ISSN: 0973-7510
E-ISSN: 2581-690X
In the present study, bioflocculant derived from bacteria are widely explored in bioremediation such as removing heavy metal and inhibiting biofilm formation. The bacteria isolated from sewage identified as Proteus mirabilis using 16S rRNA gene made up of D-Mannose, D-Fructose, D-Sucrose, and D-Galactose could be identified by Gas liquid Chromatography Mass Spectrometry (GCMS). Fourier transform infrared (FTIR) analysis of the bioflocculant indicated the presence of –COOH, -OH, and -NH groups which showed the characteristic of polysaccharide and protein. The removing percentage of heavy metals achieved by the precipitated bioflocculant were 82.63 ± 1.20, 72.076 ± 0.42, 57.36 ± 1.05, and 44.7 ± 1.053%, respectively for Zn2+, Cd2+, Cu2+, and Hg2+ at optimum conditions. Bioflocculant were exhibited excellent biofilm inhibition against P. aeruginosa where maximum reduction (73%) were achieved at the minimum amount of 80µg/ml dose. Similarly, maximum reduction (80%) were achieved at the minimum amount of 70µg/ml dose for S. aureus. It was concluded that the bioflocculant produced have the potential in metal removal and biofilm inhibition.
Heavy metals, Bioflocculant, Flocculation, Antibiofilm, Polysaccharides.
Microorganisms produced their metabolites are called as bioflocculant during their growth. Non-toxic nature of the bioflocculant are alternative to synthetic flocculant involving in bioremediation and biomedical application1,2. Several bacterial species are capable of producing bioflocculant like Alcaligenes latus, Bacillus firmus3, Aspergillus sp.4, Streptomyces and Cellulomonas5. The bioflocculants made up of microbial metabolic products such as glycoprotein and complex carbohydrates6. It is mainly affected by environmental factors such pH, Temperature, dosage of inoculum, carbon and nitrogen sources7,8. These key factors affecting bioflocculant production varies for different microbial species.
Generally, bacteria produces a variety of polysaccharides which are classified based on location such as, intracellular, extracellular, and capsular polysaccharides. They are produced via different biosynthesis pathways9. In addition, polysaccharides can be used as bioflocculant, heavy metal removing agents10, bioadsorbents, encapsulating materials. They are high molecular weight biopolymers such as polysaccharide, glycoprotein, glycolipids, and protein flocculants11,12. They described that the functional groups identification of the bioflocculant are significant factor for the flocculating activity13.
For decades, attention have focused to remove hazardous pollutants to the environment. But, currently practicing chemical and physical methods for removing the metal from the contaminated environments are not environmentally friendly, costly, non-specific14,15. In addition, many of these processes are not successful16. Therefore, bioflocculant is the best alternative to remove contaminants. Various uses of bioflocculant in bioremediation processes are involved to remove the toxic materials, and heavy metals, synthesis of nanoparticles and biomass recovery2,17-19. In addition, bioflocculant could be acted as a way of preventing biofilm formation.
Considering the significance of bioflocculant in metal sequestration and antibiofilm activity, the present study was focused at searching the strain producing the bioflocculant and simultaneously acting as an antibiofilm agent under optimum conditions. Also, amount of metal uptake and biofilm inhibition percentage of the bioflocculant was estimated. Finally, it emphasized to search over the structural and functional components responsible for flocculation and antibiofilm formation.
Identification of Bacterial strains
Totally, 20 bacterial strains were previously isolated from the wastewater. Potential strain showing highest flocculating activity were screened for further research. Test bacteria were recognized as Proteus mirabilis through partial sequencing of 16S rRNA genes. Nucleotide sequences were finally deposited in GenBank (MG996004).
Production and partial purification of bioflocculant
Bioflocculant producing media was prepared based on the methods of Zhang et al.20. Components of the media such as KH2PO4 (2.0 g), Glucose (15.0 g), (NH4)2SO4 (0.2 g), Yeast extract (0.5 g), K2HPO4 (5.0 g), MgSO4 · 7H2O (0.2 g), NaCl (0.1 g), were dissolved in 1L of distilled water (pH=7) under optimum conditions. 0.5 ml culture of P. mirabilis isolated from wastewater, inoculated into the medium and kept it for 72 hrs incubation at 30°C, 160 rpm/min. The broth was centrifuged (8000׳g, 20 min) to pelletize the cells. The upper portion of the bioflocculant were added with thrice the volume of ethanol. This washing step was repeated several times to obtain the precipitate. Thereafter, they were assayed for flocculation activity.
Characteristics of the bioflocculant
Carbohydrate content of the bioflocculant using glucose as the standard solution were measured using Phenol-sulfuric acid method21. Lowry method was assessed in measuring the protein content of the bioflocculant22. The monomeric unit of the bioflocculant was obtained by hydrolysis of the bioflocculant with HCl at 121°C for 2 hrs. They were further subjected to the analysis of FT-IR. The scanning electron microscopy (SEM: SUPRA 55 SAPPHIRE; Germany) image of the bioflocculant was obtained.
Gas Chromatograph-Mass Spectrometer
The extracted samples were subjected to GC/MS analysis (TQ8030, Shimadzu, Japan). For identifying sugars components, carrier gas used was helium the with a flow rate of 1.0 mL min-1, maintaining the temperature of 300°C with an electron ionization mode (70 eV). Compounds were identified by the NIST1 1library database using chromatograms.
Heavy metal removal rate
The bioflocculant (0.8mg/100ml) was applied for the bioflocculation test against heavy metal solutions containing Cd2+, Zn2+, Cu2+, and Hg2+ at the concentrations of 0-50 mg L-1, respectively. The amount of heavy metals were identified by inductively coupled plasma atomic emission spectroscopy (ICP-AES; iCAP 7000, Thermo Fischer Scientific, Bremen, Germany). All experiments were repeated thrice. The percentage of heavy metal removal was measured based on the following equation:
% R= (Ci –Cf) /m
Where, Ci – amount of heavy metal of test solution initially; Cf – amount of heavy metal of test solution finally; m (g) is the weight of bioflocculant.
Antibiofilm assay
Crystal violet assay was performed which was examined to quantify the biofilm inhibition percentage23.
Biofilm of Pseudomonas aeruginosa and Staphylococcus aureus grown separately on glass slides in well plates were added with the bioflocculant. They kept the incubation for 24 hrs at 37°C and washed with phosphate buffered saline (0.1 M; pH 7.4) to discard non-adherent cells. In the same way, control were also subjected to the same procedure without the bioflocculant. The glass slides were observed under confocal microscopy (CLSM, LSM 510, Carl Zeiss, and Germany) after the Acridine orange staining. All the experiments were conducted thrice.
Production of bioflocculant
Totally, 20 bacterial strains were isolated from the sewage water. In which, strain showing highest flocculating activity were selected for bioflocculant production. The strain were identified as P. mirabilis (MG996004) through 16S rRNA sequence21. Phylogenetic tree based on 16S rRNA sequences were constructed (Fig. 1). Morphological structure of P. mirabilis were identified (Fig. 2). Totally, 2 g of bioflocculant was obtained from 1L fermentation broth under optimal conditions20. The production of bioflocculant was mainly controlled by the factors, such as the components of the culture medium and the cultivation conditions24. The key factors, such as the concentration of carbon sources, temperature, shaking speed, and nitrogen sources, pH, and inoculum size have to be focused on the flocculating activity of the bioflocculant by Proteus mirabilis25.
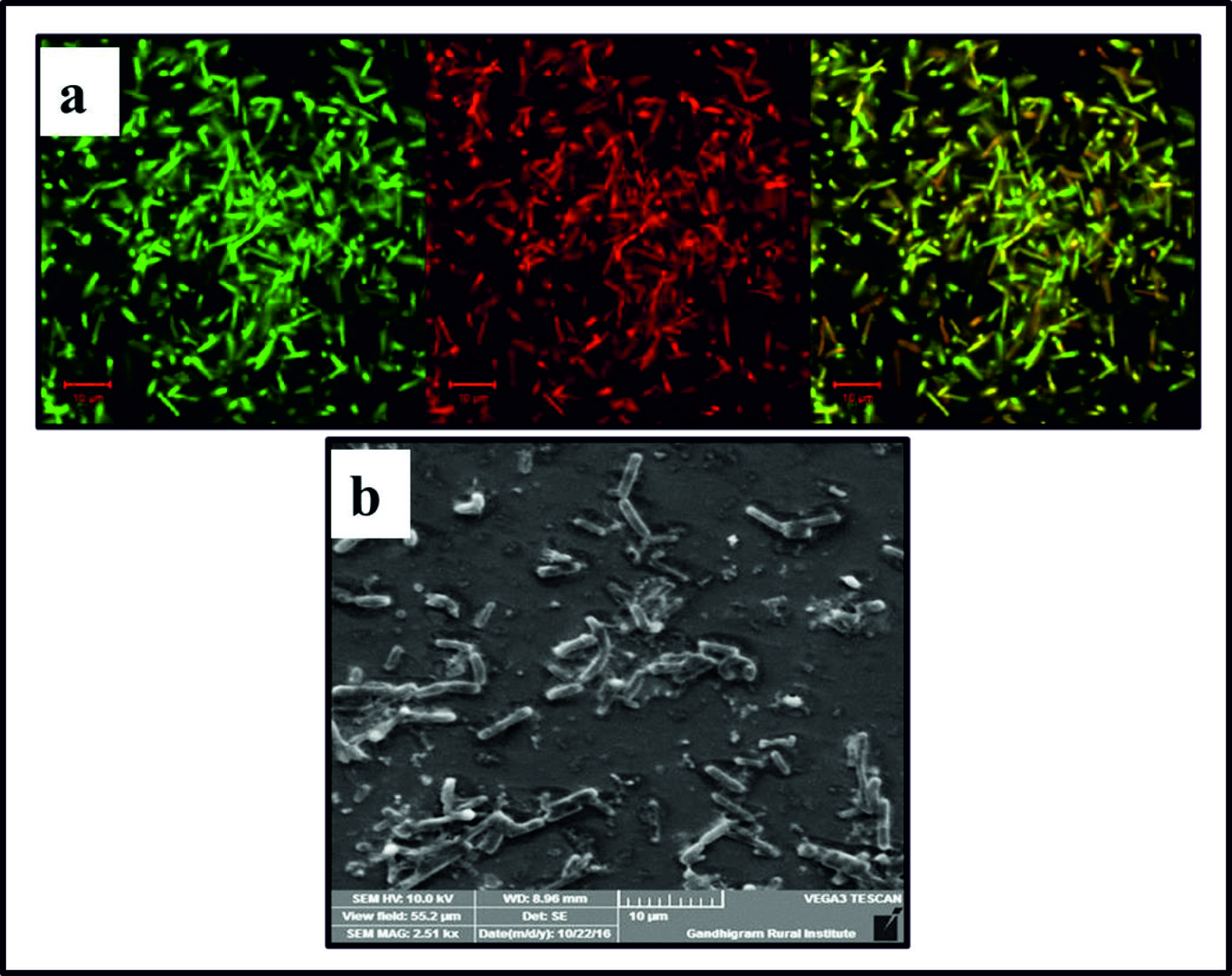 Fig. 2. a) Image of Gram negative bacteria Proteus mirabilis observed under the confocal microscope; b) SEM image of Gram negative bacteria Proteus mirabilis.
Fig. 2. a) Image of Gram negative bacteria Proteus mirabilis observed under the confocal microscope; b) SEM image of Gram negative bacteria Proteus mirabilis. Characteristics of bioflocculant
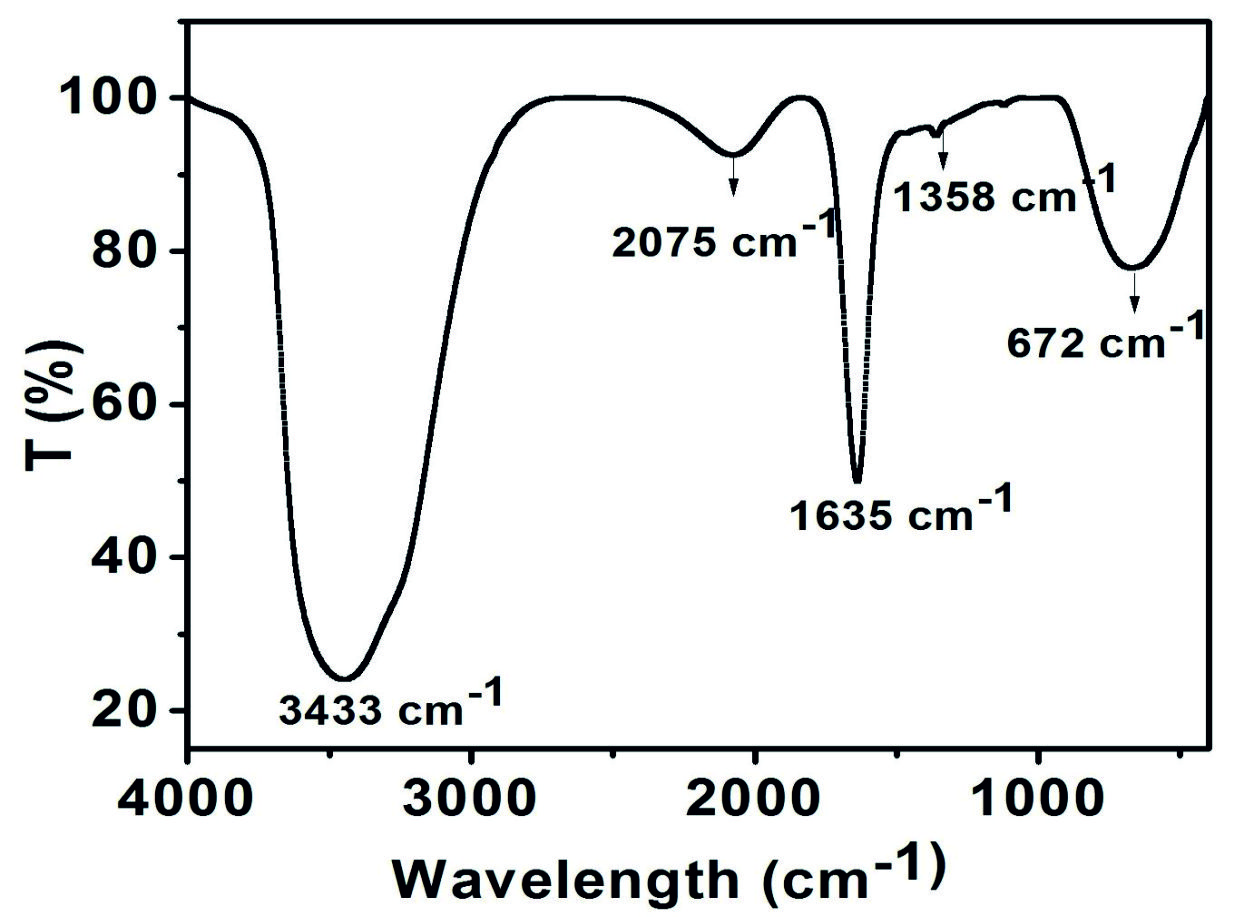
The bioflocculant produced from Proteus mirabilis (MG996004) included 68% (wt/wt) carbohydrate, and 15% (wt/wt) protein. These components were confirmed by FTIR spectroscopy by identifying the functional group of the bioflocculant (Fig. 3). The broad peak at 3433 cm-1, which was confirmed it as hydroxyl and amino group26. Sharp peak observed at 1635 cm-1 was attributed to the characteristic vibration of amide group. Weak peak at 1,358 cm-1 was correlated to the carboxylate group of the bioflocculant27. The adsorption band at 672 cm-1 could be related to the binding between sugar monomers28. Finally, spectrum revealed the peaks for polysaccharides, protein and identified the presence of the functional groups: hydroxyl (-OH), amide (-CO-NH) and carboxyl (-COO-)25. These functional groups act as a binding site for divalent cations, mainly involved in flocculating processes.
GC/MS analysis of bioflocculant was found the monomers, viz., D-Mannose, D-fructose, D-Sucrose, D-Galactose (retention times of 14.18, 15.98, 23.18, and 20.37 min, respectively) and compared with NIST Library (Fig. 4). The results indicated that the bioflocculant made up of four sugar monomers29, 30. HR-SEM showed the outer structure of the bioflocculant. Representative HR-SEM image showed well-dispersed, irregular shaped bioflocculant (Figs. 5(a)) with negatively charged zeta potential possessing best flocculating property22. The configuration of the bioflocculant were major factor involved in flocculating activity21. During flocculation, they bonded tightly and intertwined each other due to intermolecular interaction in between them22,31. The elemental analysis for the bioflocculant were shown tthe presence of C, K, Na, Mg, Cl, and O at particular proportions 29 (Fig.s 5(b)).
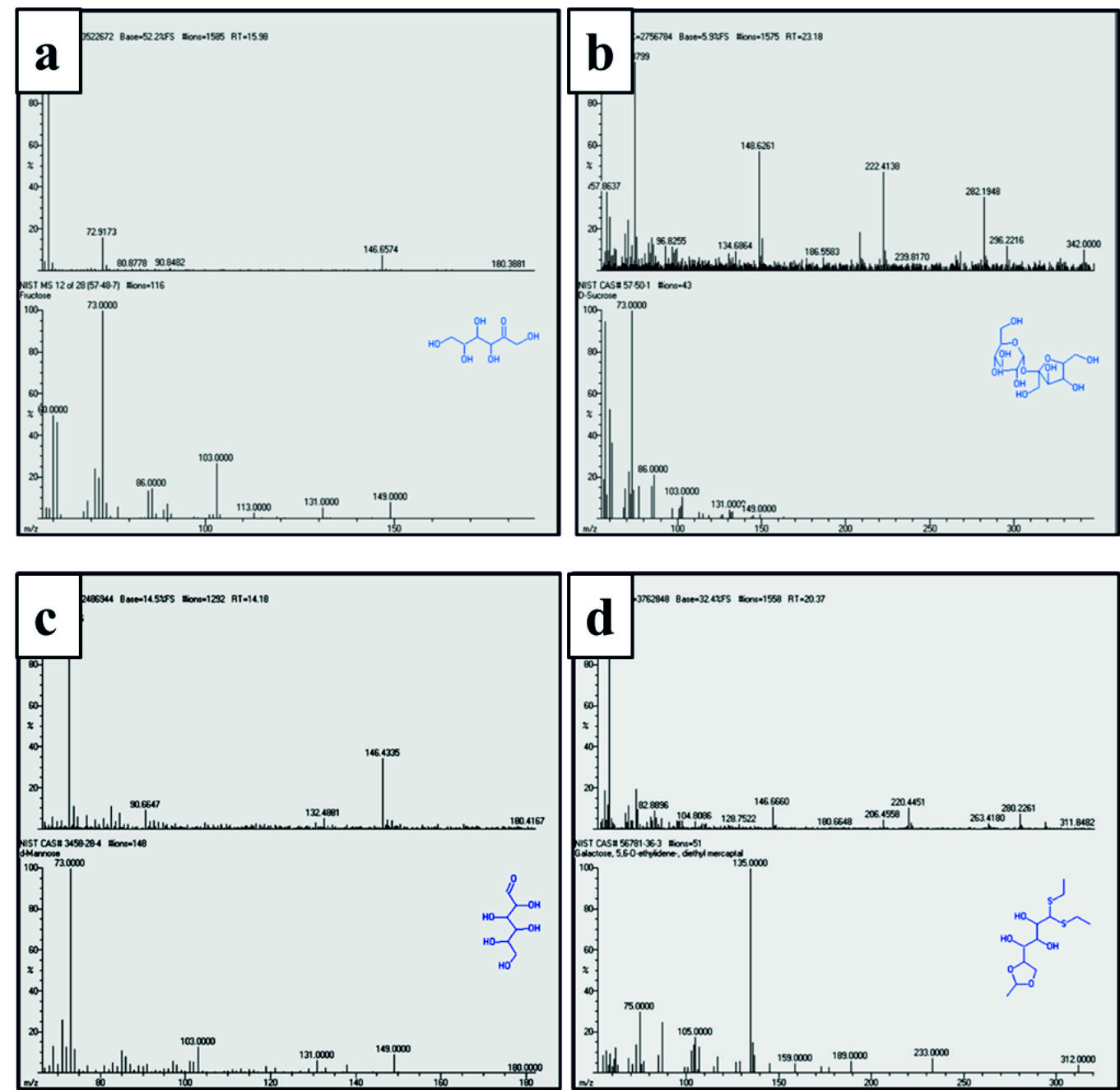 Fig. 4. GC/MS chromatogram of monomeric sugar units identified in the bioflocculant
Fig. 4. GC/MS chromatogram of monomeric sugar units identified in the bioflocculanta) D-Fructose, b) D-Sucrose, c) D-Mannose, d) D-Galactose.
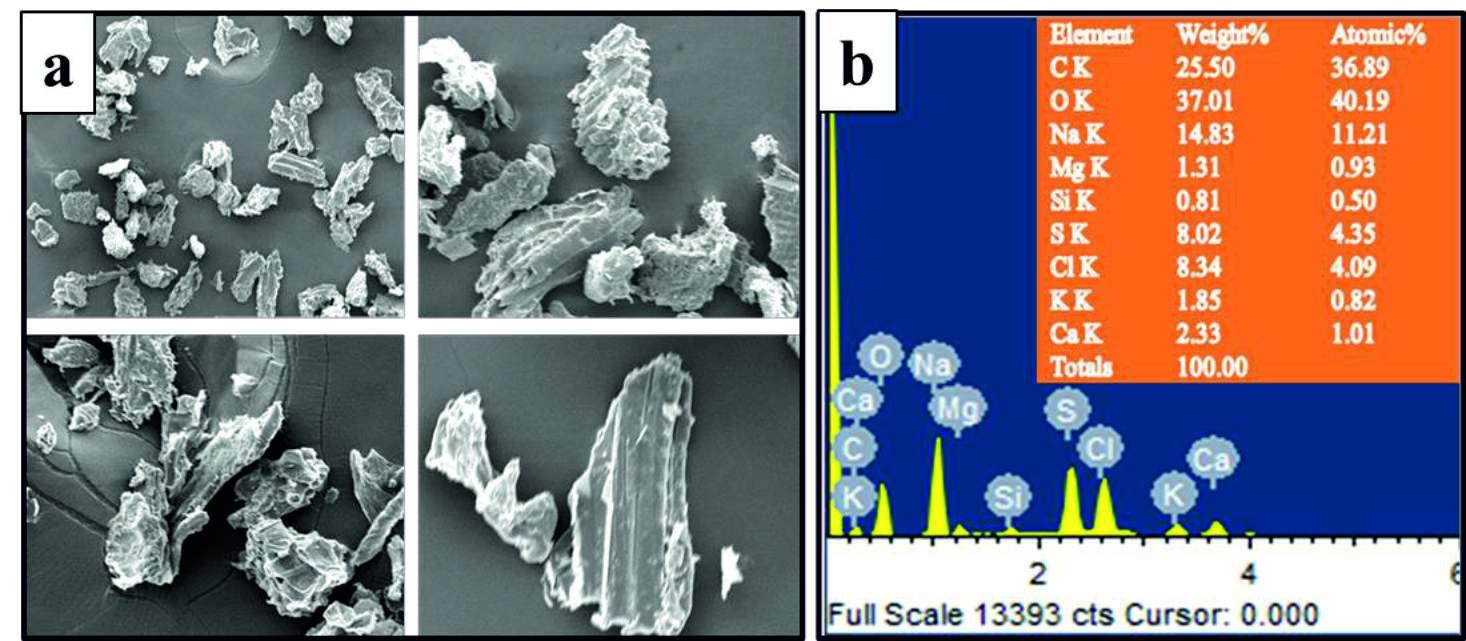 Fig. 5. (a) Irregular shape of the bio occulant; (b) SEM-EDAX spectra depicting the presence of C, K, O, Na, Ca, Mg, and Cl in the bio occulant.
Fig. 5. (a) Irregular shape of the bio occulant; (b) SEM-EDAX spectra depicting the presence of C, K, O, Na, Ca, Mg, and Cl in the bio occulant. Heavy metals removing efficiency
In the study, the metal ions such as Zn2+, Cd2+, Cu2+, and Hg2+ within the arrange of 0–50 mg L-1 in aqueous solutions could be effectively flocculated by the bioflocculant. The heavy metal removing percentage achieved by the precipitated bioflocculant was 82.63 ± 1.20, 72.076 ± 0.42, 57.36 ± 1.05, and 44.7 ± 1.053%, respectively for Zn2+, Cd2+, Cu2+, and Hg2+ at optimum conditions. When compare the treated sample with untreated sample, concentration of metals in treated sample was completely reduced that showed adsorption and bioflocculating ability of bioflocculant. In addition, concentration of bioflocculant reported was very less on comparing other reported literature32,33. It was interpreted that the electro-negative properties of the biological material and the metals regulated the metal-removal capacity. In addition, the interactions between metal cations and negatively charged functional group of bioflocculant such as carboxylic group, and the phosphate group found in bioflocculant (Fig. 6) were highly enhanced under constant optimum conditions33.
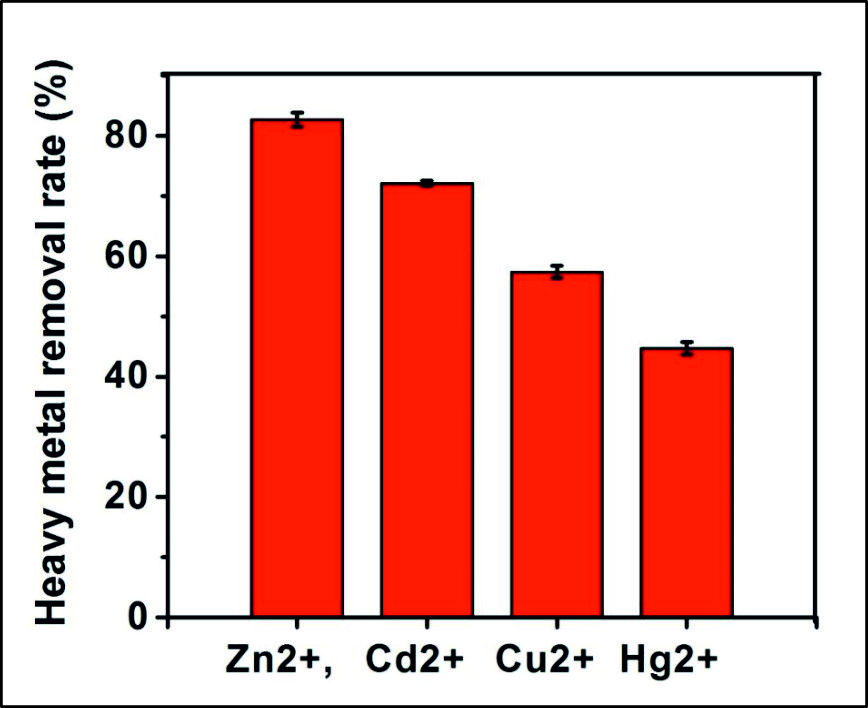 Fig. 6. Removal percentage of Cu2+, Zn2+, Hg2+, Cd2+ using bioflocculant. The given data represents the mean value of three independent experiments. Error bars indicate mean ± SD.
Fig. 6. Removal percentage of Cu2+, Zn2+, Hg2+, Cd2+ using bioflocculant. The given data represents the mean value of three independent experiments. Error bars indicate mean ± SD. Antibiofilm assay
Crystal violet staining (Biofilm assay) assay are used to quantify the measurement of biofilm. Bioflocculant were showed excellent biofilm inhibition against P. aeruginosa where maximum reduction (73%) were achieved at the amount of 80µg/ml dose. Similarly, maximum reduction (80%) were achieved at the amount of 70µg/ml dose for S. aureus34. Control (without bioflocculant) did not show any inhibiting percentage exhibited dense bacterial population (Fig. 7). High inhibition percentage and minimum dosage requirement were showed that, bioflocculant was acting as a best antibiofilm agent. They were further visualized by confocal microscope35,36. Previously reports suggested that, bioflocculant reduced cell –cell interaction by inhibiting the attachment of bacterial cells37 and also polysaccharides found in the bioflocculant might be responsible for biofilm inhibition38.
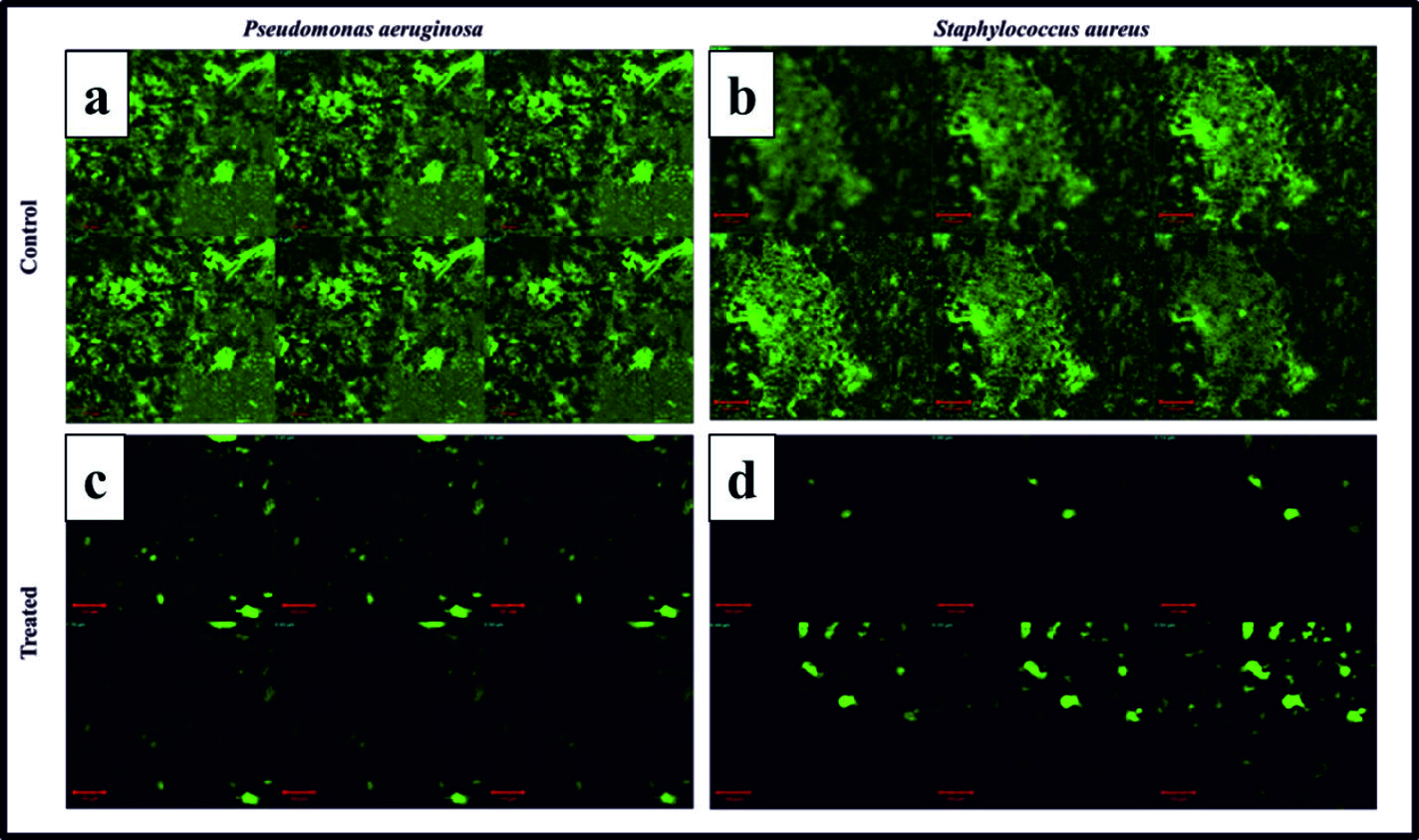 Fig. 7. (a) CLSM images of P. aeruginosa biofilm formation; (b) CLSM images of S. aureus biofilm formation; (c) CLSM images of P. aeruginosa biofilm formation treated with bioflocculant (80 µg/ml); (d) CLSM images of S. aureus biofilm formation treated with bioflocculant (70 µg/ml); Each experiment was replicated thrice.
Fig. 7. (a) CLSM images of P. aeruginosa biofilm formation; (b) CLSM images of S. aureus biofilm formation; (c) CLSM images of P. aeruginosa biofilm formation treated with bioflocculant (80 µg/ml); (d) CLSM images of S. aureus biofilm formation treated with bioflocculant (70 µg/ml); Each experiment was replicated thrice. Out of 20 bacterial strains, bacterial strain producing highest flocculating activity were chosen for further research. The main constituents of the bioflocculant were proteins, polysaccharides as the functional groups in the bioflocculant which was being confirmed by FTIR. They were mainly responsible for its flocculating activity and antibiofilm activity. Structural characteristics of the bioflocculant were confirmed by SEM analysis. Further, monomer unit of the bioflocculant were demonstrated by GCMS. The flocculating efficiency suggested that the bioflocculant was considered as an effective material for the treatment of heavy metal removing process. High inhibition percentage and minimum dosage requirement were showed that, bioflocculant was acting as a best antibiofilm agent.
Acknowledgements
I would like to acknowledge DST-PURSE (DST Sanction Order No- SR/FT/LS- 113/2009) for providing confocal microscopy images, and Centre of excellence in life science, Bharathidasan University, Trichy.
Funding
None.
Data Availability
All data generated during the study are included in the manuscript.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Abinaya M., Vaseeharan B., Divya M., Sharmili A., Govindarajan M., Alharbi N.S., Kadaikunnan S., Khaled J.M., Benelli G. Bacterial exopolysaccharide (EPS)-coated ZnO nanoparticles showed high antibiofilm activity and larvicidal toxicity against malaria and Zika virus vectors. J. Trace Elem. Med. Biol., 2018; 45:93-103.
Crossref - Aljuboori A.H.R., Idris A., Abdullah N., Mohamad R. Production and characterization of a bioflocculant produced by Aspergillus flavus. Bioresour. Technol., 2013; 127:489-93.
Crossref - Aljuboori A.H.R., Uemura Y., Osman N.B., Yusup S. Production of a bioflocculant from Aspergillus niger using palm oil mill effluent as carbon source. Bioresour. Technol., 2014; 171:66-70.
Crossref - Azizi S., Kamika I., Tekere M. Evaluation of heavy metal removal from wastewater in a modified packed bed biofilm reactor. PloS one., 2014; 11:e0155462.
Crossref - Bukhari N.A., Loh S.K., Nasrin A.B., Jahim J.M. Enzymatic hydrolysate of palm oil mill effluent as potential substrate for bioflocculant BM-8 production. Waste Biomass Valori., 2018; 1-13.
Crossref - Delattre C., Pierre G., Laroche C., Michaud P. Production, extraction and characterization of microalgal and cyanobacterial exopolysaccharides. Biotechnol. Adv., 2016; 34:1159-79.
Crossref - Drakou E.-M., Amorim C.L., Castro P.M., Panagiotou F., Vyrides I. Wastewater valorization by pure bacterial cultures to extracellular polymeric substances (EPS) with high emulsifying potential and flocculation activities. Waste Biomass Valori., 2018; 9:2557-64.
Crossref - Feoktistova M., Geserick P., Leverkus M. Crystal violet assay for determining viability of cultured cells. Cold Spring Harbor., 2016; pdb. prot087379.
Crossref - Fu F., Wang Q. Removal of heavy metal ions from wastewaters: a review. J. Environ. Manage., 2018; 92:407-18.
Crossref - Giri S.S., Harshiny M., Sen S.S., Sukumaran V., Park S.C. Production and characterization of a thermostable bioflocculant from Bacillus subtilis F9, isolated from wastewater sludge. Ecotox. Environ. Safe., 2018; 121:45-50.
Crossref - Gomaa E.Z. Production and characteristics of a heavy metals removing bioflocculant produced by Pseudomonas aeruginosa. Pol. J. Microbiol., 2018; 61:281-9.
- He N., Li Y., Chen J. Production of a novel polygalacturonic acid bioflocculant REA-11 by Corynebacterium glutamicum. Bioresour. Technol., 2018; 9:99-105.
Crossref - Kavita K., Singh V.K., Mishra A., Jha B. Characterisation and anti-biofilm activity of extracellular polymeric substances from Oceanobacillus iheyensis. Carbohyd. Polym., 2018; 101:29-35.
Crossref - Kim Y., Kim S.H. Released exopolysaccharide (r-EPS) produced from probiotic bacteria reduce biofilm formation of enterohemorrhagic Escherichia coli O157: H7. Biochem. Biophys. Res. Commun. 2018; 379:324-9.
Crossref - Kumar C.G., Joo H.-S., Choi J.-W., Koo Y.-M., Chang C.-S. Purification and characterization of an extracellular polysaccharide from haloalkalophilic Bacillus sp. I-450. Enzyme Microb. Tech., 2004; 34:673-81.
Crossref - Li C., Zhou L., Yang H., Lv R., Tian P., Li X., Zhang Y., Chen Z., Lin F. Self-assembled exopolysaccharide nanoparticles for bioremediation and green synthesis of noble metal nanoparticles. Appl. Mater. Interfaces., 2004; 9:22808-18.
Crossref - Lian B., Chen Y., Zhao J., Teng H.H., Zhu L., Yuan S. Microbial flocculation by Bacillus mucilaginosus: applications and mechanisms. Bioresour. Technol., 2004; 99:4825-31.
Crossref - Mu P., Plummer D.T. Introduction to practical biochemistry. 2004; Tata McGraw-Hill Education.
- Nguyen T.-K., Lam S.J., Ho K.K., Kumar N., Qiao G.G., Egan S., Boyer C., Wong E.H. Rational design of single-chain polymeric nanoparticles that kill planktonic and biofilm bacteria. ACS Infect. Dis., 2004; 3:237-48.
Crossref - Nwodo U.U., Green E., Mabinya L.V., Okaiyeto K., Rumbold K., Obi L.C., Okoh A.I. Bioflocculant production by a consortium of Streptomyces and Cellulomonas species and media optimization via surface response model. Colloids Surf. B., 2004; 116: 257-64.
Crossref - Oliver S., Wagh H., Liang Y., Yang S., Boyer C. Enhancing the antimicrobial and antibiofilm effectiveness of silver nanoparticles prepared by green synthesis.J. Mater. Chem. B., 2018; 6:4124-38.
Crossref - Pathak M., Devi A., Bhattacharyya K., Sarma H., Subudhi S., Lal B. Production of a non-cytotoxic bioflocculant by a bacterium utilizing a petroleum hydrocarbon source and its application in heavy metal removal. RSC Adv., 2018; 5:66037-46.
Crossref - Pathak M., Sarma H.K., Bhattacharyya K.G., Subudhi S., Bisht V., Lal B., Devi A. Characterization of a novel polymeric bioflocculant produced from bacterial utilization of n-hexadecane and its application in removal of heavy metals. Front. Microbiol. Journal., 2017; 8:170.
Crossref - Patowary K., Patowary R., Kalita M.C., Deka S. Development of an efficient bacterial consortium for the potential remediation of hydrocarbons from contaminated sites. Front. Microbiol. Journal., 2017; 7:1092.
Crossref - Rendueles O., Kaplan J.B., Ghigo J.M. Antibiofilm polysaccharides. Environ. Microbiol., 2017; 15:334-46.
Crossref - Salehizadeh H., Shojaosadati S. Removal of metal ions from aqueous solution by polysaccharide produced from Bacillus firmus. Water Res., 2003; 37:4231-5.
Crossref - Sathiyanarayanan G., Kiran G.S., Selvin J. Synthesis of silver nanoparticles by polysaccharide bioflocculant produced from marine Bacillus subtilis MSBN17. Colloids Surf. B., 2003; 102:13-20.
Crossref - Schmid J., Sieber V., Rehm B. Bacterial exopolysaccharides: biosynthesis pathways and engineering strategies. Front. Microbiol. Journal., 2003; 6:496.
Crossref - Shameer S. Biosorption of lead, copper and cadmium using the extracellular polysaccharides (EPS) of Bacillus sp., from solar salterns. 3 Biotech., 2003; 6:194.
Crossref - Sun P.-F., Lin H., Wang G., Lu L.-L., Zhao Y.-H. Preparation of a new-style composite containing a key bioflocculant produced by Pseudomonas aeruginosa ZJU1 and its flocculating effect on harmful algal blooms. J. Hazard. Mater., 2015; 284:215-21.
Crossref - Tang W., Song L., Li D., Qiao J., Zhao T., Zhao H. Production, characterization, and flocculation mechanism of cation independent, pH tolerant, and thermally stable bioflocculant from Enterobacter sp. ETH-2. PloS one., 2015; 9: e114591.
Crossref - Voica D.M., Bartha L., Banciu H.L., Oren A. Heavy metal resistance in halophilic Bacteria and Archaea. FEMS Microbiol. Lett., 2015; 363.
Crossref - Wang L., Ma F., Qu Y., Sun D., Li A., Guo J., Yu B. Characterization of a compound bioflocculant produced by mixed culture of Rhizobium radiobacter F2 and Bacillus sphaeicus F6. WorldJ. Microbiol. Biotechnol., 2011; 27:2559-65.
Crossref - Wang Z., Shen L., Zhuang X., Shi J., Wang Y., He N. & Chang Y.-I. Flocculation Characterization of a Bioflocculant from Bacillus licheniformis. Ind. Eng. Chem., 2015; 54:2894-901.
Crossref - Wu J.-Y., Ye H.-F. Characterization and flocculating properties of an extracellular biopolymer produced from a Bacillus subtilis DYU1 isolate. Process Biochem.Journal., 2007; 42:1114-23.
Crossref - Xiong Y., Wang Y., Yu Y., Li Q., Wang H., Chen R., He N. Production and characterization of a novel bioflocculant from Bacillus licheniformis. Appl. Environ. Microbiol., 2010; 76:2778-82.
Crossref - Yeo J., Lee Y.-H., Jeon S.-M., Jung U.J., Lee M.-K., Jung Y.-M., Choi M.-S. Supplementation of a novel microbial biopolymer, PGB1, from new Enterobacter sp. BL-2 delays the deterioration of type 2 diabetic mice. World J. Microbiol. Biotechnol., 2010; 17:1983-90.
- Zhang Z.-q., Bo L., Xia S.-q., Wang X.-j., Yang A.-m. Production and application of a novel bioflocculant by multiple-microorganism consortia using brewery wastewater as carbon source. J. Environ. Sci., 2007; 19:667-73.
Crossref
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.