterococcal infections with special reference to phenotypic characterization & drug resistance. Indian Journal of Medical Research, 2004; 119: 22.
- Liu, C., Bayer, A., Cosgrove, S.E., Daum, R.S., Fridkin, S.K., Gorwitz, R.J., Kaplan, S.L., Karchmer, A.W., Levine, D.P., Murray, B.E., Clinical practice guidelines by the Infectious Diseases Society of America for the treatment of methicillin-resistant Staphylococcus aureus infections in adults and children. Clinical infectious diseases, 2011; 52: e18-e55.
- Mac Faddin, J., Biochemical Tests for Identification of Medical. Bacteria. Williams and Wilkins. London 2000.
- Manikandan, S., Ganesapandian, S., Singh, M., Kumaraguru, A., Emerging of multidrug resistance human pathogens from urinary tract infections. Curr. Res. Bacteriol, 2011; 4: 9-15.
- Mazodier, P., Davies, J., Gene transfer between distantly related bacteria. Annual review of genetics 1991; 25: 147-171.
- Mohammadi, M., Ghasemi, E., Mokhayeri, H., Pournia, Y., Boroun, H., Antimicrobial resistance patterns of E. coli detected from hospitalized urine culture samples. Asian Journal of Biological Sciences 2010; 3: 195-201.
- Newman, M., Frimpong, E., Asamoah-Adu, A., Sampene-Donkor, E., Resistance to antimicrobial drugs in Ghana. The Ghanaian-Dutch Collaboration for Health research and Development. Project number 2001. GD/07. Technical report series, 2006; 5: 8-26.
- Nicolle, L.E., Uncomplicated urinary tract infection in adults including uncomplicated pyelonephritis. Urologic Clinics, 2008; 35: 1-12.
- Omoregie, E., Osagie, A., Iruolaje, E., In vitro antioxidant activity and the effect of methanolic extracts of some local plants on nutritionally stressed rats. Pharmacologyonline, 2011; 1: 23-56.
- Sapkota, A.R., Curriero, F.C., Gibson, K.E., Schwab, K.J., 2007. Antibiotic-resistant enterococci and fecal indicators in surface water and groundwater impacted by a concentrated swine feeding operation. Environmental Health Perspectives 115: 1040.
- Stamm, W.E., Catheter-associated urinary tract infections: epidemiology, pathogenesis, and prevention. The American journal of medicine, 1991; 91: S65-S71.
- Toshkova, K., Annemüller, C., Akineden, Ö., Lämmler, C., The significance of nasal carriage of Staphylococcus aureus as risk factor for human skin infections. FEMS microbiology letters, 2001; 202: 17-24.
- Wilson, J., 2006. Infection control in clinical practice. Elsevier Health Sciences.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


