ISSN: 0973-7510
E-ISSN: 2581-690X
Different samples for the isolation of cyanobacteria were collected from University garden soil patches, fresh water samples from Tigra dam near Gwalior city. Samples collected were subjected to purification and isolation of cyanobacteria in specific media (BG11) under light condition. After 15 to 20 days of incubation they were able to grow in provided BG11 media with blue green appearance in flasks. Microscopy of the samples revealed that most of the isolates obtained and purified were filamentous with several types of movements. Biochemical characterization such as catalase, oxidase and urease tests was performed in which they were observed weakly positive for urease test while negative for oxidase and catalase test. Chlorophyll and carotenoid estimation was done along with antibacterial activity of cyanobacterial extract. Molecular characterization of the isolates was carried out with the extraction of DNA from each purified cyanobacterial strain. The universal and cyanobacterial specific 16S rRNA gene was amplified and sequenced. The sequences were deposited to NCBI and phylogenetic tree was constructed to observe the relatedness among cyanobacterial isolates. After sequencing, isolates were identified as Nostoc sp., Limnothrix sp. from freshwater samples while Phormidium sp. was obtained abundantly from soil samples. Three distinct OTUs were obtained after phylogenetic tree construction showed their evolutionary relatedness. The antibacterial assay of cyanobacterial extract showed their therapeutic potential in the study.
16S rRNA gene, cyanobacteria, OTUs.
Cyanobacteria are highly diversified group of prokaryotic microbial community that exhibits photosynthesis and are oxygenic like plants with a wide evolutionary history stretching billion years back. This oxygenic group of prokaryote contribute to make environment to lesser reducing from highly reducing atmosphere and therefore triggers an evolutionary changes in biodiversity1. The nature of cyanobacteria is fairly similar with eubacteria and therefore it is resembled with prokaryotic genome. They are one of the largest bacteria present on earth2. These tiny organisms known as cyanobacteria, or blue-green algae performs photosynthesis: using sunlight, water and carbon dioxide to produce carbohydrates and, yes mainly oxygen so basically they require water, carbon dioxide and inorganic substances in the presence of light. Apart from marine, freshwater and terrestrial habitats they are present abundantly in extreme environment such as high temperature environment, salinity rich environment, psychrophilic environments and even dry ecosystem3. Photosynthesis is the most important mode of energy metabolism in cyanobacteria, they are called as photosynthetic, oxygenic prokaryotes utilizes sunlight for their growth and uses photosynthetic pigments for energy.
This group of microorganisms is either unicellular or multicellular, carryout the process of photosynthesis using photosystems I and II4. The majority of cyanobacteria are photoautotrophs like plants. Naturally, some of the cyanobacterial species can survive in a condition of complete darkness for long period of time. Some specific cyanobacteria show a separate ability for heterotrophic nutrition5. Several cyanobacterial species are reported from soil ecosystem where they are involved in nutrient element cycling process6. Cyanobacteria are assumed to be first plants to colonize bare areas of soil and rocks. Apart from fixation of CO2 through photosynthesis, cyanobacterial community is involved in the fixation of atmospheric nitrogen. The fixation of CO2 and nitrogen help in the formation of humus. Naturally, cyanobacteria produce a significant amount of polysaccharide that promote the aggregation of soil colloids and therefore soil structure and quality is improved along with improvement of water infiltration. Cyanobacteria found in the subsurface region of soil are well enough to colonize with the plant roots and produce phytohormones and beneficial metabolites, enhancing plant growth and development7.
Stanier and Van Niel8 were the first to have the opinion that blue-green algae must be classified under bacteria. Cyanobacterial community is quite different from eukaryotic microalgae, membrane bound cell organelles and a well defined nucleus, are therefore absent in cyanobacteria. They have prokaryotic ribosomal subunit 70S rather than eukaryotic 80S9,10.
In the present study cyanobacteria were isolated from fresh water and soil ecosystem of Gwalior region and molecular characterization and identification was done to see their phylogenetic relatedness along with their antimicrobial potential.
Isolation of Cyanobacteria
Soil and water samples were collected from different locations of Gwalior city in replicate. Soil samples in the form of patches or blue green mat were washed several times with sterilized water11. Water samples carrying blue green cell mass were also washed 3-4 times with sterilized water. After washing, a fraction of cell mass was transferred to sterilized fresh BG-11 broth media12.After several transfer cultures were purified and confirmed by microscopic analysis. All strains were maintained on BG11 broth media and sub-cultured to BG-11 agar test tube. Purified samples were subjected to fresh BG11 media under sufficient flux of light and 22±2 °C temperature. After every one month they were transferred to fresh BG-11 media to make them in log phase condition. To prevent clump formation, cyanobacterial cultures are shaken by hand several times on daily basis.
Microscopy and Biochemical characterization
Each cyanobacterial sample was taken from the grown cultures in flask to the clean glass slide with a drop of sterilized water. Samples were covered with coverslip and observed under light microscope at 40X magnification. Certain biochemical tests (catalase test, urease test and oxidase test) were performed on all the samples of cyanobacteria from different habitats to know the biochemical characterization of cyanobacteria along with photosynthetic pigment estimation by spectrophotometer. The Chlorophyll and carotenoid pigments were extracted from approximately 200 mg freshly harvested cyanobacterial biomass by taking methanol: water (9:1, v/v, 4ml, thrice) as solvent. Afterwards, supernatant was separated from cell debris by centrifugation at 12,000 × g for 5 min, supernatant having chlorophyll was quantified for total chlorophyll content13.
Preparation of cyanobacterial extracts
Cyanobacterial cells were filtered from the culture media with the help of sterilized filter paper and dried at 60 °C in hot air oven and then weighing of cells was done (20 mg approx). Afterwards cells are sonicated gently in sterile tubes and mixed with methanol for 2ml with shaking for 5 min. After that, the solvent was removed through separating funnel for further assay.
Antibacterial activity and screening of cyanobacterial isolates
The antibacterial activities of cyanobacterial extracts were assayed by nutrient agar disc diffusion method14. Pathogenic Gram-negative bacteria, Escherichia coli was used as tested pathogen to observe the antibacterial activity. Three wells were made in the nutrient agar plate with a 30 µl lawn of tested pathogens and wells were filled with 150 µl of cyanobacterial extract, 50 µl of known antibiotic tetracycline and 100 µl of methanol. Plates are subjected to incubation at 37 °C one-two days and zone of inhibition was measured. Antibiotic Tetracycline (50 µg) was used as positive control in the experiment.
Isolation of genomic DNA of cyanobacterial isolates
Cyanobacterial cultures obtained from different sites under study were selected for the molecular characterization. The genomic DNA from cyanobacterial cells are extracted by the protocol as described by Singh et al15 with some modifications. Freshly growing pure cultures of cyanobacteria were harvested in mid to late exponential phase (15 days ) by centrifugation (10,000 rpm ; 5 min ) in sterile microcentrifuge, supernatant discarded and the pelleted cells were washed twice with TE buffer. After washing, the content was kept in 500 µL SET buffer (75 mM NaCl, 25 mM EDTA pH 8 and 20 mM Tris). The cell suspension was added with 10 µL lysozyme solution (10 mg/mL) and incubated at 37 °C for 60 minutes. The following reagents were then added: 10 µL of sodium dodecyl sulfate (10%), and 10 µL of proteinase K (10 mg/mL). The reaction mixture was kept at 55 °C for 2 hr in water bath for the lysis of cyanobacterial cells. To this content, 50 µL NaCl (5M) was added along with equal volume of Chloroform: Isoamyl alcohol (24:1) and water saturated phenol. Again the mixture was centrifuged at 10,000 rpm for 5-10 min. Then after, aqueous layer was transferred to a fresh microcentrifuge tube carefully. To the aqueous phase, 2 volumes of chilled 95% Isopropyl alcohol was added along with 0.1 volumes of 3M Sodium acetate and kept at 4 °C for 30 minutes. The precipitated DNA was pelleted by centrifugation at 12,000 rpm for 10 min. The DNA pellet was washed with 70% ethanol and the pellet was dried by keeping at 37 °C for 10 min. The DNA was then dissolved in about 50-100 µL of milli Q water and stored at 4 °C for further analysis.
16S rRNA gene Amplification
PCR reaction was set up for the amplification of conserved prokaryotic chronometer 16S rRNA gene. The cyanobacterial specific primers cya106f (5’CGGACGGGTGAGTAACGCGTGA3’) and cya781r (5’GACTAC TGGGGTATCTA ATCCCATT3’) were used to obtain approximately 650-bp fragment of 16S rDNA16. The 100 µL PCR reaction mixture contains 50-90 ng DNA template, 1X Taq buffer, 10 mM dNTP mix solution, 1.5 mM MgCl2, and 2 U of Taq DNA polymerase (Bangalore Genei, India). The PCR reaction was performed in a thermocycler (Bio-Rad) with an initial denaturation at 94 °C for 5 minutes followed by 34 cycling of reactions with conditions consisting of 45 s at 94 °C (denaturation), 45 s at 54 °C (annealing) and 1 min at 72 °C (Primer extension) and a final extension period of 5 min at 72 °C. The PCR reaction mixture and the amplification conditions were same as described by Meena et al17. The PCR product was visualized and resolved by agarose gel electrophoresis using 1.5 % agarose (0.6 gm agarose in 50 ml 1X TAE Buffer) stained with EtBr (Ethidium bromide) and visualized and gel images were digitalized using a gel documentation system (Alpha-Imager).
16S rRNA gene sequencing and identification
The amplified PCR products (16S rRNA) of representative isolates from each sample is subjected to purification by QIA quick purification kit (Qiagen). The purified PCR product samples are then subjected to sequencing followed by sequence analysis. The nucleotide sequencing was done by using 3130xl Applied Biosystems ABI prism automated DNA sequencer by Sanger sequencing method with the help of fluorescent terminators attached to ddNTPs (Big Dye, Applied Biosystems). Phylogenetic identity of bacteria was determined by BLAST result after alignment of the nucleotide sequences using sequence alignment software i.e., clustal X18. The result of sequencing will be assessed by BLAST (Basic Local Search Alignment Tool) and other bioinformatics tools. The cyanobacterial species will be identified after BLAST analysis and other similarity searches at species level.
The nucleotide accession numbers assigned by NCBI are from KY753382 to KY753387.
Isolation of Cyanobacteria
It was observed that the cyanobacterial isolates obtained from soil and fresh water samples were able to grow and proliferate within 15 to 20 days. Cyanobacteria isolated from soil patches grow little bit slow but after 20 to 25 days of incubation they proliferate well in the autoclaved BG 11 media under proper light conditions. Fresh water samples also took 15-20 days for cyanobacterial culture proliferation. The log phase culture was then subjected to light microscopic study. A total 10 purified cyanobacterial isolates (FW1 to FW10) were obtained from freshwater and 10 from soil sample (S1 to S10) after purification through several transfers.
Biochemical characterization
The cyanobacteria observed were filamentous in structure and were motile also. Some of them were septet and some of them were non septet filamentous cyanobacteria. Whenever observing the slides, movement of the cyanobacterial cultures was also obvious. Since cyanobacterial isolates were recovered and purified from different habitats, were showing different morphological diversity. The structures of the cyanobacterial isolates were quite clear under microscope showing septet or non septet filamentous cyanobacteria. By comparing the microscopic structure with earlier studies, isolates from soil patches were identified as Phormidium sp. and Oscillatoriales class of cyanobacteria abundantly. Fresh water isolates were showing the morphology similarity with Anabaena and Nostoc sp. It was very clear by looking at the grown culture of cyanobacteria from soil having more chlorophyll content as compared to fresh water cyanobacteria. Now the biochemical analysis of chlorophyll and carotenoid reveals more chlorophyll content i.e. (0.1915 mg/g to 0.232 mg/g fresh weight) obtained in the soil cyanobacterial cell biomass extract as compared to freshwater cyanobacterial isolates (0.167 m/g to 0.1896 mg/g) and carotenoid content in soil cyanobacterial cell biomass extract (0.00161to 0.0022 mg/g) as compared to fresh water extract (0.00157 to 0.00231 mg/g).
16S rRNA gene sequencing and phylogenetic analysis
In all cyanobacterial samples, 16S rRNA gene was amplified with variation in their intensity. The gene product visualized was 600 bp in size as appeared in the agarose gel electrophoresis (Fig 2). For sequence analysis, six potent cyanobacterial strains screened by antibacterial activity were selected. This nucleic acid based tool was used to for the identification of cyanobacteria. Based on 16S rRNA gene sequencing, the strains were identified as Nostoc sp., Limnothrix sp. from fresh water samples while from soil samples Phormidium sp. were found to be abundant. The homology after BLAST analysis gives 98% to 99% similarity range. There were three distinct operational taxonomic units (OTUs) obtained from the phylogenetic tree constructed (Fig.1). The phylogeny constructed with a bootstrap value around 80 to 100 indicates a better phylogenetic representation of cyanobacterial lineages.

Fig. 1. Cyanobacterial samples purified from the garden soil and fresh water in cyanobacterial specific media BG-11
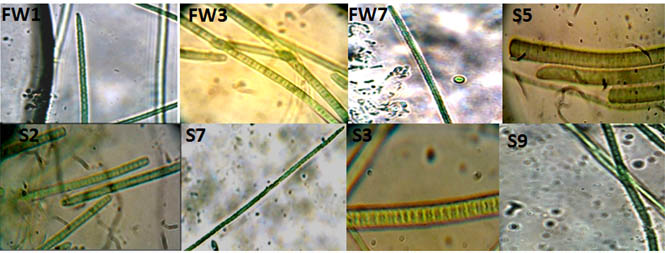
Fig. 2. Different filamentous cyanobacterial cells were obtained after isolation of pure culture. Light microscopy images of some of the cyanobacterial isolates, appearing as filamentous blue green algae with different morphological characteristics
The phylogenetic tree constructed represents the evolutionary relationship and position of the cyanobacterial taxa showing relatedness based on nucleotide sequence analysis.
Purified cyanobacterial cultures are very important and useful for their biological study. Gwalior region of Madhya Pradesh is one of the important gird agro-climatic region and still there are very few reports on isolation, characterization and diversity of blue green algae from these important habitat. The cyanobacterial cells has been reported with a distinct polymorphism and their accommodation with change in the environment, therefore diversity analysis based on their morphology is not sufficient and justified19.
Table (1):
Description of samples collected from different locations in Gwalior
S. No. |
Location |
Geographic position |
Type of the sample |
Sample pH |
|---|---|---|---|---|
1 |
University Garden soil |
26° 13′ 5.8332” N and 78° 10′ 58.1916” E |
Soil patches |
8 |
2 |
University Garden soil |
26° 13′ 5.8332” N and 78° 10′ 58.1916” E |
Soil patches |
7.8 |
3 |
Tigra Dam |
Lat. 26.218326 and Long.78.016188 |
Fresh water |
8.5 |
4 |
Tigra Dam |
Lat. 26.218326 and Long.78.016188 |
Fresh water |
8.5 |
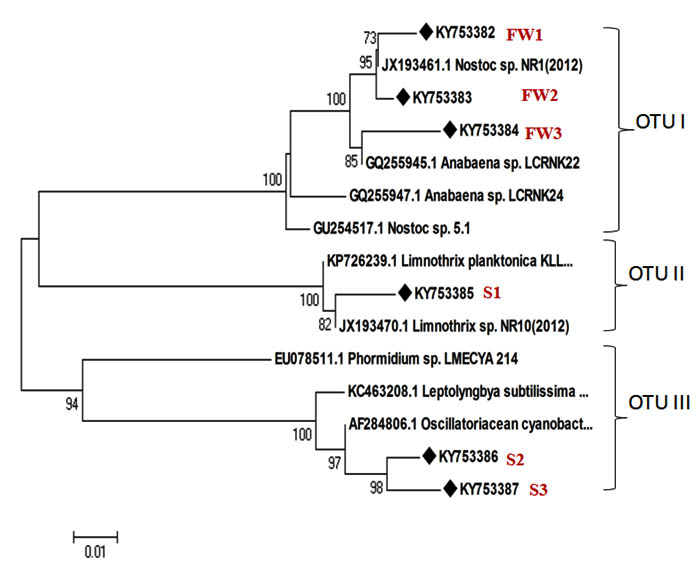
Fig. 3. Phylogenetic tree showing relatedness among different strains of cyanobacteria obtained from soil and water ecosystem (u sign shows the isolates sequenced in this study)
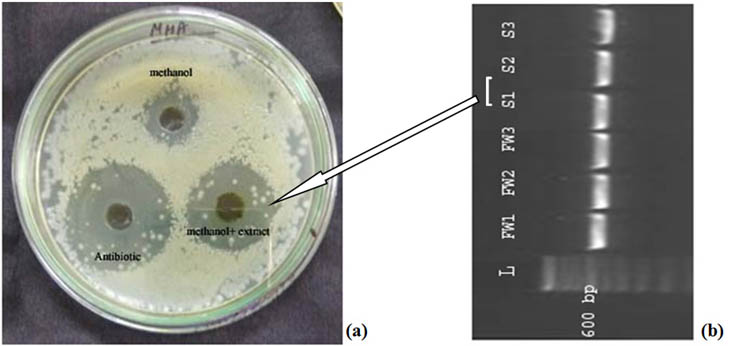
Fig. 4. (a) A significant antimicrobial activity of the isolate Limnothrix sp. (S1) was obtained against E.coli. with a prompt clear zone of inhibition in well diffusion assay. (b) 16S rRNA gene amplified products image after agarose gel electrophoresis.
Cyanobacterial isolates screened on the basis of antibacterial activity against pathogenic bacteria were selected for their identification by molecular characterization. Moreover a number of biotic interactions affect the survival of cyanobacterial cultures. Earlier reports describe the antibacterial potential of several cyanobacterial strains against numerous pathogenic bacterial strains20. In the experiment conducted by Thummajitsakul et al20, the range of zone of inhibition against E. coli was found to be between 8-10 mm while in our study, the zone of inhibition against E. coli was found to be more than 10 mm in case of Limnothrix sp. extract rather Phormidium sp. extract. Apart from E. coli other pathogenic strains such as Bacillus cereus, Klebsiella aerogenes, Micrococcus luteus, and Staphylococcus aureus are also being used to observe the antibacterial potential of cyanobacterial extracts20, 21, 22, 23. The ribosomal gene sequences obtained were utilized in the construction of phylogenetic tree to see their relatedness and evolutionary relationship. Different distinct clusters obtained in the phylogenetic tree, highlighting the cyanobacterial diversity24. The current study revealed the genetic as well as biochemical basis for the characterization of the cyanobacterial isolates obtained from freshwater and soil ecosystems. In one of the earlier investigations, the morphological and phylogenetic evaluation was done precisely for different cyanobacterial genera such as Anabaena, Aphanizomenon, Trichormus and Nostoc25 and genetic data in this study did not support the distinction of benthic Anabaena strains from planktic Anabaena and Aphanizomenon as compared to morphological data. The cyanobacteria have been reported to exhibit distinct morphological diversity. Since ribosomal gene based phylogenetic study is a reliable method for diversity analysis, provide a better understanding of their identification and classification. Moreover, the potential of cyanobacterial extract was determined, showing metabolic ability of cyanobacteria against pathogenic bacteria in the environment by showing the zone of inhibition in experiment. In one of the related investigations, the diversity analysis of cyanobacterial cells was represented from fresh water and marine ecosystem in which eight morphologically distinct cyanobacterial cells were discussed and their phylogeny was also observed using molecular chronometer 16S rRNA gene24.
The present study was an effort to isolate cyanobacteria from two different ecosystem to analyze their diversity. This diversified group of organisms was abundant in soil and water ecosystem. In the current investigation a morphological diversity of cyanobacteria was assessed along with biochemical and molecular characterization. Their ability to perform photosynthesis in presence of light differentiates from other non photosynthetic, heterotrophic group of bacteria. The variation in chlorophyll content in various cyanobacterial isolates enlightens the fact that they are diverse metabolically also. Out of several cyanobacterial isolates, few of them were found to be very promising in terms of antibacterial activity. The molecular chronometer 16S rRNA gene was selected in the study for molecular characterization. The gene sequencing and sequence analysis of this conserved universal gene will provide a suitable classification and identification of these diverse cyanobacterial strains at species level. The current investigation revealed the cyanobacterial diversity analysis along with their important antibacterial potential against pathogenic bacteria. The cyanobacterial cells obtained in this study can be exploited for the pharmaceutical industry in future as a bioactive natural compound.
ACKNOWLEDGMENTS
Authors are highly obliged to Dr. Ashok K. Chauhan, Aseem Chauhan and Amity University Madhya Pradesh, Gwalior for their research encouragement and proper facilities available at Institute of Biotechnology, AUMP.
- Olson, John M. Photosynthesis in the Archean Era. Photosynthesis Research, 2006; 88: 109-17.
- Berry, J.P., Gantar, M., Perez, M.H., Berry, G. & Noriega, F.G., Cyanobacterial toxins as allelochemicals with potential applications as algaecides, herbicides and insecticides. Marine Drugs, 2008; 6: 117-146.
- Fogg, G. E., Stewart, W. D. P., Fay, P. and Walsby, A. E. The Blue-Green Algae, Academic Press, London & New York, 1973; 459.
- Castenholz, R. W. & J. B. Waterbury, Group I. Cyanobacteria. In Staley, J. T., M. P. Bryant, N. Pfennig & J. G. Holt (eds), Bergey’s Manual of Systematic Bacteriology, Williams & Wilkins, Baltimore, 1989; 3: 1710-1727.
- Fay, P. Heterotrophy and nitrogen fixation in Chlorogloea fritschii. J. Gen. Microbiol. 1965; 39: 11-20
- Whitton, B.A. Diversity, ecology and taxonomy of the cyanobacteria. In: N.H. Mann and N.G. Carr [Eds] Photosynthetic Prokaryotes. Plenum Press, New York, 1992; pp 1-51.
- Issa, A.A., Abd-Alla, M.H. & Mahmoud, A.L.E. Effect of biological treatments on growth and some metabolic activities of barley plants grown in saline soil. Microbiol. Res., 1994; 149: 1-4.
- Stanier, R. Y., and Van Niel, C. B. The concept of a bacterium. Arch. Mikrobiol. 1962; 42: 17-35.
- Fay, P. and Van Baalen, C. [Eds]. The Cyanobacteria. Elsevier, Amsterdam, 1987; pp. 534
- Bryant, D.A. [Ed.] The Molecular Biology of Cyanobacteria. Kluwer Academic Publishers, Dordrecht, 1994; pp. 879
- Singh P., Minj R.A., Kunui K., Shaikh Z.M., Suradkar A., Shouche Y.S., Mishra A.K. and Singh S.S. A new species of Scytonema isolated from Bilaspur, Chhatisgarh, India.Journal of Systematics and evolution, 2016; 9999:1-9
- Rippka, R., Deruelles, J., Waterbury, J.B., Herdman, M. and Stanier, R.Y. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J. Gen.Microb., 1979; 111: 1-61.
- Gotz T., Windhovel U., Boger P. and Sandmann G. Protection of photosynthesis against ultraviolet-B radiation by carotenoids in transformants of the cyanobacterium Synechococcus PCC7942. Plant Physiol, 1999; 120: 599-604
- Bauer A.W., Kirby W.M.M., Sherris J.C., Turck M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol., 1996; 45(4): 493-496.
- Singh D.P., Prabha R., Kumar M. and Meena K.K. A rapid and standardized phenol – free method for the isolation of genomic DNA from filamentous cyanobacteria, Asian J. Exp. Biol. Sci., 2012; 3(3): 666-673.
- Garcia-pichel, F., Nu, U. and Muyzer, G. PCR Primers to Amplify 16S rRNA Genes from Cyanobacteria. Applied and Environmental Microbiology, 1997; 63: 3327-3332.
- Meena KK, Kumar M, Mishra S, Ojha SK, Wakchaure GC, Sarkar B. Phylogenetic study of methanol oxidizers from Chilika-Lake sediments using genomic and metagenomic approaches. Indian J Microbiol., 2015; 55:151–162.
- Jeanmougin, F., Thompson, J. D., Gouy, M. et al. ‘Multiple sequence alignment with Clustal x’, Trends Biochem. Sci., 1998; 23: 403–405.
- Dor, I. and Ehruch, A. The Effect of Salinity and Temperature Gradients on the Distribution of Littoral Microalgae in Experimental Solar Ponds, Dead Sea Area, Israel. Marine Ecology, 1987; 8: 193-205.
- Thummajitsakul S., Silprasit K. and Sittipraneed S. Antibacterial activity of crude extracts of cyanobacteria Phormidium and Microcoleus species, African Journal of Microbiology, 2012; 6(10): 2574-2579
- Rao B.D. Antibacterial activity of freshwater cyanobacteria. J. Algal Biomass Utln. 2015; 6(3): 60- 64
- Malathi, T., Ramesh Babu, M., Mounika, T., Snehalatha D. and Rao B.D. Screening of cyanobacterial strains for antibacterial activity, Phykos, 2014; 44(2): 6-11.
- Madhumathi V., Deepa P., Jeyachandran S., Manoharan C. and Vijayakumar S. Antimicrobial Activity of Cyanobacteria Isolated from Freshwater Lake, International Journal of Microbiological Research, 2011; 2(3): 213-216.
- Faldu, N., Patel, S., Vishwakarma, N.P., Singh, A.K., Patel, K. and Pandhi, N. Genetic Diversity of Marine and Fresh Water Cyanobacteria from the Gujarat State of India. Advances in Bioscience and Biotechnology, 2014; 5: 1061-1066.
- Rajaniemi, P., Hrouzek, P., Kastovská, K., Willame, R., Rantala, A., Hoffmann, L., et al. Phylogenetic and Morphological Evaluation of the Genera Anabaena, Aphanizomenon, Trichormus and Nostoc (Nostocales, Cyanobacteria). International Journal of Systematic and Evolutionary Microbiology, 2005; 55: 11-26.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


