ISSN: 0973-7510
E-ISSN: 2581-690X
J Pure Appl Microbiol, 2019, 13 (2): 933-948 | Article Number: 5525
https://dx.doi.org/10.22207/JPAM.13.2.30 | © The Author(s). 2019
Received:19/03/2019 | Accepted: 24/04/2019 | Published: 17/05/2019
Recently, probiotic lactic acid bacteria have been utilized as therapeutic supplements and food additives. Nowadays, the interest has been increased regarding the commercial utilization of probiotic LAB strains isolated from traditional fermented food products. Therefore, the present study was aimed to investigate the probiotic properties of lactic acid bacteria isolated from traditional food sources viz. dosa batter and sauerkraut. Total 7 lactic acid bacteria were obtained, 4 from dosa batter and 3 from sauerkraut, out of which one isolate from each food source were selected based on their broadest antagonistic spectrum. These strains were identified using 16S r RNA technique as Lactobacillus plantarum DB-2 (isolated from dosa batter) and Lactobacillus plantarum SK-3 (isolated from sauerkraut). The investigation of acid-bile tolerance, antibiotic sensitivity, auto-aggregation, co-aggregation, bacterial adhesion to hydrocarbons were confirmed. The results revealed normal growth of L. plantarum DB-2 and L. plantarum SK-3 in the presence of low pH, high bile salt concentration and ability to produce antimicrobial compounds viz. bacteriocin and H2O2. No gelatinase, lipase and hemolytic activity were observed. Natural susceptibility to the tested antibiotics was investigated. Thus, according to these results, L. plantarum DB-2 and SK-3 proved the good probiotic candidates as they survived during stress conditions posing to them and can be exploited for the preparation of nutraceutical products. This study also revealed the potential of using LAB and /or bacteriocin produced by them as food bio preservative to control food borne pathogenic bacteria in near future.
Lactobacillus plantarum, probiotics, acid and bile tolerance strains, bacteriocin, dosa batter, sauerkraut, auto-aggregation, antibacterial activity.
Probiotics are commonly referred as health-promoting bacteria and have been shown to improve the intestinal microbial balance1. Probiotics are defined as ‘Live microorganisms which have health benefits on host when taken in sufficient amount’2. Probiotic microorganisms are viable, non-pathogenic microorganisms which have the ability to restrict the growth of potential pathogenic microbes in the GIT, when reach in sufficient numbers and increase the multiplication of beneficial microbes and therefore, delivers the health benefit to the host3. LAB is designated as GRAS (generally regarded as safe) micro-organisms. Different geographical location across the world has diverse fermented foods which provide various lactic acid bacteria having potential probiotic properties4. The most commonly used probiotics belonging to the species of the genera Lactobacillus, Lactococcus, Pediococcus, Streptococcus, etc. and the genus Bifidobacterium5. Nowadays, lactic acid bacteria are used in probiotic preparations because it inhibit the growth of harmful bacteria, enhance good digestion, modify the balance of intestinal microflora, improve immune function and improve resistance to infection6.
A potential probiotic strain must fulfill certain selection criteria such as the ability to overcome gastric pH, the toxic effects of bile salts, phenol stability, antimicrobial production, antibiotic-resistance, and antagonistic activity and co-aggregation ability to decrease the ill-effects of pathogens. An appropriate alternative to antibiotic treatment is the use of probiotics with broad antagonistic potential. A good probiotic strain must possess the property of auto-aggregation before providing any health benefits. This property helps LAB to adhere to the intestinal epithelium and produce antimicrobial substances such as organic acid, hydrogen peroxide, and bacteriocins. Presently, traditional food products have been accepted for commercial purposes due to the probiotic significance. The probiotic strains survive the passage through the gastrointestinal tract and thus adaptable to intestinal conditions. This criterion helps the potential probiotic strain isolated from traditional food products to be used for industrial purposes other than those of animal origin7.
The main objectives of this study were to screen the probiotic potential of Lactobacilli spp. isolated from traditional fermented dosa batter and sauerkraut for its safety by antibiotic susceptibility test and assessment of probiotic qualities such as acid-bile tolerance, auto-aggregation, hydrophobicity, co-aggregation, and antagonistic potential. Tolerance studies were carried out to signify the importance of survivability of the strains in the stomach and intestine. Bacteriocin production was also investigated against several pathogenic bacteria including Escherichia coli IGMC, Staphylococcus aureus IGMC, Bacillus cereus CRI, P. aeruginosa IGMC, and S. pyogenes ATCC14289.
Collection of food samples
Both food items were prepared at home. The food samples were taken in a sterilized bag and stored at -4°C until use.
Strain isolation
1 g of food sample was added into 9 ml of normal saline. After homogenization, serial dilutions were prepared up to 10-9 with 0.85% (w/v) normal saline and 0.1 ml decimal of appropriate dilutions were plated onto de Man, Rogosa, Sharpe (MRS) agar medium (HiMedia)8. The agar plates were incubated at 35°C for 24 h under anaerobiosis. Morphologically different colonies were picked and re-streaked onto MRS agar plates up to purity. Pure strains (observed microscopically for homogeneity of cellular morphology) were preserved in glycerol stocks at -20°C.
Identification of strain
Phenotypic characterization
The morphological and cultural characteristics including gram staining9 and colonial appearance were examined.
Biochemical characterization
Indole test, MR-VP test, citrate utilization test, sugar fermentation test, gelatin hydrolysis test, lipase activity, hemolytic activity, BSH activity were employed to identify the isolated lactic acid bacteria10.
Genotypic characterization
Isolation of genomic DNA was done by following the method of11. Genomic DNA of the isolates was subjected to PCR for amplification of small 16S r RNA genes using universal primers 27F and 1492R having expected product size of 1500 bp. After amplification, PCR products were visualized using ethidium bromide (Thermo Fisher Scientific) on 1.5% agarose gel (Sigma-Aldrich)12. These have got sequenced by Bioserve Biotechnologies (India) Pvt. Ltd. to identify the isolates. BLAST software from the Genbank was used for sequence alignment. Program CLUSTAL_X was used for the multiple sequence alignment. MEGA-6.0 was used for a construction of phylogenetic tree by neighbour joining method.
Safety assessment of LAB
Safety is an important criteria for bacterial strains intended to use in the food industry.
Antibiotic sensitivity test
Study of antibiotic resistance pattern is important for selection and evaluation of safe probiotic strain. The antibiotic susceptibility of two L. plantarum strains was examined by disc diffusion technique13. The 24 h old culture was swabbed on MRS agar plates. Antibiotic impregnated discs (HiMedia) were placed onto these inoculated plates. These plates were incubated at 37°C for 24 h. Zone of inhibition was observed after 24 h14. Resistance was assessed against Ampicillin (10mg), Amoxicillin (10mg), co-Trimoxazole (30mg), Cefotaxime (30mg), Cefuroxime (30mg), Gentamycin (10mg) and Tetracycline (30mg).
Hemolytic activity
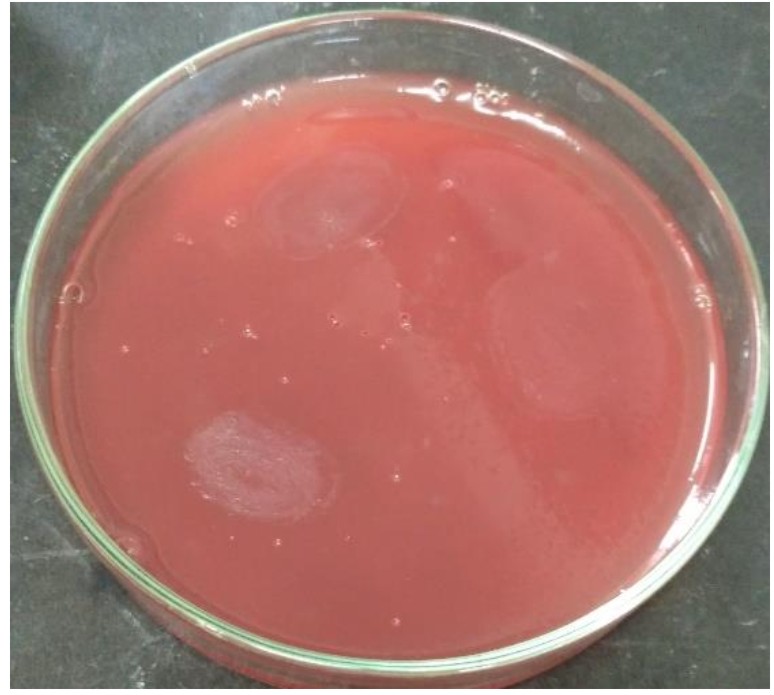
Hemolytic activity of both the strains were determined by spot inoculating overnight bacterial cultures on Blood Agar plates (HiMedia) followed by incubation of 24 h at 35°C15.
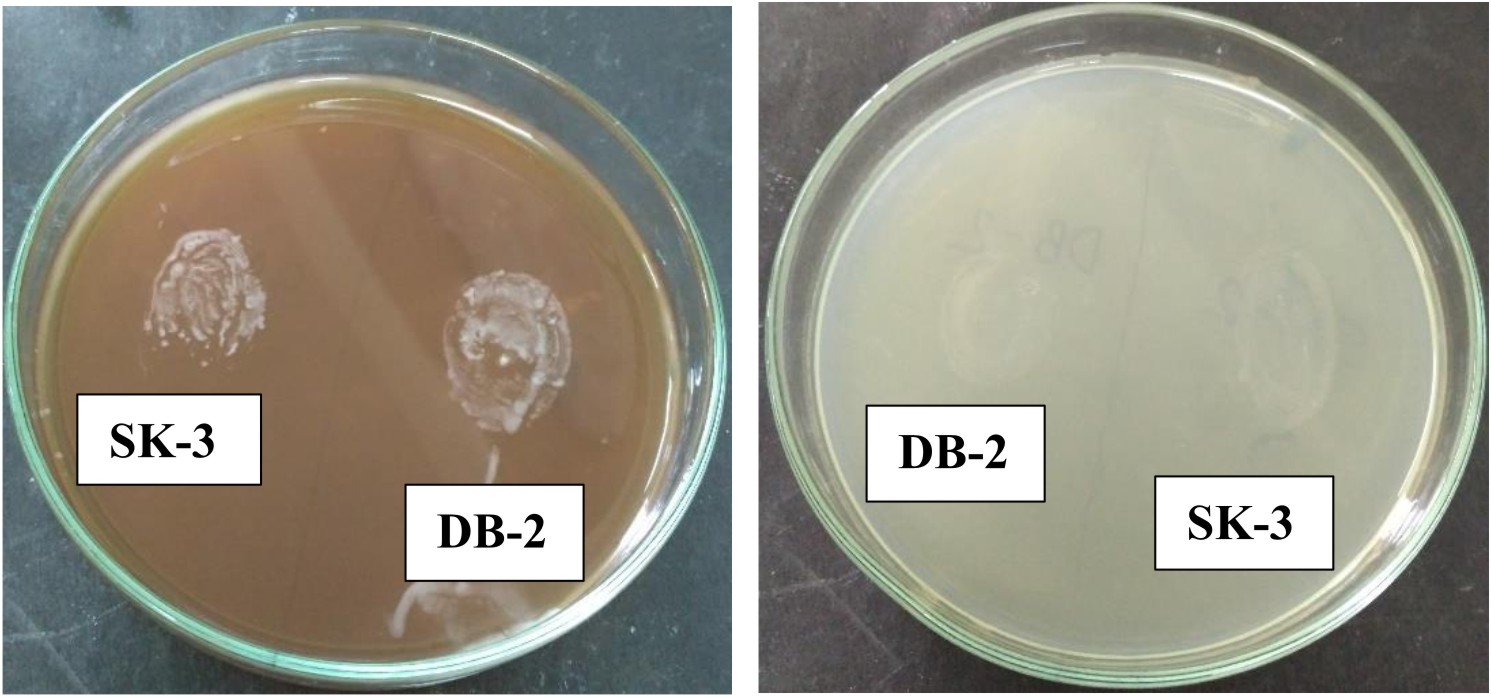
Gelatinase production
Gelatinase production was determined by streaking both the isolates on the MRS agar plates supplemented with 3% gelatin and the plates were incubated at 35°C for 24 h16.
Lipase production
Lipase enzyme production was evaluated by streaking the 24 h old culture of both the isolates on the MRS agar plates supplemented with 1% Tween 80 as a source of fatty acids. Plates were incubated at 35°C for 24 h17. The lipolytic activity was detected by the appearance of an opaque zone around the colonies18.
Assessment of probiotic attributes
Tolerance to low acid conditions
Selected isolates were grown in MRS broth at 37°C overnight. An equal amount of aliquot was taken and adjusted to pH 1.0, 2.0, 3.0, 4.0 and 5.0 with 5N HCl followed by incubation at 37°C for 3 h. Control was run alongside. 0.1 ml aliquot was taken every hour and enumerated by pour plate technique using 10-fold dilution using 0.1% peptone water. Simultaneously, the bacterial growth was monitored spectrophotometrically at OD600 at 0, 1, 2 and 3 h19.
Survivability % = X 100
Effect of bile salts on the growth rate of isolates
Selected isolates were grown in MRS broth at 37°C overnight. 0.3%, 1% and 2% (w/v) of bile salt (oxoid) was prepared and added to the 24 h old active culture of selected isolates and incubated at 37°C for 8 h. Control was run alongside. 0.1 ml aliquot was taken every hour and enumerated by pour plate technique using 10-fold dilution using 0.1% peptone water. Simultaneously the bacterial growth was monitored spectrophotometrically at OD600 at 0, 4 and 8 h20.
Survivability % = X 100
Survival in simulated in vitro digestion
The pH in the human stomach ranging from 1 (during fasting) to 4.5 (after a meal). Ingestion of food can take up to 4 h. Thus, the tolerance was assayed by determining the viable count in simulated gastric juice at different time intervals viz. 0,1 and 4 h21.
Aggregation property
Autoaggregation
Selected isolates were grown in MRS broth at 37°C overnight. After incubation, the broth was centrifuged at 10,000 rpm at 4°C for 10 min. The pellet obtained was washed twice with PBS buffer solution and re-suspend in the same solution, followed by incubation at 37°C for 5 h. An equal amount of aliquot was taken and absorbance was measured at OD600 at 0, 1, 2, 3, 4 and 5 h22.
Autoaggregation % = 1- (At / A0) x 100
Where At =Absorbance after incubation at 1, 2, 3, 4 and 5 h, A0 = Absorbance at 0 h
Co-aggregation
Mixtures were made for both the isolates with pathogenic bacteria viz. Bacillus cereus CRI, Bacillus subtilis MTCC 5981, Clostridium perfringens MTCC 1739, Escherichia coli IGMC, Pseudomonas aeruginosa IGMC, Salmonella typhimurium MTCC 3231 and Staphylococcus aureus IGMC at 1: 1 ratio. Probiotic bacterial cells and indicator microorganisms were kept as control and incubated at 35°C for 4 h. Absorbance (OD600) was observed for mixture and each of individual strain22. Co-aggregation percentage was calculated by Handley’s equation23.
Where ALAB =Absorbance of lactic acid bacterial suspension
APathogen = Absorbance of indicator microorganisms
AMixture = Absorbance of LAB suspension and indicator organisms
Adhesion property: Hydrophobicity
Selected isolates were grown in MRS broth at 37°C overnight. After incubation, the broth was centrifuged at 10,000 rpm at 4°C for 10 min. The pellet obtained was washed twice with PBS buffer solution and re-suspend in the same solution. 3 ml of cell suspension was added to 1 ml of each hydrocarbon (xylene, toluene, chloroform, n-hexadecane, n-octane and ethyl acetate). Absorbance (OD600) was taken at 0 h and after vortexing both phases for 2 min. Incubation was done for 2 h and absorbance was taken again24.
Hydrophobicity % = {(A0 – At) / A0} x 100
Where At = Absorbance at time t=2
A0 = Absorbance at time t=0
Antibacterial activity of bacteriocin producing L.plantarum
0.1 ml of indicator strains (Escherichia coli IGMC, Staphylococcus aureus IGMC, Bacillus cereus CRI, Pseudomonas aeruginosa IGMC, and Streptococcus pyogenes ATCC14289) were swabbed onto sterilized nutrient agar plates.
The selected probiotic isolates were grown overnight in TGY medium; a bacteriocin producing medium25. The isolates were centrifuged at 12,000 rpm at 4°C for 15 min. The culture supernatant was collected in sterilized test tubes and was neutralized to pH 6.5 with 1N NaOH and catalase was added at the rate of 0.1 mg/ml.
Inhibitory activity of bacteriocin was observed by well diffusion method13. The wells in the pre-swabbed nutrient plates were cut with sterile borer and 20µl of neutralized culture supernatant was placed into the wells.
AU/ml = Diameter of the zones of clearance (mm)/ x 1000
volume taken in well
Where AU = Arbitrary units / activity units of bacteriocin
H2O2 production
Both the isolates were screened for Hydrogen Peroxide by Quantitative method26. It was done by inoculating the bacterial isolate into MRS broth (25 ml) at 35°C for 24 h. After overnight incubation, 0.1 M sulphuric acid (20 ml) was added to the broth and titrated against 0.1 N KMnO4.
1 ml of KMnO4 = 1.070 mg of H2O2
BSH activity
Isolates were cultivated in MRS agar medium supplemented with 0.5% sodium salt of taurocholic acid (HiMedia) and incubated at 35°C for 24 h. The plates were observed for white precipitates27.
Statistical Analysis
All the experimental results were recorded as mean ± SD (Standard Deviation). For every observation, 3 determinations were used. Analysis of variance (ANOVA) was calculated by using one-way analysis. Duncan’s multiple range test was employed for calculating significant differences between mean. Results were statistically significant at P<0.05.
Isolation and identification of strain
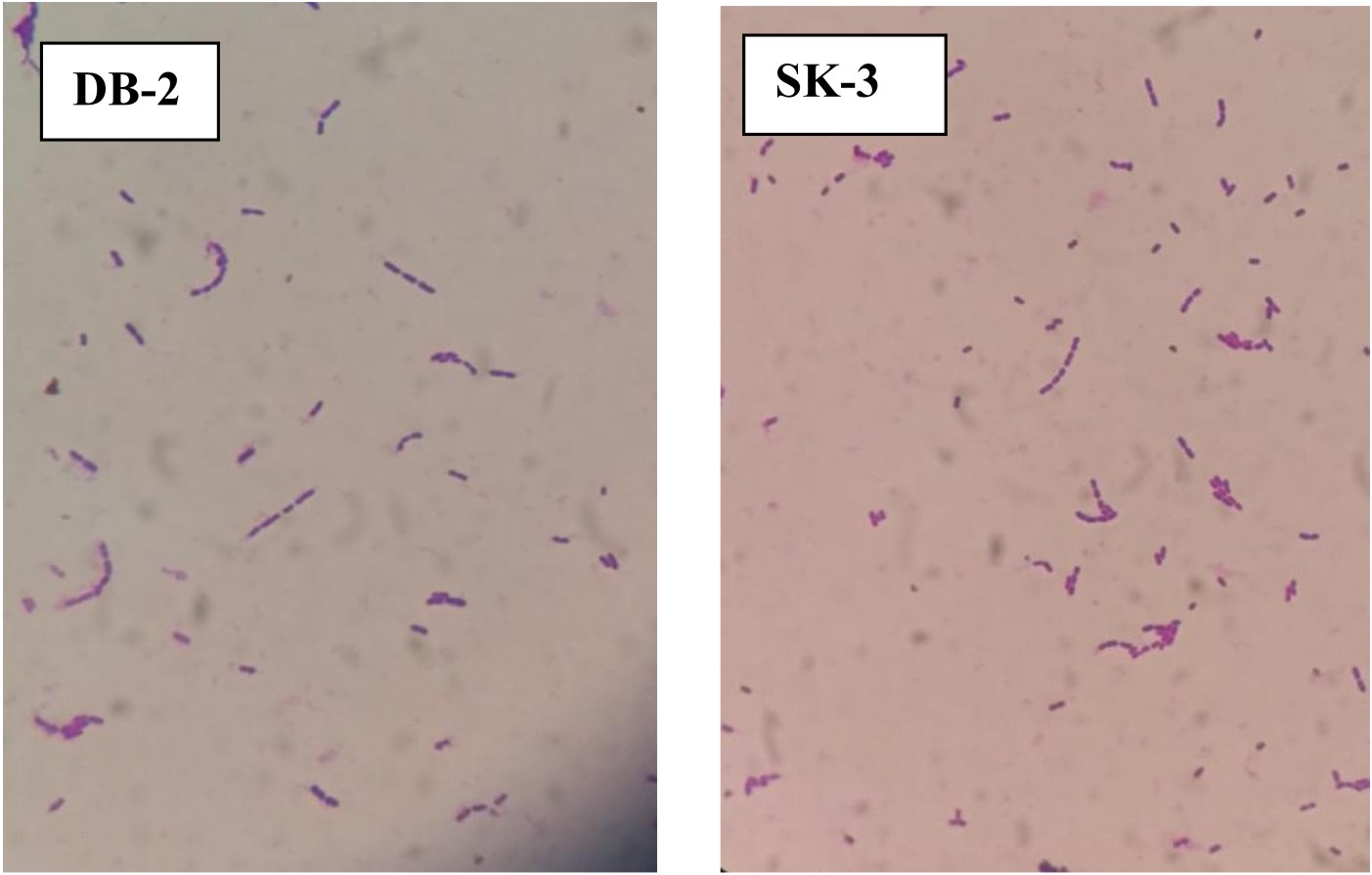
Total 7 lactic acid bacteria were isolated, 4 from dosa batter and 3 from sauerkraut. All 7 isolates were Gram-positive as examined by Gram’s staining method under an oil-immersion microscope (Fig. 1). 3 out of four dosa batter isolates were confirmed as rods while 1 was confirmed as coccus and all 3 isolates of sauerkraut were confirmed as rods as revealed by microscopic examination. All 7 isolates were non-sporulating. Out of 7 isolates, SK-3 and DB-2 gave clear halos around the indicator pathogenic organisms with widest antimicrobial spectrum and were selected for further study. P. pentosaceus VTCC-B-601 showed effective antimicrobial effect when tested against foodborne pathogens Staphylococcus aureus ATCC 25923, Salmonella typhimurium ATCC 19430, Pseudomonas aeruginosa ATCC 27853 and Micrococcus luteus ATCC 1024028.
Phenotypic characterization
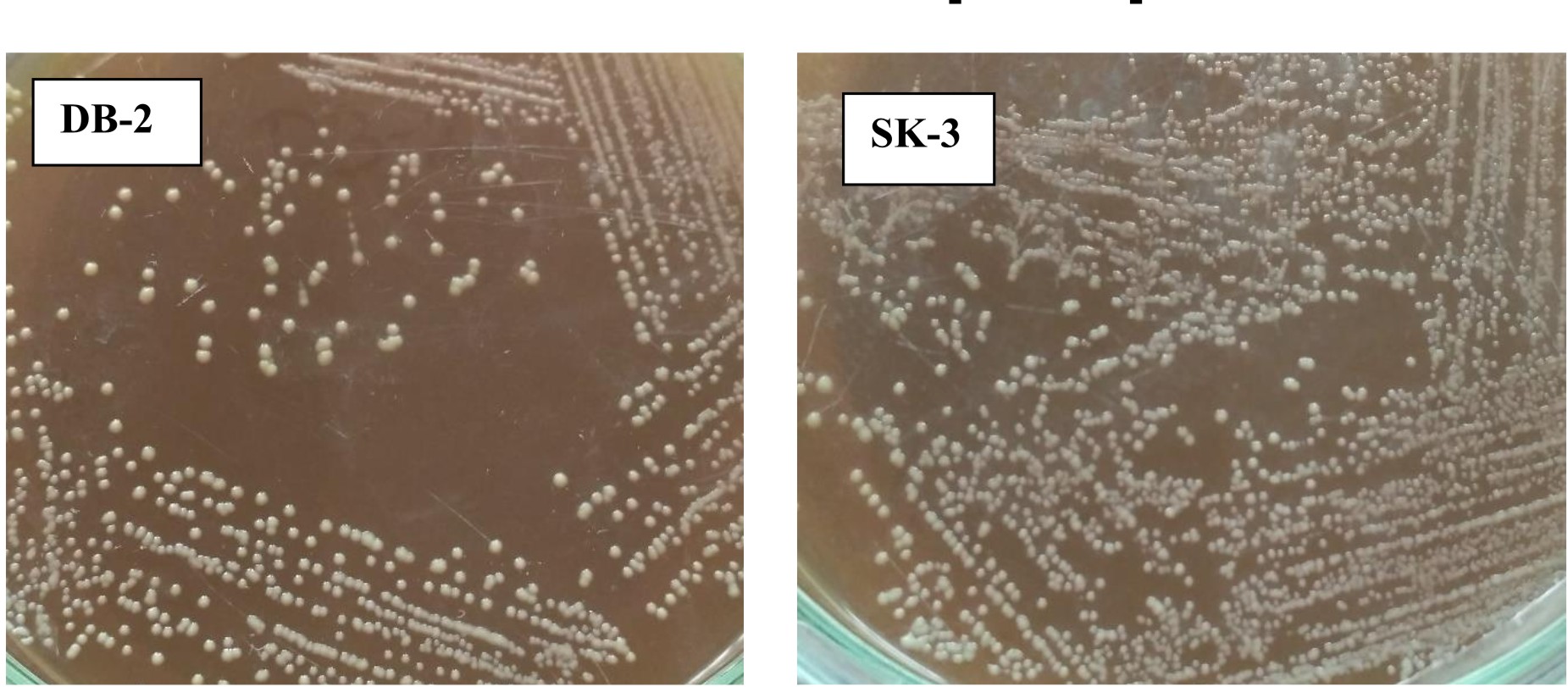
Both strains appear off-white, mucoid, raised with entire margins (Fig. 2).
Biochemical characterization
All the isolates were catalase negative, not able to utilize citrate, no casein hydrolysis, no urease production, no indole production, no lipase production and no hemolysis zone were observed (Table 1).
Table (1):
Biochemical characterization of L.plantarum DB-2 and SK-3.
| L.plantarum | Gram staining | Catalase reaction | Sugar fermentation test | Indole test | MR-VP test | Citrate utilization test | H2S production | Casein hydrolysis | Lipase production | Gelatin hydrolysis | Hemolytic activity | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SK3 | Rod | + | – | A+G– | – | +,- | – | – | – | – | – | – |
| DB2 | Rod | + | – | A+G– | – | +,- | – | – | – | – | – | – |
‘+’ indicates positive reaction; ‘-’ indicates negative reaction
Genotypic characterization
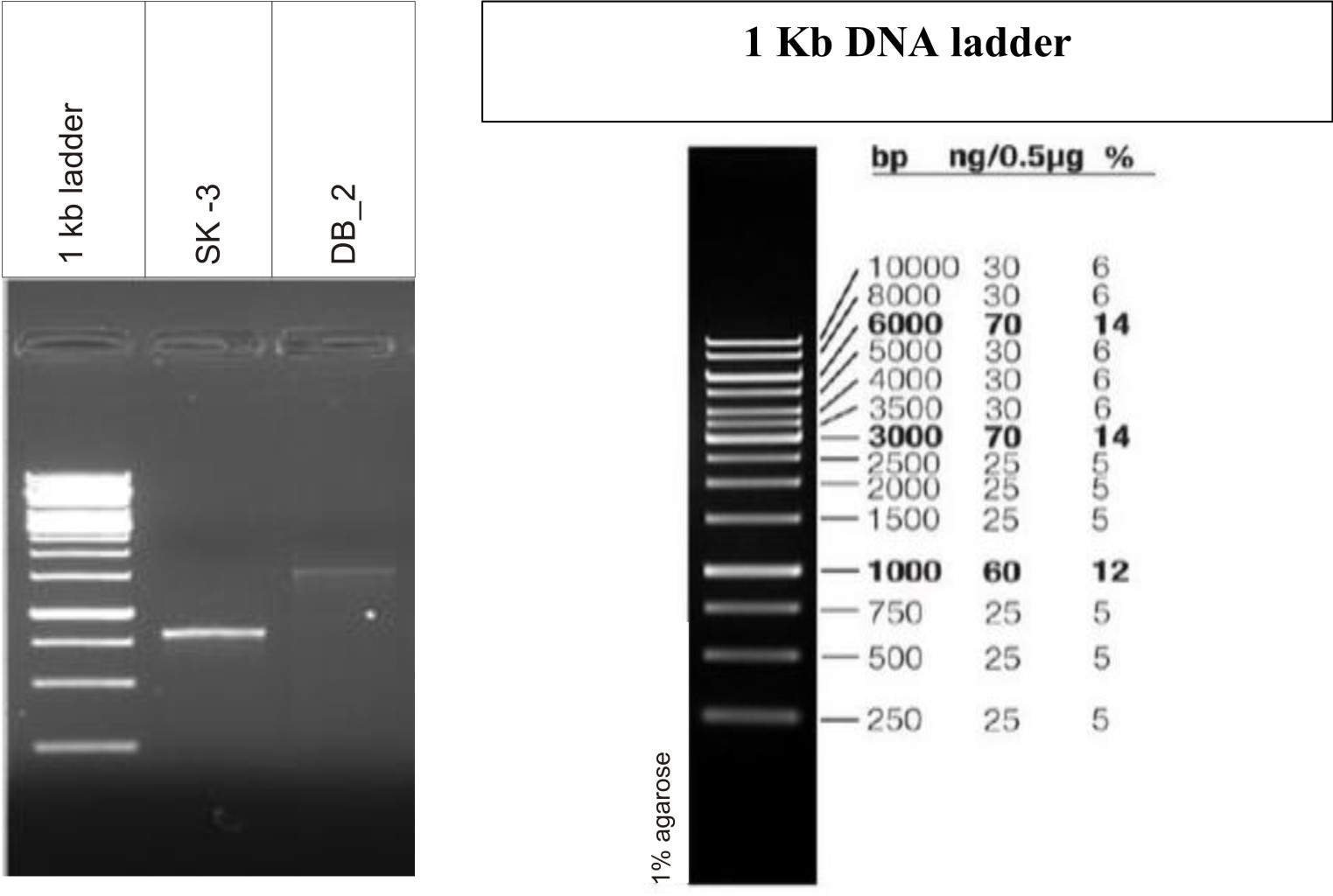
Gel electrophoresis is shown in Fig. 3. Analysis of the 16S rRNA sequences revealed that lactic acid bacteria isolated from dosa batter and sauerkraut showed 99% and 100% homology with Lactobacillus plantarum NCIMB 700965 respectively. The 16S rRNA gene sequence was submitted to Genbank and assigned accession number MK246167 and MK246169 for isolate SK3 and DB2 respectively. Neighbour-joining phylogenetic tree of L. plantarum DB-2 and SK-3 based on 16S rRNA gene sequences is shown in Fig. 4.
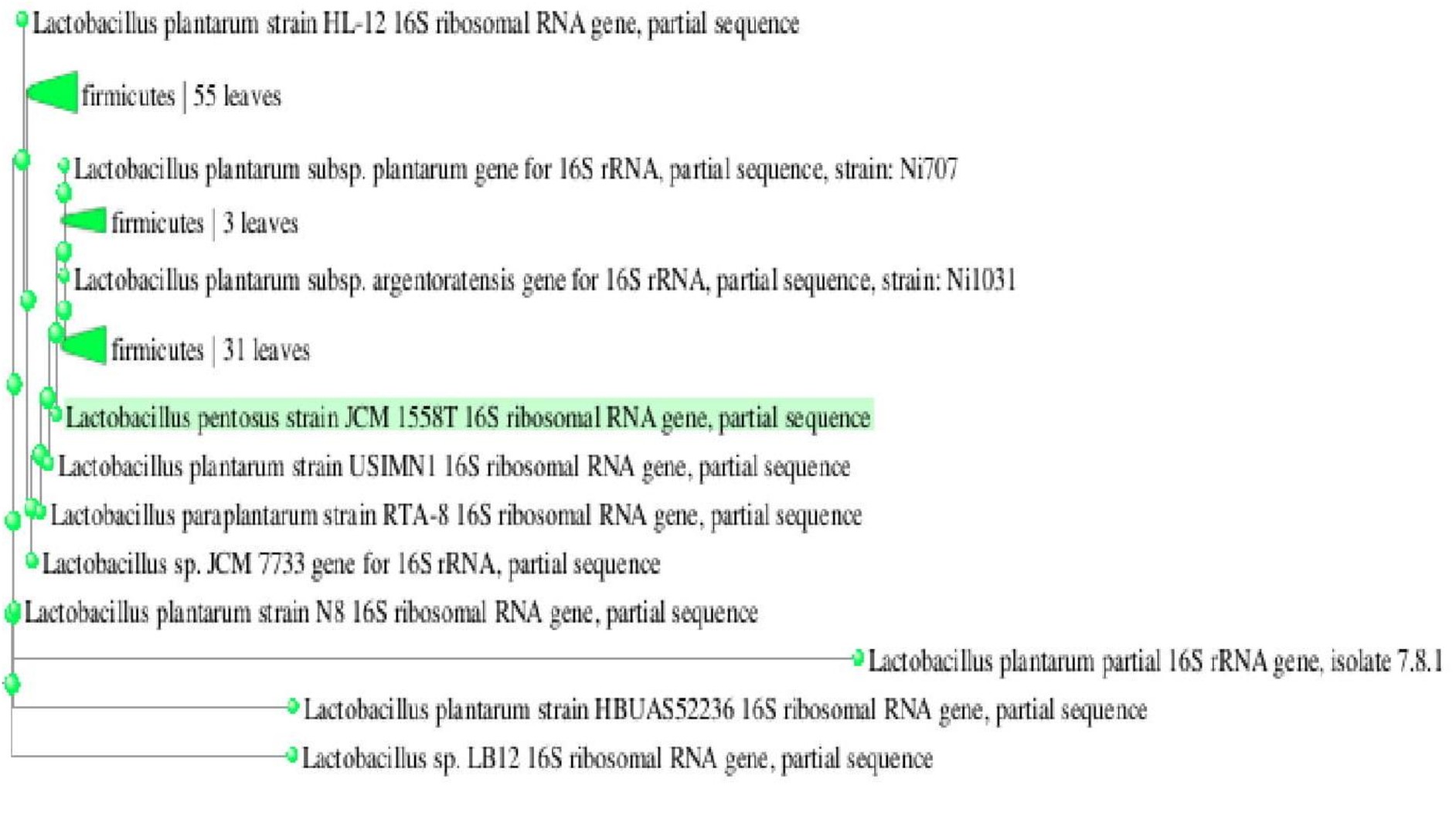

Fig. 4.A Phylogenetic dendogram of strain DB2 and related lactic acid bacterial species based on 16S r RNA gene sequence similarity
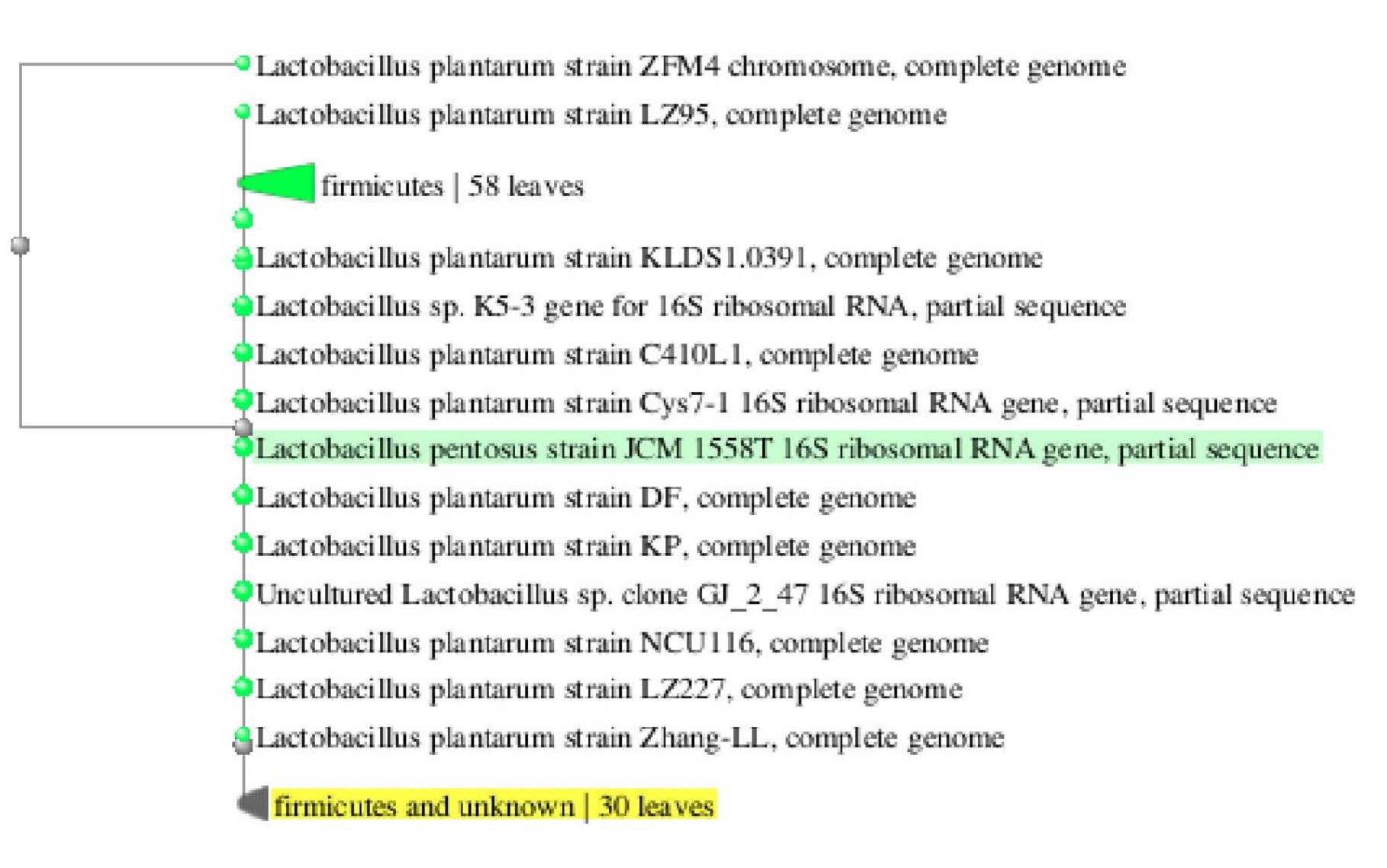

Fig. 4.B Phylogenetic dendogram of strain SK3 and related lactic acid bacterial species based on 16S r RNA gene sequence similarity
Safety assessment of selected isolates
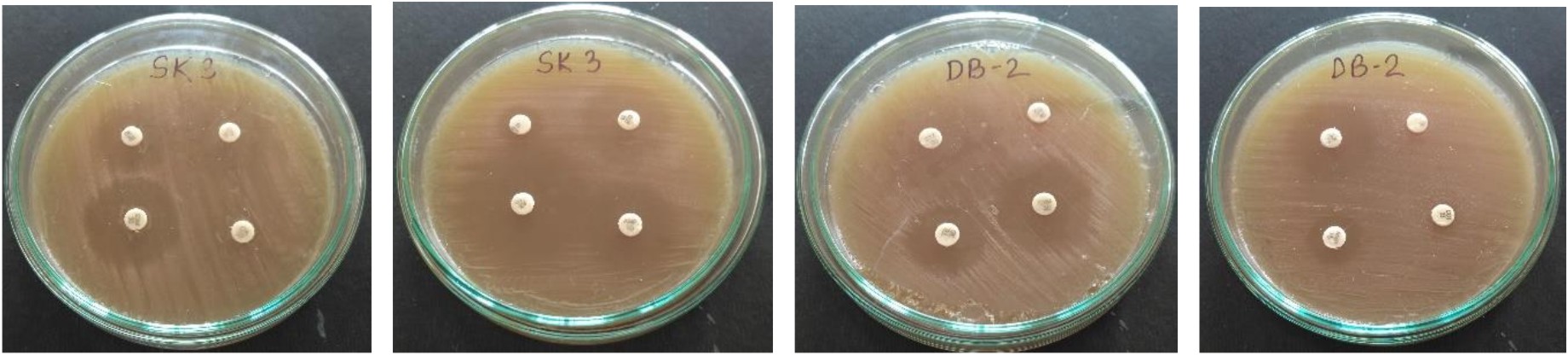
L. plantarum SK-3 and L. plantarum DB-2 exhibited 75% and 62.5% sensitivity towards the antibiotics used in this work. L. plantarum SK-3 showed resistance towards Cefotaxime CTX (30µg) and Cefoxitin CX (30µg) whereas L. plantarum DB-2 showed resistance towards Cefoxitin CX (30µg), Co-trimoxazole COT (30µg) and Gentamycin GEN (10µg) (Table 2 and Fig. 5). For some preventive and therapeutic purposes in controlling intestinal infections, the resistance of probiotic strains to some antibiotics could be used. Resistance of probiotic strains to antibiotics elucidating their potential in minimizing the negative effects of antibiotic therapy on the host bacterial ecosystem29. No hemolytic activity was shown by both the isolates as no clear zones were observed around the colonies on blood agar medium showing in Fig. 6. Both isolates showed a negative response for gelatinase as well as lipase production, showing in Fig. 7. Positive hemolytic activity (ability to breakdown red blood cells) halt the underlying epithelial layer whereas positive gelatinase activity (ability to hydrolyse gelatin) breakdown the protective lining of the GIT. Absence of hemolytic, gelatinase and lipase activity makes a strain non-virulent, indicating its selection for probiotic strain30.
Table (2):
Antibiotic sensitivity of L.plantarum SK3 and DB2.
Antibiotics used |
L.plantarum SK-3 |
L.plantarum DB-2 |
|---|---|---|
Ampicillin AMP (10 μg) |
*S |
S |
Amoxicillin AX (10 μg) |
S |
S |
Cefotaxime CTX (30 μg) |
*R |
S |
Cefoxitin CX (30 μg) |
R |
R |
Cefuroxime CXM (30 μg) |
S |
S |
Co-trimoxazole COT (30 μg) |
S |
R |
Gentamycin GEN (10 μg) |
S |
R |
Tetracycline TE (30 μg) |
S |
S |
% survivalability |
75 |
62.5 |
*Sensitive/Resistant
Probiotic properties
Tolerance to low acid conditions
A successful probiotic possesses the property of tolerating harsh acidic conditions of stomach and gut. Probiotic bacteria must pass through the stomach where the pH can be as low as 1.5 to 2, before reaching the intestinal tract31. In this study, we were able to obtain the isolates that were able to grow at minimum tested pH of 1.0 after 60 and 120 min of incubation, respectively (Table 3). Lactic acid is produced by lactic acid bacteria during fermentation metabolism thus revealed its ability to survive in the acidic environment of the stomach. However, the pH inside the gut is 2-4 in normal conditions and may reach up to pH 1 during fasting. For industrial use in food preparations, the organisms must survive the lowest possible pH32. Therefore, tolerance to low pH by L. plantarum DB-2 and L. plantarum SK-3 disclosed their survival under low acidic conditions. Moreover, different probiotic bacteria exhibit different resistance to acidic conditions and this feature is species and strain dependent33. Tolerance to acidic condition helps the Lactobacilli to reach the small intestine and thus contribute in balancing the intestinal microflora.
Table (3):
Acid tolerance of L.plantarum DB-2 & SK-3
| Incubation time (min) | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Absorbance* | Cell survival (log CFU/ml)** | %Cell survival*** | |||||||||||
| 0 | 60 | 120 | 180 | 0 | 60 | 120 | 180 | Mean | 60 | 120 | 180 | Mean | |
| DB-2 | 0.051 | 0.050 | 0.009 | 0.001 | 8.037 | 8.008 | 2.704 | 0.000 | 4.687 | 89.16 | 30.00 | 00.00 | 39.72 |
| SK-3 | 0.056 | 0.055 | 0.011 | 0.002 | 8.089 | 8.079 | 2.731 | 0.000 | 4.724 | 90.95 | 89.62 | 30.23 | 70.26 |
| DB-2 | 0.059 | 0.054 | 0.017 | 0.008 | 8.113 | 8.053 | 3.399 | 2.711 | 5.569 | 89.66 | 37.71 | 29.97 | 52.44 |
| SK-3 | 0.062 | 0.060 | 0.023 | 0.009 | 8.173 | 8.127 | 3.501 | 2.712 | 5.628 | 91.90 | 90.15 | 38.75 | 73.60 |
| DB-2 | 0.078 | 0.077 | 0.075 | 0.073 | 8.475 | 8.447 | 8.429 | 8.409 | 8.440 | 94.05 | 93.53 | 92.96 | 93.51 |
| SK-3 | 0.094 | 0.093 | 0.090 | 0.089 | 8.741 | 8.696 | 8.672 | 8.660 | 8.692 | 98.29 | 96.47 | 96.00 | 96.92 |
| DB-2 | 0.090 | 0.089 | 0.086 | 0.080 | 8.563 | 8.551 | 8.517 | 9.485 | 8.779 | 95.21 | 94.50 | 93.80 | 94.50 |
| SK-3 | 0.102 | 0.100 | 0.097 | 0.095 | 8.845 | 8.786 | 8.775 | 8.755 | 8.790 | 99.46 | 97.47 | 97.14 | 98.02 |
| DB-2 | 0.095 | 0.094 | 0.093 | 0.092 | 8.631 | 8.610 | 8.594 | 8.583 | 8.604 | 95.86 | 95.36 | 94.89 | 95.37 |
| SK-3 | 0.103 | 0.101 | 0.102 | 0.101 | 8.851 | 8.831 | 8.843 | 8.923 | 8.862 | 99.52 | 97.96 | 97.89 | 98.45 |
| DB-2 | 0.097 | 0.098 | 0.099 | 0.103 | 8.853 | 8.981 | 9.012 | 9.045 | 8.972 | 100 | 100 | 100 | 100 |
| SK-3 | 0.099 | 0.103 | 0.105 | 0.107 | 8.893 | 9.014 | 9.033 | 9.071 | 9.002 | 100 | 100 | 100 | 100 |
*Absorbance: Mean of results from three different experiments
**log CFU/ml: Mean of results from three different experiments
***Survivability: (log cfu/ml pH1, 2, 3, 4, 5/ log cfu/ml pH6.5)
Tolerance to bile salts
Bile salts are surface-active, amphipathic agents having potent antimicrobial activity. They act as detergent thus disrupts the cell membranes34. Bile acids are products of cholesterol metabolism and synthesized in the liver. It is secreted in the conjugated form (either with glycine or taurine) from the gall bladder to duodenum (500-700 ml/day). Bile acids play an important role in the digestive process (emulsification of fat). Bile concentration of intestine is 0.3% w/v. The small intestine has a low concentration of bile salts between 0.2-2 percent35. It is of great importance to evaluate the ability of probiotic strains to tolerate bile acids as these can act as antimicrobial molecules and thus influence the intestinal microflora. In this study, concentrations of 0.3%, 0.5%. 0.6%, 0.8%, 1% and 2% bile salts were used and their effect on growth rate of isolates was studied. Both the strains, L. plantarum DB-2 and L. plantarum SK-3 showed good survival after 8 h of incubation (Table 4). When the concentration of bile salt was increased up to 2%, the decrease was observed in the viable counts of isolates. It was considered that the biological cell membranes are composed of lipids and fatty acids and bile salts increase the permeability of these membrane.
Table (4):
Bile tolerance of L.plantarum DB-2 & SK-3
| Bile salt concentration (%) | Incubation time (h) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Absorbance* | Cell survival (log CFU/ml)** | %Cell survival*** | |||||||||
| 0 | 4 | 8 | 0 | 4 | 8 | Mean | 4 | 8 | Mean | ||
| 0.3 | DB-2 | 0.076 | 0.074 | 0.070 | 8.445 | 8.411 | 8.346 | 8.400 | 93.27 | 92.31 | 92.79 |
| SK-3 | 0.094 | 0.093 | 0.092 | 8.717 | 8.688 | 8.664 | 8.689 | 96.18 | 95.42 | 95.80 | |
| 0.5 | DB-2 | 0.071 | 0.068 | 0.063 | 8.397 | 8.334 | 8.294 | 8.341 | 92.42 | 91.73 | 92.08 |
| SK-3 | 0.087 | 0.079 | 0.077 | 8.522 | 8.495 | 8.451 | 8.489 | 94.04 | 93.08 | 93.56 | |
| 0.6 | DB-2 | 0.040 | 0.036 | 0.021 | 7.842 | 7.765 | 6.903 | 7.503 | 86.11 | 76.35 | 81.23 |
| SK-3 | 0.048 | 0.033 | 0.023 | 7.948 | 7.716 | 7.049 | 7.571 | 85.42 | 77.64 | 81.53 | |
| 0.8 | DB-2 | 0.031 | 0.027 | 0.019 | 7.673 | 7.536 | 6.803 | 7.337 | 83.57 | 75.24 | 79.41 |
| SK-3 | 0.030 | 0.028 | 0.022 | 7.635 | 7.584 | 6.944 | 7.387 | 83.95 | 76.48 | 80.22 | |
| 1 | DB-2 | 0.026 | 0.024 | 0.020 | 7.394 | 7.181 | 6.856 | 7.143 | 79.63 | 75.83 | 77.73 |
| SK-3 | 0.027 | 0.026 | 0.020 | 7.408 | 7.225 | 6.857 | 7.163 | 79.98 | 75.52 | 77.75 | |
| 2 | DB-2 | 0.021 | 0.020 | 0.012 | 6.944 | 6.805 | 6.380 | 6.709 | 75.46 | 70.56 | 73.01 |
| SK-3 | 0.024 | 0.020 | 0.015 | 7.017 | 6.856 | 6.505 | 6.792 | 75.89 | 71.64 | 73.77 | |
| Control | DB-2 | 0.096 | 0.100 | 0.103 | 8.791 | 9.017 | 9.041 | 8.949 | 100.0 | 100.0 | 100.00 |
| SK-3 | 0.098 | 0.102 | 0.104 | 8.853 | 9.033 | 9.079 | 8.988 | 100.0 | 100.0 | 100.00 | |
*Absorbance: Mean of results from three different experiments
**log CFU/ml: Mean of results from three different experiments
***Survivability: (log cfu/ml 0.3, 0.5, 0.6, 0.8, 1, 2% bile salt/ log cfu/ml 0% bile salt)
Tolerance to simulated gastric conditions
Probiotics must have the ability to survive passage through the stomach and small intestine as they are usually administered orally. Therefore, the survival of the probiotic strain in simulated conditions establishing in the stomach and the duodenum was examined by incubating selected cultures in MRS supplemented with pepsin (pH 2.0 and 3.0) and pancreatin (pH 8.0) for 4 h at 35°C. The survival of L. plantarum DB-2 and L. plantarum SK-3 at pH 2.0 and 3.0 containing pepsin (stomach conditions) and pH 8.0 containing pancreatin (intestinal conditions) was observed for different time intervals upto 4 h. L. plantarum DB-2 and L. plantarum SK-3 showed good survival at pH 2.0 (log CFU/ml 3.711 and 4.879) after 1 h of incubation whereas both the isolates did not show the survival after 4 h of incubation in pepsin at pH 2.0. At pH 3.0 of simulated gastrointestinal juice, both the strains survived after 4 h of incubation (log CFU/ml 5.798 and 3.711). Both isolates exhibited good survival at pH 8.0 of simulated gastrointestinal juice (log CFU/ml 7.437 and 7.920) (Table 5). Probiotic bacteria exert their health-promoting effects as metabolically viable active cells when they reach the colon by first surviving the transit through stomach followed by the intestine36. Ability to resist gastrointestinal conditions should be tested as survival through the GI tract is an important criterion for the selection of probiotic lactic acid bacteria. The simulated gastrointestinal and pancreatic digestions have been tested because of the independent action of strains to each of them and lead to a suitable global selection of probiotic bacteria37. Variation in acid resistance of probiotic lactic acid bacteria during the transit through gastrointestinal tract might be because of the changing pH values of the gastric juice (pH 2.0 to pH 3.5) depending on food components and feeding time38. In the present study, both lactic acid bacteria resisted the effects of pepsin and pancreatin during the transit in GI tract, therefore could be suggested as a potential probiotic candidate for further use in food preparations to improve the health of the gut.
Table (5):
Resistance of isolate SK-3 and DB-2 to simulated gastrointestinal juices
|
Gastro-intestinal juices |
Isolates | Incubation time (h) | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Absorbance* | Cell survival (log CFU/ml)** | %Cell survival*** | |||||||||
| 0 | 1 | 4 | 0 | 1 | 4 | Mean | 1 | 4 | Mean | ||
| pH 2 | DB-2 | 0.015 | 0.009 | 0.003 | 5.218 | 3.711 | 0.000 | 2.976 | 41.16 | 00.00 | 20.58 |
| SK-3 | 0.016 | 0.010 | 0.003 | 5.580 | 4.879 | 0.000 | 3.486 | 54.01 | 00.00 | 27.00 | |
| pH 3 | DB-2 | 0.025 | 0.021 | 0.018 | 7.099 | 6.901 | 5.798 | 6.599 | 76.55 | 64.17 | 70.36 |
| SK-3 | 0.022 | 0.015 | 0.009 | 6.917 | 5.218 | 3.711 | 5.282 | 57.76 | 40.87 | 49.32 | |
| pH 8 | DB-2 | 0.047 | 0.032 | 0.027 | 7.922 | 7.689 | 7.437 | 7.683 | 85.29 | 82.31 | 83.80 |
| SK-3 | 0.060 | 0.053 | 0.047 | 8.125 | 8.128 | 7.920 | 8.058 | 89.98 | 87.23 | 88.60 | |
| Control | DB-2 | 0.098 | 0.100 | 0.102 | 8.858 | 9.015 | 9.035 | 8.969 | 100.0 | 100.0 | 100.0 |
| SK-3 | 0.098 | 0.102 | 0.104 | 8.853 | 9.033 | 9.079 | 8.988 | 100.0 | 100.0 | 100.0 | |
*Absorbance: Mean of results from three different experiments
**log CFU/ml: Mean of results from three different experiments
***Survivability: (log cfu/ml pH2, 3, 8/ log cfu/ml pH6.5
Aggregation property
Autoaggregation
Interaction of the bacterial strain with itself (clumping of the cell) determines the auto-aggregation capability. Probiotic bacteria should adhere to the enterocytic cellular lines of oral cavity and GIT in order to exhibit their beneficial effects39. Bacterial aggregation depends on the amount of biofilm production which helps in adhesion of the cell40. The exact mechanism is not known of autoaggregation. Autoaggregation was investigated on the basis of sedimentation rate. The sedimentation rate was observed over 5 h of incubation. The ability of strains to auto-aggregate increased with increasing incubation time. L. plantarum DB-2 and L. plantarum SK-3 showed 97.6% and 98.2% aggregation after 5 h of incubation (Table 6). L. plantarum ST16Pa showed aggregation percentage as 37.05% after 60 min incubation41. The observed auto-aggregation could be due to cell surface component as they were not lost after washing and suspending of the cells in phosphate saline buffer42.
Table (6):
Estimation of auto-aggregation of selected Lactobacillus strains
| Isolates | OD600# | **Autoaggregation (%) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1h | 2h | 3h | 4h | 5h | Mean | 1h | 2h | 3h | 4h | 5h | Mean | |
| L.plantarum DB-2 | 0.023 | 0.020 | 0.017 | 0.016 | 0.012 | 0.017 | 95.4 | 96.0 | 96.6 | 96.8 | 97.6 | 96.48 |
| L.plantarum SK-3 | 0.02 | 0.017 | 0.015 | 0.013 | 0.009 | 0.014 | 96.0 | 96.6 | 97.0 | 97.4 | 98.2 | 97.04 |
*Autoaggregation in terms of sedimentation rate
#OD600 = Mean of the results from three different experiments
**Autoaggregation % = 1- (At=1, 2, 3, 4 and 5h/ A0h) x 100
Co-aggregation
As autoaggregation is the important property for adhesion to the mucoid lining of epithelial cells. In the same manner, co-aggregation posing a barrier in preventing the colonization of pathogenic microorganisms43,44. Co-aggregation is the process of joint aggregation of probiotic and pathogenic bacteria45. Probiotics have the ability to coaggregate with pathogenic microorganisms, inhibit their growth and finally kill them by secreting antimicrobial compounds which directly attack the cells of pathogenic bacteria46. L. plantarum SK-3 exhibited higher co-aggregation ability with pathogenic bacteria (71.25%) and the lower co-aggregation potential with pathogens was exhibited by L. plantarum DB-2 (59.34%). L. plantarum DB-2 and L. plantarum SK-3 showed highest co-aggregation potential against S.aureus (77.13% and 84.00%, respectively) (Table 7). The effective co-aggregation potential of probiotic bacteria against Gram-positive bacteria depend on the same cell wall morphology. Both have a thick peptidoglycan layer and bond get stronger by their hydrophobic nature47. The co-aggregation ability of lactic acid bacteria portrayed the high potential to kill undesirable microorganisms as they produce antimicrobial substances in close proximity to pathogenic bacteria. Lactic acid bacteria having co-aggregation potential have a significant role in human gut as they inhibit the growth of pathogenic strains by coaggregate with them in the gastrointestinal tracts48. During the process of co-aggregation, lactic acid bacteria secretes antimicrobial substance in higher concentration and thus maintaining the environment around the pathogens49 which behaves as an important host defence mechanism in the gut against the foodborne infections50. L. plantarum Lp-115 exhibited higher co-aggregation with E. sakazakii and good co-aggregation ability with S. aureus51.
Table (7):
Evaluation of Co-aggregation ability of L.plantarum DB-2 and SK-3 with test indicators
| Indicators | OD600# | **Co-aggregation (%) | ||||
|---|---|---|---|---|---|---|
| L. plantarum | Mean |
L. plantarum |
Mean | |||
| DB-2 | SK-3 | DB-2 | SK-3 | |||
| Bacillus cereus | 0.340 | 0.230 | 0.285 | 57.50 | 71.25 | 64.38 |
| Bacillus subtilis | 0.382 | 0.214 | 0.298 | 52.25 | 73.25 | 62.75 |
| Clostridium perfringens | 0.292 | 0.221 | 0.257 | 63.50 | 72.38 | 67.94 |
| Escherichia coli | 0.322 | 0.234 | 0.278 | 59.75 | 70.75 | 65.25 |
| Pseudomonas aeruginosa | 0.412 | 0.265 | 0.339 | 48.50 | 66.88 | 57.69 |
| Salmonella typhimurium | 0.346 | 0.318 | 0.332 | 56.75 | 60.25 | 58.50 |
| Staphylococcus aureus | 0.183 | 0.128 | 0.156 | 77.13 | 84.00 | 80.57 |
Adhesion property: Hydrophobicity
Cell surface hydrophobicity is the non-specific interaction between host and bacterial cells. Cell surface property of lactic acid bacteria is the key component for adhesion. Initially, the interaction is weak but gets stronger by adhesion process, mediated by cell surface proteins and lipoteichoic acids52,53. Bacterial adhesion to xylene, toluene, chloroform, ethyl acetate, n-Hexadecane and n-Octane was tested to study the Lewis acid-base characteristics of the bacterial cell surfaces. Out of the six solvents, chloroform is a monopolar acidic solvent, ethyl acetate is monopolar basic solvent, xylene, toluene, n-Hexadecane and n-Octane is the non-polar solvent. Determination of bacterial adhesion to xylene is a valid qualitative phenomenological approach54. The result of this study showed that the probiotic strains exhibited strong hydrophobicity towards non-polar solvents viz., chloroform, xylene, toluene, n-hexadecane and n-octane, exhibiting hydrophobic cell surface, which is a highly desirable probiotic trait. Electron donor and electron acceptor properties of bacteria were also regarded as a measure of hydrophobicity obtained with chloroform and ethyl acetate55. All the selected isolates proved to be a strong electron acceptor and weak electron donor as they exhibited strong affinity towards basic solvents such as ethyl acetate and low affinity towards acidic solvent such as chloroform. L. plantarum DB-2 and L. plantarum SK-3 exhibited highest adhesion towards toluene (85.6% and 87.6%, respectively and lowest adhesion towards chloroform (66.0% and 80.8%, respectively) (Table 8). Higher hydrophobicity is required for the colonization and adhesion of bacteria to the epithelial cells of the gut56. It was hypothesized that the presence of S-layer proteins on the cell wall of lactobacilli which have high isoelectric point showed strong affinity towards non-polar solvent. It has been suggested that cell surface properties play key role in autoaggregation as well as hydrophobicity. Adherence to epithelia helps in evaluating the surface hydrophobicity towards the non-polar and polar solvent. A good probiotic must possess high autoaggregation and strong hydrophobicity.
Table (8):
Adhesion of L. plantarum DB-2 and SK-3 to different hydrocarbons
Isolates |
OD600* |
% Hydrophobicity ** |
||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Xylene |
Toluene |
Chloroform |
n-Octane |
n-Hexadecane |
Ethyl Acetate |
Mean |
Xylene |
Toluene |
Chloroform |
n-Octane |
n-Hexadecane |
Ethyl Acetate |
Mean |
|
L.plantarum DB-2 |
0.091 |
0.072 |
0.170 |
0.080 |
0.096 |
0.083 |
0.098 |
81.8 |
85.6 |
66.0 |
84.0 |
80.8 |
83.4 |
80.2 |
L. plantarum SK-3 |
0.083 |
0.062 |
0.096 |
0.082 |
0.086 |
0.075 |
0.080 |
83.4 |
87.6 |
80.8 |
83.6 |
82.8 |
85.0 |
83.9 |
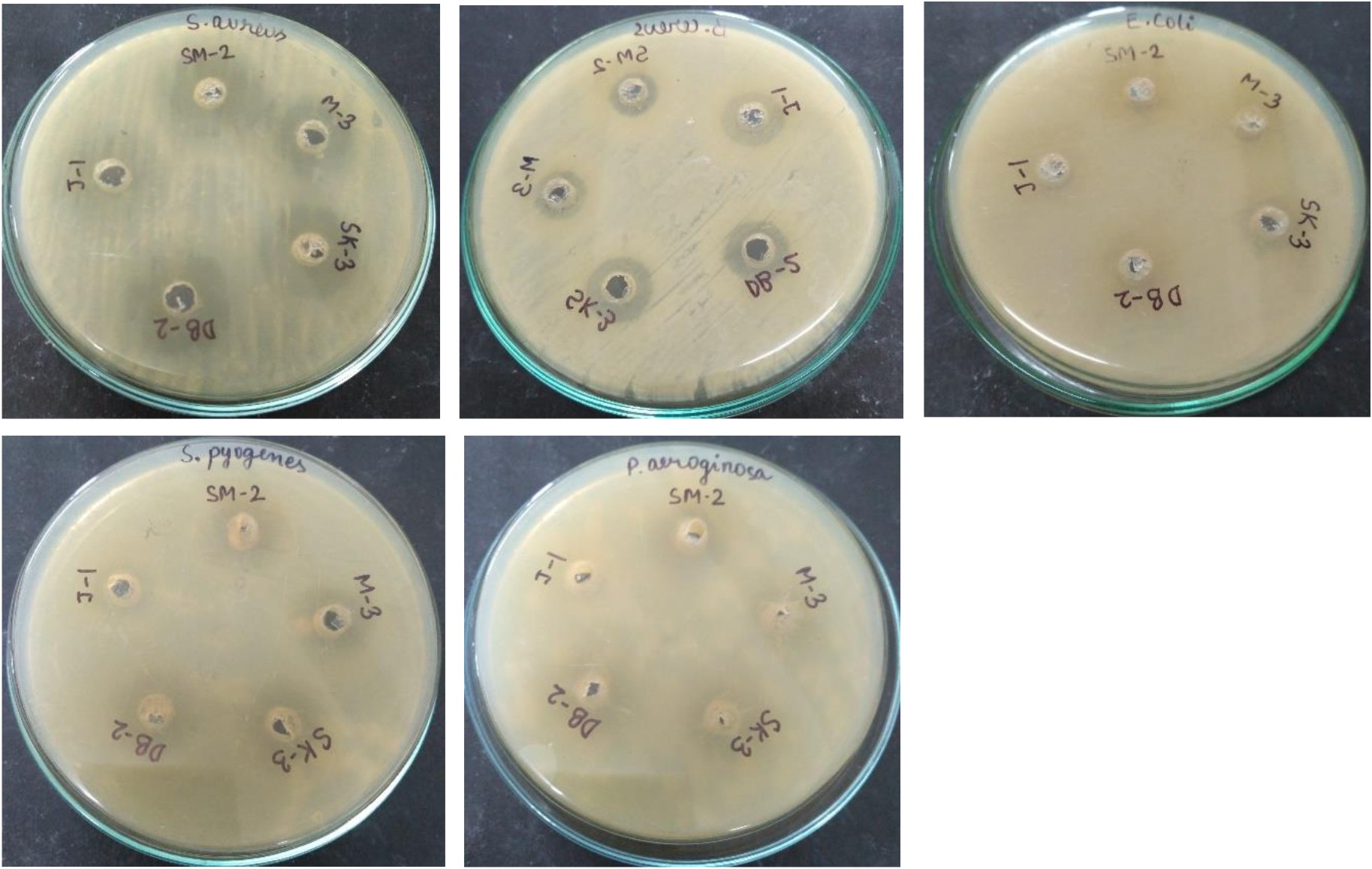
Antibacterial activity of bacteriocin producing L. plantarum
The antagonism exhibited by lactic acid bacteria occur due to the production of volatile short chain fatty acids such as lactic acid, acetic acid, propionic acid, hydrogen peroxide and specific inhibitory substances such as bacteriocin57. The bacteriocin activity of both isolates were measured using serial two-fold dilutions of acid neutralized and catalase treated cell-free culture supernatant against S. aureus, B. subtilis, E. coli, S. pyogenes and P. aeruginosa. The activity was lost after treatment with trypsin, this suggests that the activity was solely because of the bacteriocin production. Maximum bacteriocin production was observed during 18 h growth cycle of L. plantarum SK-3 and L. plantarum DB-2 with 745 AU/ml and 710 AU/ml against test indicator S. aureus followed by 685 AU/ml and 680 AU/ml against S. pyogenes, 455 AU/ml and 545 AU/ml against E.coli, 490 AU/ml and 440 AU/ml against P. aeruginosa, 410 AU/ml and 400 AU/ml against B. cereus, respectively (Fig. 8). Bacteriocins from L.gasseri inhibited various food borne pathogens58. The multi-drug resistance need to be solved with bacteriocin-producing lactic acid bacteria59. Both probiotic strains could be exploited for their implementation in controlling foodborne pathogenicity and safe bio-preservation of food products using natural antimicrobial agent i.e. bacteriocin as this study demonstrated the ability of lactic acid bacteria to inhibit the growth of foodborne pathogens through the production of bacteriocin.
H2O2 production
Probiotic isolates L. plantarum DB-2 and L. plantarum SK-3 were screened for production of hydrogen peroxide. L. plantarum DB-2 and L. plantarum SK-3 have been reported to produce 0.56 g L-1 and 0.52 g L-1 respectively. Normally, hydrogen peroxide is produced by vaginal lactobacilli isolates but may also be associated with intestinal lactobacilli60. In this study, L. plantarum DB-2 and L. plantarum SK-3 have been reported to produce H2O2 as an antimicrobial agent against food spoilage pathogens. Thus, this attribute can be used beneficially in improving vaginal health and in preventing urogenital infections. It can be hypothesized that the antagonism of these strains depends on lactic acid, H2O2 and bacteriocin thus exhibiting its potential and safe use as a bio-preservative in the food and fermentation industry.
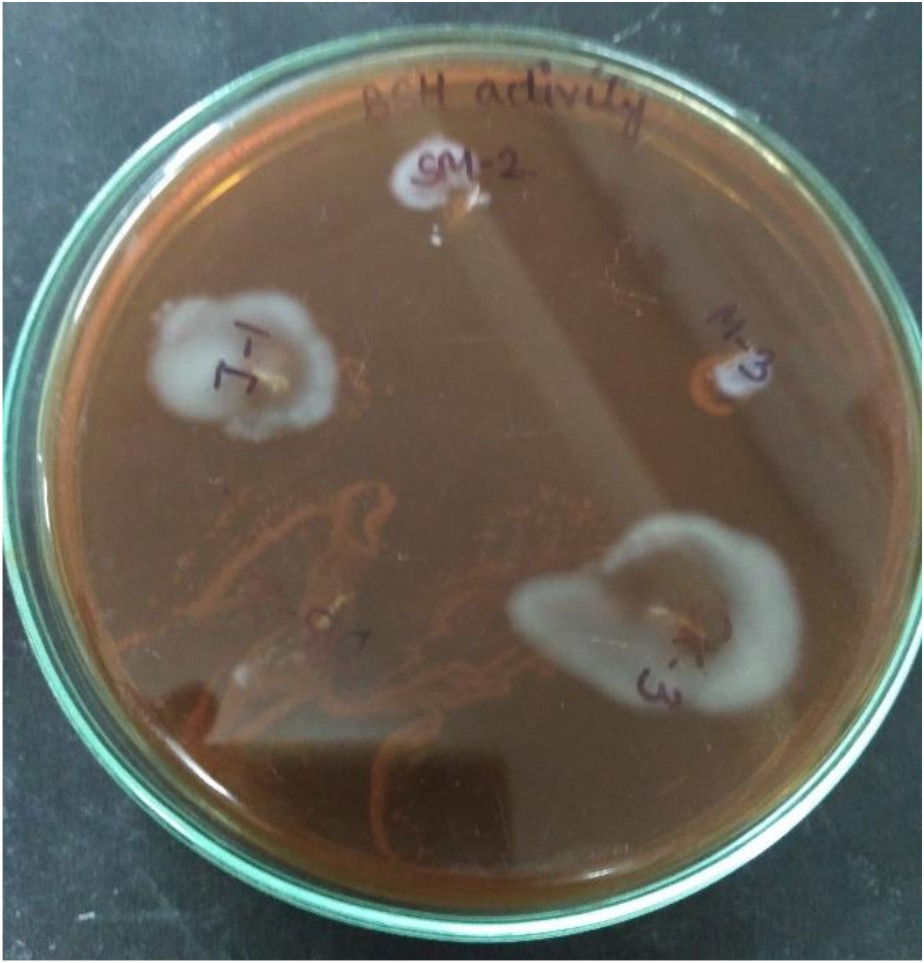
BSH activity
BSH catalyses the deconjugation of bile salts and deconjugated bile salts have lower solubility at low pH and thus precipitate as a result of the fermentative metabolism of lactic acid bacteria61. Removal of cholesterol from the medium (broth) in the presence of bile might be ascribed to co-precipitation with deconjugated bile salts62. In this study, isolate SK-3 deconjugated the bile acids (growth was observed on plates) while isolate DB-2 was unable to conjugate the bile acids (no growth was observed on plates) (Fig. 9).
All the probiotic attributes tested in this study revealed the safe status of both the isolates for further use in the food and fermentation industry. However, further evaluation of their beneficial effects on human beings will promote the application of both the strains in the pharmaceutical and cosmetic industry.
Compliance with Ethics Requirements
Acknowledgments
None
Conflict Of Interest
The authors declare that there is no conflict of interest.
Authors’ Contribution
All authors have made substantial, direct and intellectual contribution to the work and approved it for publication.
Funding
None
Data Availability
All datasets generated or analyzed during this study are included in the manuscript.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Mattila-Sandholm T., Alander M., Satokari R., Korpela R., Saxelin M., Vilpponen-Salmela T., von Wright A. Persistence of colonization of human colonic mucosa by a probiotic strain Lactobacillus rhamnosus GG, after oral consumption. Appl. Environ. Microbiol., 1999; 65(1): 351-4.
- Food and Agriculture Organization and World Health Organization (FAO/WHO), Health and nutritional properties of probiotics in food including powder milk with live lactic acid bacteria, FAO and WHO Joint Expert Committee Report, 2002.
- Hawaz E. Isolation and identification of probiotic lactic acid bacteria from curd and in vitro evaluation of its growth inhibition activities against pathogenic bacteria. African Journal of Microbiology Research, 2014; 8(13): 1419-1425.
- Guasch-Jane M., Andres-Lacueva C., Jauregui O., Lamuela-Raventos R. First evidence of white wine in ancient Egypt from Tutankhamun’s tomb. Journal of Archaeological Science, 2005; 33: 1075-1080.
- Saad N, Delattre C., Urdaci M., Schmitter J.M., Bressollier P. An overview of the last advances in probiotic and prebiotic field. LWT – Food Science and Technology, 2013; 50(1): 1-16.
- Ahn Y.T., Lim K.L., Ryu J.C., Kang D.K., Ham J.S., Jang Y.H., Kim H.U.. Characterization of Lactobacillus acidophilus isolated from piglets and chicken. Asian Journal of Animal Science, 2002; 15(12): 1790-1797.
- Lindstrom C., Holst O., Nilsson L., Oste R., Andersson K.E. Effects of Pediococcus parvulus 2.6 and its exopolysaccharide on plasma cholesterol levels and inflammatory markers in mice. AMB Express, 2012; 2: 66.
- de Man J., Rogosa M. and Sharpe M. A medium for the cultivation of lactobacilli. Journal of Applied Bacteriology, 1960; 3; 13-135.
- Gram H.C. ber die isolierte Fהrbung der Schizomyceten in Schnitt- und Trocke-nprהparaten. Fortschritte der Medizin, 1884; 2: 185–189.
- Aneja K.R. Experiments in Microbiology, Plant pathology and Biotechnology. Biochemical activities of microorganisms, 2003, pp. 245-275. 4th Ed. Newage International Publishers, New Delhi.
- Hashem S., Sabit H.H., Amin M., Tawakkot W. and Shamseldin A.F. Molecular characterization of Egyptian isolates of Lactobacillus and Bifidobacterium. Journal of American Science, 2010; 6(11): 959-964.
- Bollag D.M. and Edelstein S.J. Protein concentration determination. In: Bollag DM and Edelstein SJ, eds. Protein Methods, Wiley-Liss, New York, 1991.
- Kimura H., Sashihara T., Matsusaki H., Sonomoto K. and Ishizaki A. Novel bacteriocin of Pediococcus sp. ISK-1 isolated from well – aged bed of fermented rice bran. Annals of New York Academy of Science, 1998; 864: 345-348.
- Thirabunyanon M., Boonprasom P. and Niamsup P. Probiotic potential of lactic acid bacteria isolated from fermented dairy milks on antiproliferation of colon cancer cells. Biotechnology Letters, 2009; 31: 571–576.
- Linaje R., Coloma M.D., Perez-Martinez, G., Zuniga M. Characterization of faecal enterococci from rabbits for the selection of probiotic strains. Journal of Applied Microbiology, 2004; 96: 761-771.
- Harrigan W.F., McCance, M.E. Laboratory Methods in Food and Dairy Microbiology. Academic Press, London, 1990.
- Barrow G.I., Feltham R.K.A., Cowan and Steel’s manual for the identification of medical bacteria, 3rd ed, Cambridge: Cambridge University Press, 1993.
- Tanasupawat S., Phoottosavako M. and Keeratipibul. Characterization and lipolytic activity of lactic acid bacteria isolated from Thai fermented meat. Journal of Applied Pharmaceutical Sciences, 2015; 5(03): 006-012.
- Liong M.T. and Shah N.P. Acid and bile tolerance and cholesterol removal ability of Lactobacilli strains. Journal of Dairy Science, 2005; 88: 55-56.
- Gilliland S.E. and Walker D.K.. Factors to consider when selecting a culture of L.acidophilus as a dietary adjunct to produce a hypercholesterolemic effect in humans. Journal of Dairy Science, 1990; 73: 905-909.
- Charteris W.P., Kelly P.M., Morelli L., Collins J.K. Development and application of an in vitro methodology to determine the transit tolerance of potentially probiotic Lactobacillus and Bifidobacterium species in the upper human gastrointestinal tract. J. Appl. Microbiol., 1998; 84: 759-768.
- Del Re B., Sgorbati B, Miglioli M., Palenzona D. Adhesion, autoaggregation and hydrophobicity of 13 strains of Bifidobacterium longum . Lett. Appl. Microbiol., 2000; 31: 438-442.
- Handley P.S., Harty D.W., Wyatt J E, Brown C.R., Doran J.P. A comparison of the adhesion, co-aggregation and cell-surface hydrophobicity properties of fibrillar and fimbriate strains of Streptococcus salivarius. J. Gen. Microbiol., 1987; 133: 3207–3217.
- Rosenberg M., Gutnick D. and Rosenberg E. Adherence of bacteria to hydrocarbons: a simple method for measuring cell surface hydrophobicity. FEMS Microbiology Letters, 1980; 9: 29-33.
- Papagianni M. and Anastasiadou S. Pediocins: the bacteriocins of Pediococci: sources, production, properties and applications. Microbial Cell Factories, 2009; 8: 3.
- AOAC. Official methods of analysis of association of official analytical chemists, 16th edn. Association of Official Analytical Chemists.Arlington, Virginia, USA, 1995.
- Begley M., Hill C., Gahan C.G. Bile salt hydrolase activity in probiotics. Appl Environ Microbiol, 2006; 72: 1729-1738.
- Nghe D. and Nguyen T. Characterization of antimicrobial activities of Pediococcus pentosaceus Vtcc-B-601. Journal of Applied Pharmaceutical Sciences, 2014; 4(05): 061-064.
- El-Naggar M.Y.M. Comparative study of probiotic cultures to control the growth of Escherichia coli O157:H7 and Salmonella typhimurium. Asian Network for Scientific Information Biotechnol, 2004; 3(2): 173-180.
- Marroki A., Bousmaha-Marroki L. Lactobacilli isolated from Algerian goat’s milk as adjunct culture in dairy products. Brazilian Archives of Biology and Technology, 2014; 57: 410-420.
- Dunne C., O’Mahony L., Murphy L., Thonton G., Morrissey D., O’Halloran S., Feeney M., Flynn S, Fitzgerald G., Daly C., Kiely B., O’Sullivan G.C., Shanahan F., Collins J.K. In vitro selection criteria for probiotic bacteria of human origin: correlation with in vivo findings. Am. J. Clin. Nutr. 2001; 73: 386S-392S.
- Subhashini. Bioprospecting of Lactic acid bacteria for potentiality as probiotics. International Journal of Microbiological Research, 2014; 5(2): 90-97.
- Fontana L., Bermudez-Brito M., Plaza-Diaz J., Muסoz-Quezada S., Gil A. Sources, isolation, characterisation and evaluation of probiotics. Br J Nutr 109 Suppl, 2013; 2: S35-50.
- Lebeer S., Vanderleyden J., De Keersmaecker S.C. Genes and molecules of lactobacilli supporting probiotic action. Microbiology and Molecular Biology Reviews, 2008; 72: 728-764.
- Gunn J.S. Mechanisms of bacterial resistance and response to bile. Microbes Infection, 2000; 2: 907-913.
- Malek R.A., Hamdan S., El Enshasy H., Othman N.Z., Zainol N.A., Sarmidi M.R. and Aziz R. Production of Lactobacillus salivarius, a new probiotic strain isolated from human breast milk, in semi-industrial scale and studies on its functional characterization. 2010, pp. 1196-1204. In: Mיdez-Vilas A, ed. Current Research, Technology and Education Topics in Applied Microbiology and Microbial Biotechnology, Vol. 2, Spain: Formatex Research Centre, Badajoz.
- Bautista-Gallega J., Arroyo-Lopez F.N, Rantsiou K., Jimenez-Dia R., Garrido-Fernandez A. and Cocolin L. Screening of lactic acid bacteria isolated from fermented table olives with probiotic potential. Food Research International, 2013; 50: 135-142.
- Petsuriyawong B. and Khunajakr N. Screening of probiotic lactic acid bacteria from piglet feces. Kasetsart Journal, 2011; 45: 245-253.
- Nikolic M., Jovcic B., Kojic M. and Topisirovic L. Surface properties of Lactobacillus and Leuconostoc isolates from homemade cheeses showing auto-aggregation ability. European Food Research and Technology, 2010; 231: 925-931.
- Rickard A.H., Gilbert P, High N.J., Kolenbrander P.E. and Handley P.S. Bacterial co-aggregation: an integral process in the developments of multispecies biofilms. Trends Microbial, 2003; 11: 94-100.
- Vaz-Velho M. and Todorov S.D. Potential probiotic evaluation of bacteriocin producing Lactobacillus plantarum ST16PA isolated from Papaya (Carica Papaya). 12th ASEAN Food Conference, Bangkok, Thailand, 2011.
- Kos B., Suskovic J., Vukovic S., Simpraga M., Frece J., Matosic S. Adhesion and aggregation ability of probiotic strain Lactobacillus acidophilus M92. Journal of Applied Microbiology, 2003; 94: 981-987.
- Reid G., McGroarty J.A., Domingue P.A., chow A.W., Bruce A.W., Eisen A. and Costerton J.W.1. Co-aggregation of urogenital bacteria in vitro and in vivo. Current Microbiology, 1990; 20: 47-52.
- Boris S., Suarez J.E., Vasquez F. and Barbes C. Adherence of human vaginal Lactobacilli to vaginal epithelial cells and its interaction with uropathogens. Infection Immunity, 1998; 66:1985-1989.
- Surono I.S. Probiotic, fermented milk and healthy, 2004, Pp. 20- 40. YAPPMI, Jakarta, Indonesia.
- Bao Y., Zhang Y., Zhang Y., Lui and Wang S. Screening of potential probiotic properties of Lactobacillus fermentum isolated from traditional dairy products. Food Control, 2010; 21: 695-701.
- Arief I.I., Jenie B.S.L., Astawan M., Fujiyama K. and Witarto A.B. Identification and probiotic characteristics of lactic acid bacteria isolated from Indonesian local beef. Asian Journal of Animal Sciences, 2015; 9(1): 25-36.
- Botes M., Van Reenen C.A. and Dicks L.M.T. Evaluation of Enterococcus mundtii ST4SA and Lactobacillus plantarum 423 as probiotics by using a gastrointestinal model with infant milk formulations as substrate. International Journal of Food Microbiology, 2008; 128: 362–370.
- Li Q., Liu X., Dong M., Zhou J. and Wang Y. Aggregation and adhesion abilities of 18 lactic acid bacteria strains isolated from traditional fermented food. International Journal of Agricultural Policy and Research, 2015; 3(2): 84-92.
- Reid G., McGroarty J.A., Angotti R. and Cook R.L. Lactobacillus inhibitor production against Escherichia coli and co-aggregation ability with uropathogens. Canadian Journal of Microbiology, 1988; 34: 344–351.
- Collado M.C., Meriluoto J. and Salminen S. Adhesion and aggregation properties of probiotic and pathogen strains. European Food Research and Technology, 2008; 222: 1065- 1073.
- Rojas M., Ascencio F. and Conway P.L. Purification and characterization of a surface protein from Lactobacillus fermentum104R that binds to porcine small intestinal mucus and gastric mucin. Applied and Environmental Microbiology, 2002; 68: 2330-2336.
- Ross S. and Jonsson H. A high-molecular mass cell surface protein from Lactobacillus reuteri1063 adheres to mucus components. Microbiology, 2002; 148: 433-442.
- Kiely L.J. and Olson N.F. The physicochemical surface characteristics of Lactobacillus casei. Food Microbiology, 2000; 17: 277-291.
- Bellon-Fontaine M.N., Rault J. and van Oss C.J. Microbial adhesion to solvents: a novel method to determine the electron donor/electron acceptor or lewis acid-base properties of microbial cells. Colloids and Surfaces B: Biointerfaces, 1996; 7: 47–53.
- He T., Priebe M.G., Zhong Y., Haung C., Harmsen H.J., Raangs G.C., Antonie J.M., Welling G.W. and Vonk R.J. Effects of yogurt and bifidobacteria supplementation on the colonic microbiota in lactose intolerant subjects. Journal of Applied Microbiology, 2008; 104(2): 595-604.
- 57. Havenaar R. and Huis in’t Veld J.H.J. Probiotics: a general view. In: Wood B.J.B., ed. The lactic acid bacteria in health and disease, London: Elsevier Applied Science, 1992; 1: 151- 71.
- Itoh T., Fujimoto Y., Kawai Y., Toba T., Saito T. Inhibition of food borne pathogenic bacteria by bacteriocins from Lactobacillus gasseri. Lett Appl. Microbiol, 1995; 21: 137-141.
- Barefoot S.F., Klaenhammer T.R. Detection and activity of lactacin B, a bacteriocin produced by Lactobacillus acidophilus. Appl Environ Microbiol, 1983; 45: 1808-1815.
- Martםn R. and Suבrez J.E. Biosynthesis and degradation of H2O2 by vaginal lactobacilli. Appl. Environ. Microbiol., 2010; 76: 400-405.
- Dashkevicz M.P., Feighner S.D. Development of a differential medium for bile salt hydrolase-active Lactobacillus spp. Appl. Environ. Microbiol., 1989; 55: 11-16.
- Tahri K., Crociani J., Ballongue J. and Schneider F. Effects of three strains of Bifidobacteria on cholesterol. Letters in Applied Microbiology, 1995; 21: 449-451.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.