ISSN: 0973-7510
E-ISSN: 2581-690X
Halophilic bacteria are excellent source of enzymes not only salt stable but also it can withstand and carry out reactions very efficiently under extreme conditions. Screening of bacteria from different hyper saline environment in Kanyakumari Coast, South India led to the isolation of total 111 culturable heterotrophic bacteria. Screening of Halophiles for the DNase production led to 23 isolates and from which maximum DNase producing 4 strains were selected through DNase well diffusion method. The potential isolate KVCMST-8A 12 showed maximum zone of inhibition at 35mm after 72 hours, which were able to grow optimal in media with 2-40% of salt. According to its phenotypic characteristics and comparatively partial 16S rRNA sequence, the halophilic KVCMST-8A 12 was identified as member of Bacillus cereus family.
DNase, Halophilic, Bacillus cereus, in vitro assay, Substrate gel.
Marine environment is the prime reservoir of biological diversity and the marine microorganisms are recognized to be rich source of novel compounds. In India about 1000 natural products were derived from marine microbes (Suthindhiran et al., 2010). Marine microorganisms are mostly unexplored and marine environment is rich source of microorganisms producing novel and efficient compounds (Sujatha et al., 2005). Microbes form the topmost source of industrially important enzyme producers due to their rapid doubling time when compared to plants or animals (Kumar and Takagi, 1999). The ability of marine bacteria to synthesis variety of enzymes with good stability at higher temperature, alkaline conditions and high salinity (Berdy 2005) are attractive phenomenon to accomplish extremophilic organisms in biotechnological processes.
DNase catalyses the hydrolysis of deoxyribonucleic acid by breaking down phosphodiester bonds (Nishino and Marikawa, 2002), consequently DNase is considered to have a potential role in DNA utilization and nutrient cycling in the environment (Mulcahy et al., 2010). The balance of extracellular DNA in the marine environment is largely regulated by DNase, which is important for the functioning of deep sea ecosystems (Dell anno and Danovaro 2005). The role of DNases in cellular metabolism of nitrogen and phosphorous suggests that DNase secretion may also play an important role in the fate of extracellular DNA in the natural environment, particularly with regarded to the dynamics and stability of marine aggregates (Aisha et al., 2019).
Halophiles are the most likely source of enzymes, because not only is their enzyme salt tolerant but also may be active at high temperature, and pH values (Gomez and Steiner 2004). The isolation of moderate halophiles with capacity to produce extracellular enzymes will provide the possibility to have optimal activities at different salt concentrations (Ventosa et al., 1998). Halophiles are now gaining more access to industrial microbiology and biotechnology because halophiles grow at high salt concentration and this minimize the risk of contamination during cultivation (Oren, 2006), also it has great potential uses in industrial processes (Ventosa and Nieto, 1995).
Marine bacteria, especially the benthic microbes could utilize the DNA from dead and degraded organism as sole source of energy. Hence, the attempt to isolate the marine bacteria with potential and unique DNase activity from sediments off Kanyakumari coast, Southern tip of India was initiated. This is a preliminary report on the isolation and characterization of marine bacteria with DNase activity in the largely untapped Kanyakumari coastal areas.
Isolation of marine bacteria
Sediment samples were collected from 3 sampling points, off coast Kanyakumari and transferred aseptically to lab. The samples were serially diluted and plated on Zobell marine agar and incubated at room temperature for 48 h. The bacterial isolates were selected at random on the basis of their prominence and distinctness in colony morphology and pigmentation. The isolates were screened for their capacity to produce DNA hydrolyzing activity on DNase test agar plate.
DNase well diffusion method
The well diffusion method was followed for DNase test employing the standard procedure (Sanchez and Colom, 2010). Petri plates were prepared by pouring 20 ml of DNase test agar and allowed to solidify. The 48 h culture broths inoculated with the bacterial isolates were added to the wells on the agar. After incubation at 37°C for various time points (24 h, 48 h, & 72 h) the zone of inhibition was measured.
In vitro DNase assay
The activity of DNase was studied by the reaction of cleavage of calf thymus DNA (20µg/ml) in 10mM Tris -HCl buffer ( pH 7.5) containing 2.5mM MgCl2, 0.1mM CaCl2, and DNase (0.1-2U) at 33°C for 1-15 min. The reaction mixture was supplemented with EDTA (final concentration 2mM) and incubated at 65°C for 10 min. The degradation products were analyzed by electrophoresis in 1% agarose gel stainined with ethidium bromide.
DNase Activity Gel System
For substrate gel, the methodology of Shikawa et al., 1994 was followed. In brief, after SDS – PAGE, the gel was washed with 10mM Tris – HCl (pH 7.8) / 5mM mercaptoethanol at 50°C for 1 h to remove SDS, and then with 10 mM Tris – HCl (pH 7.8), at 4°C overnight. Then the gels were incubated in 10 mM Tris – HCl (pH 7.8), containing 1mM mercaptoethanol and indicated concentration of MgCl2, CaCl2 at 37°C for 3 h. When the gel was stained with 0.5µg/ml ethidium bromide, apparent endonuclease activity was detected as dark areas on fluorescent background by UV transilumination of the gel.
Effect of NaCl concentration on DNase production of potential isolate
The activity of crudes in potential strains KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6 and KVCMST-8A 12 in presence of NaCl (2-40%) was measured by standard assay method and DNase activity through well diffusion method.
Characterization of isolate
Characterization of the isolate was carried out by traditional biochemical methods and molecular characterization of the 16S rRNA gene of the potential isolate was done through commercial sequencing and the organism after analysis with CLUSTAL X, the phylogenetic relationship was established using phylip and MEGA 4 and submitted to Genbank.
Marine environment is bestowed with high microbial consortium and biologically productive ecosystem inspite of the extreme environmental conditions. To thrive in these extreme conditions of physical and chemical changeovers, the marine bacteria have developed strategies to adapt to this environment. To maintain their structure and physiology, they are compelled to produce specialized metabolites and potent extracellular hydrolytic enzymes to assimilate the nutrients available around them. Although utilization of dissolved proteins has been investigated to some extent (Hollibaugh and Azam, 1983), the mechanism of utilization of dissolved macromolecules in the marine environment has not been extensively studied. Few references are available on the rapid hydrolysis of extracellular DNA in the marine environment by cell associated and extracellular nuclease. In the present work, the bacterial isolates obtained from the sediment samples collected from various depths off coast Kanyakumari was analyzed for their ability to produce extracellular enzyme, DNase.
Isolation of marine microbe (s)
In the deep sea environment, sediments form a reservoir of microbes with enzymatic action with potential biotechnological applications. Kaboyashi et al., 2008 reported a number of cultivable aerobic microbes with variety of enzyme activities from the deep sea floor sediment off shore Shimokita Penishla, Japan. In a view to isolate microbes with novel enzymes activities in peninsular India, sediment samples at 3 different sampling points off coast Kanyakumari were collected and transported aseptically to the laboratory and labeled as 6A, 7A, and 8A respectively. The collected samples were serially diluted, plated on Zobell marine agar and the culturable heterotrophic aerobic bacteria were isolated and stored for further studies. Totally, 111 isolates (Table.1) were obtained from the sampling points 6A (41), 7A (43) and 8A (27) respectively. The isolated strains were screened for DNase activity by streaking onto a DNase test agar plate. Out of 111 isolates, 23 isolates which showed clearing zone around the streak with pink halos on DNase test agar were selected as DNase producing isolates and further to pick up the best strain, well diffusion method was employed.
Table (1):
Isolation of marine microbes.
Sample No. |
Number of Isolates Obtained |
Number of Isolates Positive for DNase |
|---|---|---|
6A |
41 |
11 |
7A |
43 |
8 |
8A |
27 |
4 |
Total |
111 |
23 |
Screening for extracellular DNase
The culture filtrate of bacterial isolates positive on DNase test agar were analyzed for extracellular DNase production employing agar well diffusion method. Although, all the 23 isolates demonstrated extracellular DNase activity, 2 strains from sampling point 6A designated as KVCMST-6A 20, KVCMST-6A 23, one strain from sampling point 7A denoted by KVCMST-7A 6, and yet another strain from sampling point 8A named KVCMST-8A 12 were shown to produce maximum DNase activity and these strains were used for further studies (Table. 2). DNase test agar is generally used for screening of microbes for DNase production. A thermophilic fungus was isolated and found to produce extracellular DNase employing this method (Landry et al., 2014).
Table (2):
Screening and isolation of extracellular DNase activity by different isolates.
S. No. |
Baterial Isolates |
DNase Test Zone (mm) after 72 Hours |
|---|---|---|
1. |
KVCMST- 6A 2 |
23(a) |
2. |
KVCMST- 6A 3 |
28(b) |
3. |
KVCMST -6A 6 |
21(a) |
4. |
KVCMST -6A 8 |
24(a) |
5. |
KVCMST- 6A 9 |
28(b) |
6. |
KVCMST -6A 13 |
26(b) |
7. |
KVCMST -6A 20 |
34(c) |
8. |
KVCMST -6A 21 |
27(b) |
9. |
KVCMST -6A 23 |
31(c) |
10. |
KVCMST -6A 25 |
27(b) |
11. |
KVCMST -6A 31 |
28(b) |
12. |
KVCMST -7A 1 |
28(b) |
13. |
KVCMST -7A 2 |
24(a) |
14. |
KVCMST -7A6 |
33(c) |
15. |
KVCMST -7A 7 |
24(a) |
16. |
KVCMST -7A 17 |
25(a) |
17. |
KVCMST-7A 18 |
25(a) |
18. |
KVCMST- 7A 19 |
28(b) |
19. |
KVCMST- 7A 25 |
25(a) |
20. |
KVCMST- 8A 2 |
20(a) |
21. |
KVCMST -8A1 2 |
35(c) |
22. |
KVCMST -8A13 |
27(b) |
23. |
KVCMST -8A15 |
25(a) |
Values are mean of trplicate experiments Duccants multiple analysis were carried out.
a=20-25mm, b=26-30mm, c=>30mm
Time – Course study for DNase production
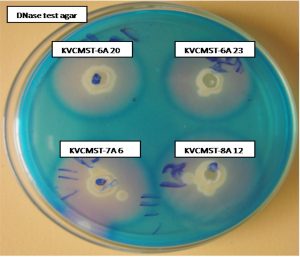
Based on the preliminary screening, four strains that produce maximum zone of > 30mm on 72 hours (Fig. 1) were further analyzed for production of extracellular deoxyribonuclease activity at different time points on DNase test agar employing agar well diffusion method. Test agar plates were incubated 72 h to analyze the production of DNase and the zone measured at 24, 48, and 72 hours. The zone of DNase activity by the strains KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6 and KVCMST-8A 12 were shown in Table. 3. Among the four strains, KVCMST-8A 12 demonstrated a prominent zone right from 24 hours up to 72 hours compared to other strains. However, the other three strains KVCMST-6A 20, KVCMST-6A 23 and KVCMST-7A 6 were not to be excluded as they produce a zone on par with KVCMST-8A 12.
Fig. 1. Extracellular DNase activity of the selected isolates on DNase test agar with toluidine blue.
Table (3):
DNase production of selected bacterial isolates on DNase test agar.
| Bacterial Isolates | DNase Test Zone Diameter in mm | ||
|---|---|---|---|
| 24hrs | 48hrs | 72hrs | |
| KVCMST 6A 20 | 14 | 26 | 34 |
| KVCMST 6A 23 | 16 | 25 | 31 |
| KVCMST 7A6 | 16 | 26 | 33 |
| KVCMST 8A1 2 | 20 | 28 | 35 |
Values are observation of mean ± triplicate experiments
In vitro DNase activity
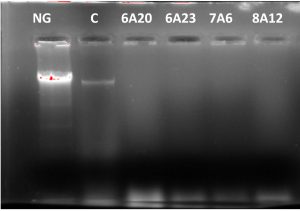
DNA cleavage is one of the important mechanism to arrest the growth of bacteria and viruses, in control of diseases particularly cancer, there is a considerable interest in development of enzymes suitable to cleave DNA. The selected strains KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6, and KVCMST-8A 12 were subjected to DNA cleavage assay on prokaryotic bacterial DNA to demonstrate the in vitro DNase activity employing electrophoretic technique. When the crude extracts of the four isolates were incubated with DNA in tris buffer for 30 minutes and electrophoresed on 0.8% agarose, there was clear cut degradation of DNA (Fig. 2, lane 3 to 6) compared to the control DNA (lane 2) and extract without sample DNA (lane 1). The DNA cleavage potential of the isolates was evidenced by the complete disappearance of DNA. Song and Zhang (2008) have reported a novel – nonspecific nuclease from thermophilic bacteriophage. However, optimum rate of conditions like substrate concentration, enzyme activity, enzyme concentration, specificity, temperature, incubation period, pH, effect of cations and inhibitors need to be carried out.
Fig. 2. Degradation of prokaryotic DNA demonstrated by in vitro DNase activity on agarose gel by the selected bacterial isolates. NG- negative control, C- control DNA
Substrate gel analysis of DNase
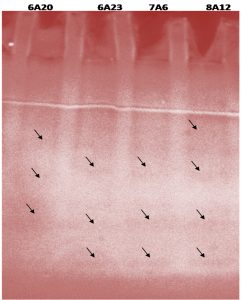
For demonstration of in vitro enzyme activities, substrate gel substrate system plays a vital role. It helps not only to locate the presence of enzyme, but also indicates whether the enzyme is in active state, thus to isolate and purify this enzyme, substrate gel analysis was carried out. In activity gel analysis, the crude extracts of the bacterial isolates KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6, and KVCMST-8A 12 were separated by SDS polyacrylamide gel in which Calf thymus DNA was incorporated. After electrophoresis, the gel was washed and incubated in appropriate buffer and DNase activities were revealed by staining with ethidium bromide. The enzyme activity which appeared as dark band (Fig. 3) was shown in arrow marks.
Fig. 3. Activity gel analysis of DNase activity exhibited by the crude extracts of selected bacterial isolates.
Effect of NaCl on growth and DNase production of potential strains
There are only few reports on halophilic nuclease production by halophiles, including Micrococcus variants, Var halophilus (Kamekura and Onishi 1978) and Bacillus halophilus (Onishi et al., 1983, Ventosa 2015). In order to know the halophilic nature and capacity of DNase production of all strains it is necessary to check the growth and DNase production at various NaCl concentrations (Table. 4 to Table. 7). The isolates were grown at various NaCl concentrations and the overnight grown culture was centrifuged and supernatant was assayed for DNase production on toludine blue DNA agar, DNA hydrolysis zone were noted. The strains exhibited maximum DNase production and growth at 7.5% except KVCMST-8A 12 it showed maximum DNase production at 8% (1.36M) concentration but the strain was capable of producing DNase upto 40% NaCl concentration tested (6.81M) (Fig.4). Onishi et al., 1983 identified a moderate halophile, Bacillus sp N23-2 produced an extracellular nuclease during the growth in 1 to 2M NaCl. KVCMST-8A 12 is a true halophilic microbe as it possess extracellular deoxyribonuclease even at 6.81M.
Table (4):
DNase production for KVCMST-6A20 at various NaCl concentrations on DNase test agar.
| NaCl concentrations (%) | DNase test Zone diameter in mm | Optical Density (600nm) at 24 hrs | ||
|---|---|---|---|---|
| 24 hrs | 48 hrs | 72 hrs | ||
| 2 | 14 | 26 | 34 | 0.832 |
| 4 | 12 | 20 | 22 | 0.744 |
| 6 | 10 | 19 | 21 | 0.547 |
| 8 | 10 | 19 | 21 | 0.421 |
| 10 | 8 | 18 | 20 | 0.156 |
| 12 | 8 | 18 | 20 | 0.143 |
| 14 | 8 | 18 | 20 | 0.133 |
| 16 | 6 | 14 | 16 | 0.127 |
| 18 | 6 | 14 | 16 | 0.105 |
| 20 | 4 | 10 | 12 | 0.085 |
| 22 | 4 | 10 | 12 | 0.067 |
| 24 | 4 | 10 | 12 | 0.043 |
Table (5):
DNase production for KVCMST-6A23 at various NaCl concentrations on DNase test agar.
| NaCl concentrations (%) | DNase test Zone diameter in mm | Optical Density (600nm) at 24 hrs | ||
|---|---|---|---|---|
| 24 hrs | 48 hrs | 72 hrs | ||
| 2 | 16 | 25 | 31 | 0.891 |
| 4 | 12 | 18 | 20 | 0.369 |
| 6 | 12 | 18 | 20 | 0.265 |
| 8 | 10 | 12 | 18 | 0.243 |
| 10 | 10 | 12 | 18 | 0.214 |
| 12 | 10 | 12 | 18 | 0.207 |
| 14 | 8 | 11 | 13 | 0.193 |
| 16 | 8 | 11 | 13 | 0.187 |
| 18 | 6 | 8 | 10 | 0.057 |
| 20 | 6 | 8 | 10 | 0.044 |
| 22 | 6 | 8 | 10 | 0.026 |
| 24 | 6 | 8 | 10 | 0.015 |
Table (6):
DNase production for KVCMST-7A6 at various NaCl concentrations on DNase test agar.
| NaCl concentrations (%) | DNase test Zone diameter in mm | Optical Density (600nm) at 24 hrs | ||
|---|---|---|---|---|
| 24hrs | 48hrs | 72hrs | ||
| 2 | 16 | 26 | 33 | 0.756 |
| 4 | 15 | 20 | 22 | 0.360 |
| 6 | 14 | 16 | 19 | 0.372 |
| 8 | 14 | 16 | 19 | 0.254 |
| 10 | 12 | 14 | 16 | 0.277 |
| 12 | 12 | 14 | 16 | 0.234 |
| 14 | 10 | 12 | 14 | 0.174 |
| 16 | 10 | 12 | 14 | 0.162 |
| 18 | 8 | 10 | 12 | 0.125 |
| 20 | 8 | 10 | 12 | 0.116 |
| 22 | 6 | 8 | 10 | 0.109 |
| 24 | 6 | 8 | 10 | 0.108 |
Table (7):
DNase production for KVCMST-8A12 at various NaCl concentrations on DNase test agar and Effect of NaCl on growth of KVCMST-8A12.
| NaCl concentrations (%) | DNase test Zone diameter in mm | Optical Density (600nm) at 24 hrs | ||
|---|---|---|---|---|
| 24hrs | 48hrs | 72hrs | ||
| 2 | 20 | 28 | 35 | 0.756 |
| 4 | 20.5 | 29 | 30.5 | 0.773 |
| 6 | 21 | 29 | 36 | 0.821 |
| 8 | 22 | 30 | 37 | 0.937 |
| 10 | 18 | 25 | 31 | 0.687 |
| 12 | 16 | 23 | 29 | 0.564 |
| 14 | 14 | 21 | 28 | 0.421 |
| 16 | 13 | 19 | 24 | 0.362 |
| 18 | 10 | 16 | 21 | 0.359 |
| 20 | 10 | 16 | 21 | 0.267 |
| 22 | 8 | 14 | 18 | 0.192 |
| 24 | 8 | 14 | 18 | 0.178 |
| 26 | 7 | 11 | 14 | 0.152 |
| 28 | 7 | 11 | 14 | 0.143 |
| 30 | 7 | 11 | 14 | 0.127 |
| 32 | 6 | 9 | 11 | 0.097 |
| 34 | 6 | 9 | 11 | 0.095 |
| 36 | 6 | 9 | 11 | 0.081 |
| 38 | 6 | 9 | 11 | 0.072 |
| 40 | 6 | 9 | 11 | 0.052 |
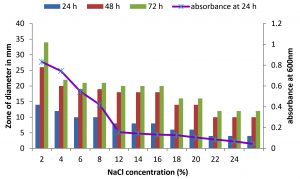
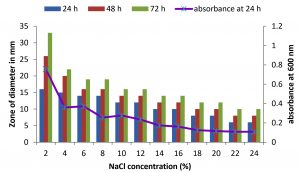
Fig. 4. Effect of NaCl on growth and DNase production of KVCMST-6A20.The zone of clearance in DNase test agar at various time of incubation indicated by bars: 24 hours, 48 hours, 72 hours and its growth indicated by its absorbance at 24 hours.
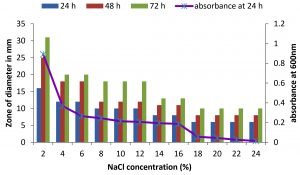
Fig. 5. Effect of NaCl on growth and DNase production of KVCMST-6A23. The zone of clearance in DNase test agar at various time of incubation indicated by bars: 24 hours, 48 hours, 72 hours and its growth indicated by its absorbance at 24 hours.
Fig. 6. Effect of NaCl on growth and DNase production of KVCMST-7A6. The zone of clearance in DNase test agar at various time of incubation indicated by bars: 24 hours, 48 hours, 72 hours and its growth indicated by its absorbance at 24 hours.
Identification of strain
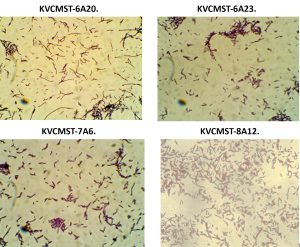
Having known that the crude extracts from the bacterial isolates were (KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6 and KVCMST-8A 12) capable of producing extra cellular DNase and its in vitro DNase activity have been demonstrated both in in vitro DNase activity on agarose gel and activity gel analysis on SDS-PAGE, it is necessary that the taxonomical identity of the bacterial isolates have to be established. Therefore, employing various biochemical tests (Table.3 and Table.4), these isolates were tentatively identified as Bacillus Sp belonging to Bacillus cereus group. They were gram positive (Fig.8) anaerobic, spore forming, rod shaped organisms with smooth, convex, round, glossy and white colony morphology. From which KVCMST-8A 12 tolerated up to 40% NaCl and growth occurs on Mac Conkey agar. All strains showed positive for hydrolysis of starch, casein and DNA, but negative for hydrolysis of gelatin, urease, indole and negative for oxidase.
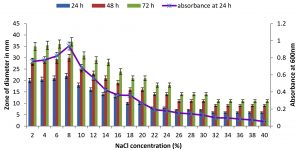
Fig. 7. Effect of NaCl on growth and DNase production of KVCMST-8A12.The zone of clearance in DNase test agar at various time of incubation indicated by bars: 24 hours, 48 hours, 72 hours and its growth indicated by its absorbance at 24 hours.
Fig. 8. Gram staining characteristics of the bacterial strains KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6 and KVCMST-8A 12.
Table (8):
Biochemical Characteristic of the isolates.
| S. No. | Phenotypic/Biochemical | Strains | |||
|---|---|---|---|---|---|
| KVCMST 6A 20 | KVCMST 6A 23 | KVCMST 7A 6 | KVCMST 8A 12 | ||
| 1 | Grams Reaction | G+ rods | G+ rods | G+ rods | G+ rods |
| 2 | Motility | Motile | Motile | Motile | Motile |
| 3 | Pigmentation | Colorless | Colorless | Colorless | Colorless |
| 4 | Colony Morphology | Large Spherical | Large Spherical | Large Spherical | Large Spherical |
| 5 | Spore Formation | + | + | + | + |
| 6 | Growth at | 4ºC | 4ºC | 4ºC | 4ºC |
| 7 | Optimum Temperature | 37ºC | 37ºC | 37ºC | 37ºC |
| 8 | Optimum pH | 7.0 | 7.0 | 7.0 | 7.0 |
| 9 | Optimum % NaCl | 2% | 2% | 2% | 8% |
| 10 | Lactose fermentation on MacKonkey | LF | LF | LF | LF |
| 11 | Indole Production | – | – | – | – |
| 12 | Methyl Red | – | – | – | – |
| 13 | Voges –Prousker | + | + | + | + |
| 14 | Citrate Utilization | + | + | + | + |
| 15 | Starch Hydrolysis | + | + | + | + |
| 16 | Urea Hydrolysis | – | – | – | – |
| 17 | TSI Agar test (1)Acid |
+ | + | + | + |
| (2)Alkaline | + | + | + | + | |
| (3)Gas production | – | – | – | – | |
| (4)H2S | + | + | + | + | |
| 18 | Catalase | + | + | + | + |
| 19 | Oxidase | – | – | – | – |
| 20 | Nitrate Reduction | + | + | + | + |
| 21 | Casein Hydrolysis | + | + | + | + |
| 22 | Gelatin Hydrolysis | – | – | – | – |
| 23 | Acid Fast staining | + | + | + | + |
| 24 | Utilization of Sugars | + | + | + | + |
| 25 | Utilization of Amino acids | + | + | + | + |
(+ present / – absent / W- Weak positive)
Fig. 9. Neighbor joining tree based on 16srRNA sequences showing the phylogenetic relationship between KVCMST-6A 20 and other related species of the genus Bacillus and related reference microorganisms (Accession no MG786828).
Table (9):
Nucleotide informations of KVCMST-6A 20.
| Sequence information | Nucleotide distribution table | |||
|---|---|---|---|---|
| Sequence type | DNA | Nucleotide | Count | Frequency |
| Length | 653bp | Adenine (A) | 166 | 0.254 |
| Name | KVCMST_6A_20 | Cytosine (C) | 140 | 0.214 |
| Weight (single-stranded) | 202.878 kDa | Guanine (G) | 193 | 0.296 |
| Weight (double-stranded) | 403.527 kDa | Thymine (T) | 154 | 0.236 |
| C + G | 333 | 0.510 | ||
| A + T | 320 | 0.490 | ||
Fig. 10. Neighbor joining tree based on 16srRNA sequences showing the phylogenetic relationship between KVCMST-6A 23 and other related species of the genus Bacillus and related reference microorganisms (Accession no MH938091).
Table (10):
Nucleotide informations of KVCMST-6A 23.
| Sequence information | Nucleotide distribution table | |||
|---|---|---|---|---|
| Sequence type | DNA | Nucleotide | Count | Frequency |
| Length | 625bp | Adenine (A) | 171 | 0.274 |
| Name | KVCMST_6A_23 | Cytosine (C) | 145 | 0.232 |
| Weight (single-stranded) | 193.956 kDa | Guanine (G) | 178 | 0.285 |
| Weight (double-stranded) | 386.23 kDa | Thymine (T) | 131 | 0.210 |
| C + G | C + G | 323 | ||
| A + T | A + T | 302 | ||
On the basis of phenotypic characteristics and the comparison of partial 16S rRNA gene sequences, all the isolates KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6 and KVCMST-8A 12 were identified as member of the Bacillus cereus family (accession no MG786828, MH938091, MG786829, MG786830 respectively) (Fig.9 to Fig.12). Bacillus cereus group comprises 11 closely related species, including B. anthracis, along with B. cereus, B. thuringiensis, B. mycoides, B. weihenstephanensis, B. pseudomycoides and the recently identified B. gaemokensis, B. manliponensis, B. cytotoxicus, B. toyonensis, B. bingmayongensis (Liu et al., 2017). Rohban et al., (2009) have isolated hydrolyzing enzymes belonging to the members of genera such as Bacillus and Halobacillus. In their study most of the DNase produces were of the genera Halomonas, however in our study, the best DNase producing isolate was found to be Bacillus Sp.
Fig. 11. Neighbor joining tree based on 16srRNA sequences showing the phylogenetic relationship between KVCMST-7A 6 and other related species of the genus Bacillus and related reference microorganisms (Accession no MG786829).
Table (11):
Nucleotide informations of KVCMST-7A 6.
| Sequence information | Nucleotide distribution table | |||
|---|---|---|---|---|
| Sequence type | DNA | Nucleotide | Count | Frequency |
| Length | 710bp | Adenine (A) | 184 | 0.259 |
| Name | KVCMST_7A_6 | Cytosine (C) | 158 | 0.223 |
| Weight (single-stranded) | 220.56 kDa | Guanine (G) | 211 | 0.297 |
| Weight (double-stranded) | 438.755 kDa | Thymine (T) | 157 | 0.221 |
| C + G | C + G | 369 | ||
| A + T | A + T | 341 | ||
Fig. 12. Neighbor joining tree based on 16srRNA sequences showing the phylogenetic relationship between KVCMST-8A 12 and other related species of the genus Bacillus and related reference microorganisms (Accession no MG786830).
Table (12):
Nucleotide informations of KVCMST-8A 12.
| Sequence information | Nucleotide distribution table | |||
|---|---|---|---|---|
| Sequence type | DNA | Nucleotide | Count | Frequency |
| Length | 1,557bp | Adenine (A) | 394 | 0.253 |
| Name | KVCMST-8A_12 | Cytosine (C) | 354 | 0.227 |
| Weight (single-stranded) | 483.84 kDa | Guanine (G) | 478 | 0.307 |
| Weight (double-stranded) | 962.15 kDa | Thymine (T) | 331 | 0.213 |
| C + G | C + G | 832 | ||
| A + T | A + T | 725 | ||
In conclusion, the strains KVCMST-6A 20, KVCMST-6A 23, KVCMST-7A 6 and KVCMST-8A 12 isolated from the sediment sample collected at a depth of 26.4 m off Kanyakumari coast possess a strong DNase activity as evidenced by the DNase test agar, in vitro DNase assay and gel activity system and with the help of phylogenetic analysis all strains were founds to be belongs to Bacillus cereus family. Among which the strain KVCMST-8A 12 can be regarded as a halophilic, valuable source of the enzyme DNase and demands the purification and characterization of this enzyme which can be used in various biotechnological and biomedical fields. In our present study, we were able to demonstrate a zone greater than the purified enzyme as well as to that of the results obtained by Kample et al., (2011) by employing only the crude culture supernatant of KVCMST-8A 12. This indicates a positive approach that this enzyme can be purified in future and this isolate would promise a valuable source of the enzyme DNase for commercial exploitation.
ACKNOWLEDGMENTS
The authors thank UGC, New Delhi for the financial support. Asha acknowledges DST for the fellowship under Inspire Program (DST/ INSPIRE FELLOWSHIP/2014/69 dt.15.09.2014).
CONFLICT OF INTEREST
The authors declares that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
University Grant Commission, New Delhi, India (F. No. 41-572/2012 (SR) dt.18.07.2012).
ETHICS STATEMENT
Not applicable.
AVAILABILITY OF DATA
All datasets generated or analyzed during this study are included in the manuscript and/or the Supplementary Files.
- Kamekura M and Onishi H. Properties of the halophilic nuclease of moderate halophile Micrococcus varians subsp halophilus. J.Bac., 1978; 133: 59-65.
Crossref - Hiroshi Onishi, Tatsuro Mori, Setsuo Takeuchi and Keiko Tani. Halophilic Nuclease of a Moderately Halophilic Bacillus sp. Production, Purification, and Characterization. Applied and environmental microbiology, 1983; 45(1): 24-30.
Crossref - Hollibaug JT, Azam F. Microbial degradation of dissolved proteins in seawater Limnol. Oceanogr, 1983; 28: 1104-1116.
Crossref - Ventosa A. Taxonomy of new species of moderately halophilic eubacteria, In F.R. Valera. General and Applied Aspects of Halophilic Microorganisms. Plenum Press, New York. 1991; 45-51.
Crossref - Shikowa D, Ohyma H, Yamada TT, Akahashi K, Tanuma S. Identification of an endonuclease responsible for apoptosis in rat thymocytes. Eur. J. Biochem., 1994; 226: 23-30.
Crossref - Ventosa A, Nieto JJ. Biotechnological applications and potentialities of halophilic microorganisms. World J Microbiol Biotechnol., 1995; 11: 85–94.
Crossref - Ventosa A, Nieto JJ and Oren A. Biology of moderately halophilic aerobic bacteria. Microbiology and Molecular Biology Reviews, 1998; 62: 504–544.
Crossref - Ganesh Kumar C and Hiroshi Takagi. Microbial alkaline proteases: From a bio industrial viewpoint. Biotechnol. Adv., 1999; 17: 561-59.
Crossref - Nishino T, Morikawa K. Structure and function of nucleases in DNA repair shape grip and blade of the scissors, Oncogene., 2002; 21: 9022-9032.
Crossref - Oren A. Halophilic Microorganisms and their Environments. Berlin: Kluwer Academic Publishers. 2002.
Crossref - Berdy J. Bioactive microbial metabolites, J Antibiot (Tokyo), 2005. 58: 1-26.
Crossref - Dell Anno A and Danovaro R. Extracellular DNA plays a key role in deep-sea ecosystem functioning. Science, 2005; 309: 2179.
Crossref - Sujatha P, Bapi Raju KVVSN, T Ramana. Studies on a new marine Streptomycete BT-408 producing polyketide antibiotic SBR-22 effective against methicillin resistant Staphylococcus aureus. Microbiol Res., 2005; 160: 119-126.
Crossref - Oren A. The order Haloanaerobiales. In The Prokaryotes. A Handbook on the Biology of Bacteria Volume 4. 3rd edition. Edited by: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E. New York: Springer. 2006. 804-817.
- Kobayashi H, Fumi K, Tadashi K, Norio I. Nuclease Lel 1 and 2 Study Biosci Biotechnol. Biochem., 2008; 64(5): 948-957.
Crossref - Zhang XY, Zhang YJ, Chen XL, Qin QL, Zhago DL, Li TG, Dang HY and Zhang Z. Myroides profundi sp isolated from deep sea sediment of the southern Okinawa Trogh FEMS. Microbiol. Lett., 2008; 287: 108-112.
Crossref - Rohban R, Amoozegar MA and Ventosa A. Screening and isolation of halophilic bacteria producing extracellular hydrolyses from Howz Soltan Lake, Iran. J Ind Microbiol Biotechnol., 2009; 36: 333–340.
Crossref - Mulcahy H, Charron-Mazenod L and Lewenza S. Pseudomonas aeruginosa produces an extracellular deoxyribonuclease that is required for utilization of DNA as a nutrient source. Environ. Microbiol., 2010; 12: 1621–1629.
Crossref - Krish Suthindhiran and Krishnan Kannabiran. Diversity and exploration of bioactive marine actinomycetes in the Bay of Bengal of the Puducherry coast of India. Indian J Microbiol., 2010; 50(1): 76–82.
Crossref - Sanchez M and Colom F. Extracellular DNase activity of Cryptococcus neoformans and Cryptococcus gatti. Rev. Lberom. Micol., 2010; 1: 10-13.
Crossref - Kamble KD, Kample LH, Musaddiq M. Optimization of pH and temperature for deoxyribonuclease producing bacteria obtained from soil P G Department of Biology Shri Shivagi College of Arts Commerce and science Akola (MS) – 444 001, India, 2011.
- Kyle S Landry, Andrea V and Robert E Levin. Purification of an Inducible DNase from a Thermophilic Fungus. Int. J. Mol. Sci., 2014; 15: 1300-1314.
Crossref - Ventosa A, de la Haba R R, Sanchez-Porro C, Papke R T. Microbial diversity of hypersaline environments: a metagenomic approach. Curr Opin Microbiol., 2015; 25: 80 – 87.
Crossref - Yang Liu, Qiliang Lai, Juan Du and Zongze Shao. Genetic diversity and population structure of the Bacillus cereus group bacteria from diverse marine environments. Scientific Reports, 2017; 7: 689.
Crossref - Aisha SM Al-Wahaibi, Emilia Lapinska, Nithyalakshmy Rajarajan, Sergey Dobretsov, Robert Upstill-Goddard and J. Grant Burgess. Secretion of DNases by Marine Bacteria: A Culture Based and Bioinformatics Approach. Frontiers in Microbiology, 2019; 10: 969.
Crossref
© The Author(s) 2020. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.