ISSN: 0973-7510
E-ISSN: 2581-690X
Plastics contribute significantly to global environmental pollution due to their non-biodegradable nature. Biopolymers like polyhydroxybutyrate (PHB), synthesized by microorganisms, offer an eco-friendly alternative. This study reports the isolation and characterization of PHB-producing bacteria from untreated sewage, identifying one potent isolate as Bacillus tequilensis via 16S rRNA sequencing. Preliminary screening using Sudan Black B confirmed PHB accumulation. Optimization of growth conditions revealed that glucose and tryptone were the most effective carbon and nitrogen sources, respectively, yielding 0.10 g of PHB under optimal conditions (pH 7, 37 °C, 48 hours). Notably, substituting glucose with watermelon rind, a sustainable fruit waste, enhanced PHB production to 0.34 g, demonstrating a 3.4-fold increase. The extracted PHB was characterized using FT-IR, confirming the presence of functional groups typical of PHB. This study highlights a dual environmental benefit: valorization of sewage-derived bacteria and agro-industrial waste for sustainable bioplastic production. These findings present a novel approach for cost-effective and eco-friendly PHB synthesis.
Polyhydroxybutyrate, PHB, Plastics, Bacillus tequilensis
Traditionally, plastics have been synthesized using carbon-based polymers derived from non-renewable resources, commonly petroleum. Because plastics are relatively inexpensive, easy to produce, and versatile, their demand is always increasing. Numerous chemicals employed in plastic production are exceedingly hazardous and present a significant risk to all life on Earth.1 It has been demonstrated that certain components of plastic, such as benzene and vinyl chloride, are cancer-causing agents, while others, such as gases and liquid hydrocarbons, are harmful to the environment. These toxic constituents are released during the production and recycling of plastics. Not only do they cause damage to the fragile ecosystem, but they are also a source of a range of ailments, including congenital anomalies to malignancies like cancer, affecting a variety of tissues and systems in humans.2 Thus, it is necessary to provide an eco-friendly option with properties similar to those of plastics.
Biodegradable polymers synthesized using biological sources have been going on for a long period, and polyhydroxybutyrate synthesized from microorganisms has been considered a good substitute for petrochemical plastics.3 Polyhydroxybutyrate (PHB) is a thermostable polyester that degrades quickly and is manufactured by bacterial fermentation. It exhibits an elevated crystallinity of 50%-70%, good gas barrier performance (water vapor permeability is approximately 560 g/Sm2/day), and physical aspects resembling those of polypropylene. In terms of tensile strength at break, PHB has a strength of 25 MPa, and its elastic modulus is 3GPa. However, PHB is not without its drawbacks, including its high fragility (showing 3 -5% tensile elongation at break) and its limited thermal stability (with deterioration becoming apparent above 200 °C). Due to its non-toxic, biocompatible, and biodegradable qualities, PHB is now an attractive option for uses in packaging, medicine, agriculture, and the automotive industry.4,5 However, despite these remarkable attributes, the manufacturing of PHBs is still at an early stage due to their high production costs and is not capable of competing with conventional plastics on a global commercial scale.6 In addition to the microbe strain and substrate type, other growth parameters, such as pH, duration, and temperature, play a pivotal role in the formulation and development of PHB. Therefore, further study is required to find novel species with significantly greater production capacities and to optimize the circumstances for maximum PHB synthesis.7,8
Numerous carbon sources, including glucose, fructose, and lactose, have been extensively investigated as feedstocks for PHB production, as raw materials contribute to nearly half of the total manufacturing cost.9 Bacillus spp. have been particularly noted for their ability to accumulate PHB under optimized conditions.10 In addition to these pure sugars, a wide range of industrial byproducts and waste-derived carbon sources such as molasses, fruit waste, potato peels, waste frying oil, sugarcane bagasse hydrolysate, and various wastewater streams have been explored for their potential to support microbial PHB synthesis.11-20 Fruit wastes, rich in carbohydrates and sugars, represent a sustainable and low-cost carbon source that can be valorized for PHB production.21 Among these, watermelon peel an abundant and nutrient-rich fruit waste has shown significant potential as an effective carbon substrate for PHB-accumulating microbes such as Limnothrix planktonica, a cyanobacterial strain isolated from mangrove ecosystems.12,22,23 Watermelon peel can not only reduce production costs but also address environmental concerns associated with agro-industrial waste disposal. Studies indicate that such fruit waste hydrolysates support substantial microbial growth and synthesis of PHB, offering an eco-friendly and economically sustainable substitute to traditional carbon sources.12,23,24 While glucose often yields the highest PHB accumulation,24,25 the vaporization of watermelon peel provides a promising route to sustainable bioplastic production, reducing reliance on pure sugars and costly substrates.9 The enumeration and isolation of PHB-producing bacteria were conducted using a sewage sample, which is recognized for its high nutrient content derived from waste.15,26 The production of polyhydroxybutyrate was characterized using FT-IR and was subjected to optimization studies using different parameters like temperature, incubation time, carbon and nitrogen sources, etc. PHB production of the isolate was also compared with the cultures of Bacillus subtilis and Pseudomonas aeruginosa as positive controls.
Collection of the sewage sample and isolation of PHB-producing bacterial strains
The bacterial strains that produce PHB that were utilized in this investigation were isolated from a sewage sample obtained from the premises of Kristu Jayanti (Deemed to be University) (Latitude 13.0579° N, Longitude 77.6430° E). For bacterial strain isolation, we mixed 1 g of soil with 100 mL of sterile distilled water from the collected sewage sample. After vigorous mixing, to obtain a 10-1 dilution, 1 mL of the suspension was mixed with 9 mL of sterile distilled water. We repeated this serial dilution to get dilutions 10-2 to 10-6. From the respective dilutions, 0.1 mL was plated onto a nutrient medium by the spread plate method and incubated at 30 °C for 48 hours. The total number of colonies formed was enumerated by colony-forming unit (CFU), and distinct colonies were identified and streaked in separate media plates to obtain pure cultures.7,14
Screening for PHB-producing strains by the Plate assay
Bacterial isolates were cultured individually on Luria Bertani (LB) agar plates, each forming a distinct colony.27 Following this, the plates were kept at 37 °C for a 48 hour incubation. These plates were incubated at 37 °C for a duration of 48 hours. Following the incubation period, Sudan Black B stain (0.3% w/v in ethanol) was evenly distributed over the plate on top of the bacterial colonies and left undisturbed for 30 minutes. For the removal of excess stain adhering to the bacterial colonies, ethanol solution (96%) was added gently over the colonies on the plates and observed for the presence of pigment. Colonies that retain the black color after ethanol treatment are identified as PHB-producing colonies.28,29
Cell staining with Sudan Black for the selection of isolates
Following the preparation and heat-fixation of the bacterial smears, the slides were subjected to Sudan Black B staining solution (3% w/v in 70% ethanol) for 10 minutes, subsequently washed with ethanol and distilled water until the slides were completely decoloured. Safranin was used as a counterstain for ten seconds, followed by a water wash and drying. The slide was viewed under a 100x microscope after adding a few drops of immersion oil onto the slide. Bacterial cells that appear black are identified as PHB-producing bacteria, whereas those that picked up the counterstain are indicated as negative for producing PHB.28,30
Morphological characterization of isolates by Gram staining
The bacterial smear was prepared on a glass slide fixed to which the primary stain, Crystal violet, was applied. Following a staining period of 60 seconds, the slide was tilted to remove the stain, and any excess stain was washed away with water. Mordant iodine solution was then poured over the smear as a mordant for 60 seconds to fix the dye. Running water was used to rinse the iodine solution. After shaking off any remnant water, a decolorizer, like an ethanol and acetone mixture, was added for 5 seconds and immediately rinsed with water. After that, safranin was applied and fixed for approximately 40-60 seconds to counterstain the smear. After washing off the safranin, the slide was dried and examined under a 100x magnification.
Identification of PHB-producing isolate by 16S rRNA sequencing
The bacterial genomic DNA was isolated using the PCI (phenol, chloroform, isoamyl) separation technique after inoculating PHB-producing isolates in fresh nutrient medium and incubating at 37 °C for 24 hours. The 27F/1492R primer was used for the PCR amplification of the 16S rRNA gene. The amplified 16S rDNA was sent for sequencing at BioCipher Labs, Bangalore. The sequence obtained from the isolate was submitted to GenBank with the accession number PP340962. GenBank has deposited the sequence obtained from the isolate with the accession number PP340962. Through the use of the BLAST search tool, the partial sequences were compared to the NCBI-BLAST database (https://blast.ncbi.nlm.nih.gov/Blast.cgi).31 The sequences were also compared in the form of a phylogenetic tree in MEGA software to understand the evolutionary relationship and similarities within the genus of the isolate.
Optimization of medium components and incubation conditions for obtaining high PHB yield
The isolates that produce PHB are selected for subsequent extraction and quantification. Out of the two isolates producing PHB, the one producing a higher yield is chosen and taken up for further growth optimization. Each microorganism requires its own optimum carbon, nitrogen, pH, temperature, and incubation time for growth. These external factors are essential in the PHB yield of the bacteria as well, and thus need to be optimized according to the isolate utilized.7,17
Carbon source optimization
Carbon is an essential nutrient for bacterial growth and directly influences PHB accumulation.32 To evaluate the impact of various carbon sources on PHB yield, maltose, glucose, sucrose, glycerol, and starch were supplemented in minimal media, and PHB production was quantified. In this study, glucose derived from treated watermelon peel waste a rich and sustainable fruit waste substrate was used to assess its efficacy as a cost-effective source of carbon for upscaling the production of PHB. Fruit wastes such as watermelon peel contain abundant sugars and carbohydrates, supporting robust microbial growth and biopolymer synthesis, as demonstrated in similar studies with Limnothrix planktonica.12 The minimal medium with 1% concentrations of each selected carbon source was incubated at 30 °C in a shaker incubator for 48 hours. After incubation, PHB was extracted and quantified to determine yield differences across substrates.11
Nitrogen source optimization
Following the carbon source optimization, the medium containing the optimal carbon source was added with various nitrogen sources, such as tryptone, peptone, and gelatin, at a concentration of 1%. The cultures were incubated for 48 hours in a rotary shaker at 30 °C, followed by quantification to select the optimal nitrogen source for high PHB yield.33
Optimization of pH
Optimization of pH is done by culturing the isolate in minimal media containing the optimal carbon and nitrogen source at various pH levels of 5, 5.8, 6.4, 6.8, 7, 7.4, 7.8, and 8.2. The culture media were incubated for 48 hours in a shaker incubator, followed by the quantification of PHB yield using a UV spectrophotometer.28
Optimization of temperature
The optimal temperature for procuring high PHB yield is achieved by supplementing the minimal media with the selected best carbon and nitrogen source and inoculating the isolated strains. The cultures were cultured at different levels of temperatures 24 °C, 30 °C, 37 °C, 40 °C, 45 °C, and 50 °C for a constant period of 48 hours in a rotary shaker. After incubation at their respective temperature, PHB yield was quantified by a UV spectrophotometer.28
Optimization of Incubation Time
Optimal incubation time is determined by placing the cultures in different periods, such as 12, 24, 48, and 72 hours. Following the incubation, bacterial growth and PHB yield are determined by spectroscopic techniques like UV spectrophotometry.28
Use of natural waste – watermelon peels as a carbon source
Watermelon peels were collected and rinsed with water to eliminate dirt and debris. The fruit waste was subsequently chopped or blended into small pieces to enhance the surface area for enzymatic hydrolysis. A solution of amylase enzyme was prepared by dissolving the appropriate amount in water following the manufacturer’s instructions. The amylase solution was added to the fruit waste in a container and thoroughly mixed. The mixture was then heated to a temperature of 60-70 °C for 2 hours. Occasional stirring was performed to ensure even heating. The pH of the mixture was monitored using a pH meter, aiming for an optimal pH of 7.0. If necessary, the pH was adjusted by adding the required acid or base solution. Upon completion of the enzymatic reaction, the glucose solution was separated from the solid fruit waste using filtration and centrifugation. Subsequently, the glucose solution underwent purification techniques like distillation to eliminate impurities and obtain pure glucose.34,35
Extraction of PHB from the bacteria and its quantification
The dispersion method was utilized to extract PHB, using sodium hypochlorite and chloroform, with a few minor modifications, as reported by Balakrishnan and Prabakaran.36 Centrifugation was done to separate the PHB-positive cells at 5000 rpm for 5 minutes at room temperature, and then they were washed in PBS (Phosphate buffer solution). The weight of the cell pellets was calculated after they had been air-dried for 20 minutes at room temperature. Chloroform and 6% sodium hypochlorite were added to the cell pellet in an amount of 12.5 µL each. The mixture was then incubated overnight at 30 °C after this addition. After centrifugation at 6000rpm in room temperature where a three-phase solution was obtained. The bottom chloroform phase contains PHB. The chloroform phase (which contains the PHB) was then carefully transferred to a fresh tube and kept for drying. After the drying process, PHB was scraped out, washed with ethanol, and quantified. Quantification is done using spectroscopic techniques like a UV Spectrophotometer at 235 nm after mixing the filtered pellet with concentrated. H2SO4 (sulphuric acid). The PHB granules are converted into crotonic acid by H2SO4, which is then estimated.17,28,36
Characterization of the extracted PHB
Functional/chemical groups like CH3, CH2, C=O, OH, are considered crucial molecules that decide the presence of PHB, and advanced spectroscopic techniques like FTIR (Fourier transform infrared spectroscopy) can provide a means for characterizing these molecules. One milligram of extracted PHB was dissolved in chloroform, and the chloroform-pellet mixture was thoroughly mixed with potassium bromide (KBr). This was followed by drying at 100 °C for approximately four hours. The FT-IR spectra were recorded between 4000 and 600 cm-1 using a Spectrum 65 FT-IR.37,38
Isolation of PHB-producing bacteria
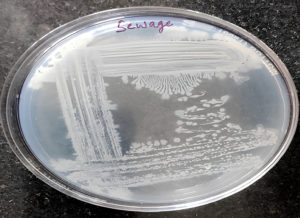
By serial dilution, independent colonies of bacteria were obtained from the sewage sample. Aliquots were taken in order from 10-2 to 10-6 and plated on LB agar. Seven isolates were found in the 10-6 place, out of which two isolates produced PHB, confirmed by the Sudan black plate assay.39 Out of these two, one produced PHB significantly and was chosen as the study isolate (Figure 1). Two positive controls- Bacillus subtilis and Pseudomonas aeruginosa were compared with this isolate as well.
Isolate identification
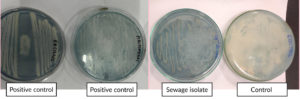
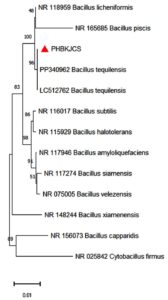
The PHB-producing isolate was cultured separately and taken for identification. Before its identification, the isolate of the smear was made on a microscopic slide and stained with Sudan black B dye to confirm PHB production, where PHB granules present inside the cytoplasm were stained blue-black, confirming PHB production in the isolate.40 Sudan black plate assays were also done in the isolate, positive, and negative control plates, as seen in Figure 2. Sudan black B plate assay was conducted similar to the work of Lathwal et al., and Susianingsih et al.,28,41 which involved incubation of the culture plates with Sudan Black B stain and subsequent rinsing with alcohol, thereby producing bluish black colonies if the strain produces PHB, which was complementary to the results obtained (positive result with bluish black colonies). The isolate identification was done by identifying the morphological characterization of the bacteria by traditional Gram staining. Gram staining revealed the bacteria as Gram-positive rod-shaped bacteria. This was subsequently followed up with 16S rRNA sequencing, and the bacterial strain was found to be Bacillus tequilensis, confirmed by BLAST analysis of a 99% match with the isolate sequence. The phylogenetic relationships were established with the Neighbor-Joining approach,42 and the most suitable tree configuration is presented. Bootstrap values, derived from 500 replicates, are displayed above the branches to signify the confidence level in the taxonomic categorization.43 The tree is proportional, with branch lengths reflecting evolutionary distances. Using the Maximum Composite Likelihood method,44 these distances were calculated as the number of substitutions per site. Thirteen nucleotide sequences were included in this analysis. Ambiguous sites were excluded through pairwise deletion. A total of 1,342 nucleotide positions were aligned to the final alignment. All evolutionary studies were conducted utilizing the MEGA11 software45 (Figure 3).
Figure 2. Sudan Black plate assay showing bluish black colonies in the isolate and positive controls
Optimization of growth conditions for PHB-producing isolate
Effect of nutrient components on PHB production
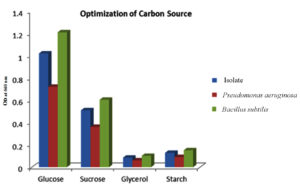
Carbon and nitrogen derived from various types were studied to optimize the minimal media for the isolate- Bacillus tequilensis, while simultaneously comparing with optimization done in the two positive controls. Prior research has established that the choice of carbon source exerts a significant influence on PHB yield. These findings, depicted in Figure 4, showcase the impact of the various carbon sources, like glucose, sucrose, glycerol, and starch, on the production of PHB. These results underscore the variability in the ability of bacteria to utilize different carbon sources, a characteristic dependent on multiple factors. Specifically, the investigation revealed that glucose stands as the most favorable carbon source for promoting PHB yield, with sucrose and starch following as relatively effective options. This suggests that monosaccharides such as glucose are readily metabolized by bacteria, thereby enhancing the production of PHB, whereas a complex molecule, such as starch, on the other hand, poses a significant challenge for the production of PHB.
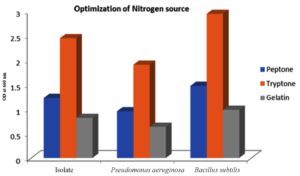
In Figure 5, the impact of various nitrogen sources, namely peptone, tryptone, and gelatin, on PHB production was visually represented. It was observed that the strain achieved its highest PHB yield when cultured in a production medium enriched with tryptone. Hence, it was evident that complex nitrogen sources like tryptone had a beneficial effect on bacterial growth. In contrast, the utilization of alternative sources for nitrogen resulted in a notable reduction in PHB production. Conversely, bacteria of the same genus, Bacillus wiedmannii, exhibited the highest PHB yield when cultured in a production medium supplemented with a simple nitrogen source like peptone, implying the importance of nutrient optimization for each isolate for higher activity.46
The effect of pH media on the production of PHB
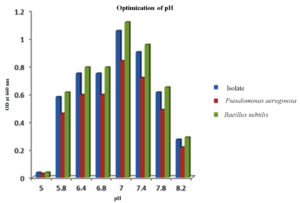
Various levels of pH ranging from 5.0 to 8.2 were systematically assessed with the PHB isolates along with the controls to identify the optimal pH conditions for PHB production. The results revealed that from pH 5.8 to 7.8, the selected isolates exhibited the capacity to synthesize PHB. However, the highest yield of PHB was achieved when the pH was maintained at 7.0 (Figure 6). Interestingly, at pH 5.0, no discernible PHB production was observed.
The effect of temperature on the production of PHB
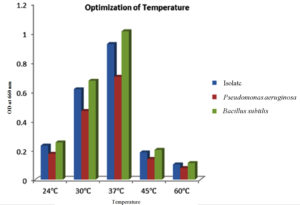
The isolate, along with the two positive controls, were subject to varied temperature incubations 24 °C, 30 °C, 37 °C, 45 °C and 60 °C after the media and pH optimization. The highest quantity of PHB was produced at 37 °C, after which there is a decline in PHB production as seen in Figure 7. It may be due to the fact that the enzyme that is responsible for PHB synthesis may be more active at mesophilic temperatures than at extreme temperatures, resulting in diminished enzyme and protein function.46
The effect of incubation time on the production of PHB
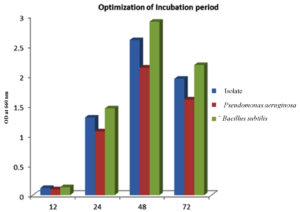
The isolate, along with the controls, was cultivated in the optimized media enriched with glucose, tryptone with a pH of 7 at 37 °C for various durations, specifically 12, 24, 48, and 72 hours. It was observed that the PHB yields exhibited a time-dependent increase, with the highest yield achieved after 48 hours of growth.47 The optimal incubation duration for PHB yield was determined to be 48 hours, post which the yield decreases, as seen in Figure 8.
Enhancement in the production of PHB using watermelon rinds
The Bacillus tequilensis cultures, when grown under optimal parameters, i.e., in media supplemented with glucose and tryptone as the nutrient source, incubated for 48 hours at 37 °C, yielded 0.1 g of PHB48 and are illustrated in Figure 9. It has been demonstrated that the use of fruit and vegetable waste as substrate offers an effective way to increase the yield of PHB and has been studied in a certain number of Bacillus species.49 Watermelon rinds were blended in media, and the produced PHB yield was compared with the PHB produced in conventional optimized media. The yield of the PHB was now found to be 0.34 g, as shown in Figure 10. This further supports the usage of watermelon rinds/peels as a carbon source can aid in increased PHB biomass production, serving as an economical strategy to enhance PHB production in such microorganisms.
Characterization of the extracted PHB using FT-IR Analysis
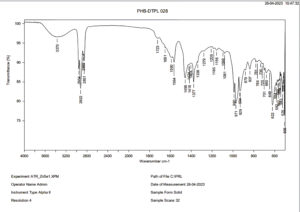
The FT-IR spectra analysis indicates the presence of aliphatic functional groups within PHB. Notably, the spectrum exhibits a prominent absorption peak in the range of 1723-1338 cm-1, as illustrated in Figure 11. Furthermore, the spectrum also reveals absorption bands corresponding to C-H stretching, which occur within the range of 2954-2851 cm-1, as well as O-H stretching, which occurs within the region of 3370-3200 cm-1. These findings are consistent with a study conducted by Thapa et al.,7 wherein similar results were obtained when analyzing PHB extracted from various bacteria, including from Bacillus species like Bacillus pasteurii and Bacillus cereus, as well as Arthrobacter sp. and Micrococcus luteus.
Plastics are widely used across industries, such as the Healthcare industry, the packaging industry, and the automotive industry, to name a few. Approximately 311 million tonnes of plastic were produced worldwide in 2014, and the number is expected to increase by two-fold within two decades, continuing to grow towards 2050, according to a study reported by Khairul et al.50 Plastic pollution poses significant environmental and health hazards. There is indeed a pressing need to switch to safer alternatives to mitigate the hazards caused by such non-degradable polymers. One such approach adopted in our study is the usage of watermelon waste. Watermelon is considered a tropical fruit rich in sugars, and watermelon wastes contain nearly 7% sugars and trace quantities of proteins and lipids, with the remaining constituents comprising mainly water.51 These watermelon peel wastes are mainly composed of lignocellulosic fibers and are a substance of high nutrient profile, as stated by Hasanin and Hashem.34 Therefore, watermelon peels serve as an excellent substrate not only for the sustenance of microorganisms but also aid in the intracellular synthesis of PHB (a biodegradable polymer), which is a safer, non-toxic polymer in certain organisms like Bacillus sp.21 PHB has numerous applications, including the development of agriculture and construction materials, car interior components, electrical gadgets, bottles, and containers. Due to its compatibility with bodily tissues, PHB finds use in pharmaceutical and medical fields, including surgical sutures, wound dressings, and eye devices. PHB can be utilized as a packaging material. Its impact as a packaging material is supported with studies in meat salad packaging.52-55
While PHB production using Bacillus species and agro-waste substrates has been reported previously, this study is novel in isolating Bacillus tequilensis from untreated sewage and demonstrating a 3.4-fold enhancement in PHB yield using watermelon rind waste under optimized conditions. The integration of sewage-derived bacteria with fruit waste valorization provides a dual environmental advantage: it utilizes low-cost, abundant waste streams while promoting sustainable bioplastic synthesis. To our knowledge, this is the first report combining sewage isolation, watermelon rind supplementation, and optimization of B. tequilensis for enhanced PHB production, which could serve as a model for eco-friendly, scalable biopolymer production.
This study successfully isolated and characterized a polyhydroxybutyrate-producing bacterium, Bacillus tequilensis, from untreated sewage samples. Optimization of growth parameters identified glucose and tryptone as the most effective carbon and nitrogen sources, with maximum PHB accumulation observed at pH 7, 37 °C, and 48 hours of incubation. Notably, replacing conventional carbon sources with watermelon rind waste resulted in a 3.4-fold increase in PHB yield, highlighting the potential of agro-waste valorization for enhanced biopolymer production. FT-IR analysis confirmed the characteristic functional groups of PHB, validating successful polymer synthesis. Overall, the integration of sewage-derived bacterial isolates with fruit waste substrates presents a cost-effective and environmentally sustainable approach for PHB production, with promising substrate for scaling up bioplastic manufacturing.
ACKNOWLEDGMENTS
The authors are thankful to the Department of Life Sciences, Kristu Jayanti (Deemed to be University), for the continued support and for providing facilities during the preparation of this research article.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Saha M, Naik A, Desai A, et al. Microplastics in seafood as an emerging threat to marine environment: A case study in Goa, west coast of India. Chemosphere. 2021;270:129359.
Crossref - Wang JG, Bakken LR. Screening of Soil Bacteria for Poly-β-Hydroxybutyric Acid Production and Its Role in the Survival of Starvation. Microb Ecol. 1998;35(1):94-101.
Crossref - Khosravi-Darani K, Mokhtari ZB, Amai T, Tanaka K. Microbial production of poly(hydroxybutyrate) from C1 carbon sources. Appl Microbiol Biotechnol. 2013;97(4):1407-1424.
Crossref - Atakav Y, Pinar O, Kazan D. Investigation of the Physiology of the Obligate Alkaliphilic Bacillus marmarensis GMBE 72T Considering Its Alkaline Adaptation Mechanism for Poly(3-hydroxybutyrate) Synthesis. Microorganisms. 2021;9(2):462.
Crossref - McAdam B, Brennan Fournet M, McDonald P, Mojicevic M. Production of Polyhydroxybutyrate (PHB) and Factors Impacting Its Chemical and Mechanical Characteristics. Polymers. 2020;12(12):2908.
Crossref - Pachekoski WM, Agnelli JAM, Belem LP. Thermal, mechanical and morphological properties of poly (hydroxybutyrate) and polypropylene blends after processing. Mater Res. 2009;12(2):159-164.
Crossref - Thapa C, Shakya P, Shrestha R, Pal S, Manandhar P. Isolation of Polyhydroxybutyrate (PHB) Producing Bacteria, Optimization of Culture Conditions for PHB production, Extraction and Characterization of PHB. Nepal J Biotechnol. 2019;6(1):62-68.
Crossref - Preethi Rathna R, Kulandhaivel M. Microbial Production of Biopolymer Polyhydroxybutyrate (PHB): Current Challenges and its Application. Asian J Chem. 2023;35(10):2289-2300.
Crossref - Dey P, Rangarajan V. Improved fed-batch production of high-purity PHB (poly-3-hydroxy butyrate) by Cupriavidus necator (MTCC 1472) from sucrose-based cheap substrates under response surface-optimized conditions. 3 Biotech. 2017;7(5):310.
Crossref - Bhagowati P, Pradhan S, Dash HR, Das S. Production, optimization and characterization of polyhydroxybutyrate, a biodegradable plastic by Bacillus spp. Biosci Biotechnol Biochem. 2015;79(9):1454-1463.
Crossref - El-Kadi SM, Elbagory M, EL-Zawawy HAH, et al. Biosynthesis of Poly-β-Hydroxybutyrate (PHB) from Different Bacterial Strains Grown on Alternative Cheap Carbon Sources. Polymers. 2021;13(21):3801.
Crossref - Mishra P, Panda B. Polyhydroxybutyrate (PHB) accumulation by a mangrove isolated cyanobacteria Limnothrix planktonica using fruit waste. Int J Biol Macromol. 2023;252:126503.
Crossref - Amir M, Rizvi SF, Asif M, et al. Polyhydroxybutyrate (PHB) bioplastic characterization from the isolate Pseudomonas stutzeri PSB1 synthesized using potato peel feedstock to combat solid waste management. Biocatal Agric Biotechnol. 2024;57:103097.
Crossref - Amadu AA, Qiu S, Ge S, et al. A review of biopolymer (Poly-β-hydroxybutyrate) synthesis in microbes cultivated on wastewater. Sci Total Environ. 2021;756:143729.
Crossref - Wei YH, Chen WC, Wu HS, Janarthanan OM. Biodegradable and Biocompatible Biomaterial, Polyhydroxybutyrate, Produced by an Indigenous Vibrio sp. BM-1 Isolated from Marine Environment. Mar Drugs. 2011;9(4):615-624.
Crossref - Morgan-Sagastume F, Karlsson A, Johansson P, et al. Production of polyhydroxyalkanoates in open, mixed cultures from a waste sludge stream containing high levels of soluble organics, nitrogen and phosphorus. Water Res. 2010;44(18):5196-5211.
Crossref - Narayanan M, Kandasamy S, Kumarasamy S, Gnanavel K, Ranganathan M, Kandasamy G. Screening of polyhydroxybutyrate producing indigenous bacteria from polluted lake soil. Heliyon. 2020;6(10):e05381.
Crossref - Rathna RP, Kulandhaivel M. Sustainable Synthesis of Biopolymer Polyhydroxybutyrate (PHB) from Agro-residue by Brevibacterium casei with Emphasis on Degradation Analysis. J Pure Appl Microbiol. 2024;18(1):347-366.
Crossref - Nikodinovic-Runic J, Guzik M, Kenny ST, Babu R, Werker A, O Connor KE. Carbon-Rich Wastes as Feedstocks for Biodegradable Polymer (Polyhydroxyalkanoate) Production Using Bacteria. Adv Appl Microbiol. 2013;84:139-200.
Crossref - Sirohi R, Pandey JP, Gaur VK, Gnansounou E, Sindhu R. Critical overview of biomass feedstocks as sustainable substrates for the production of polyhydroxybutyrate (PHB). Bioresour Technol. 2020;311:123536.
Crossref - Sosa AR, Barragan LAP, Reyes AR, Carvajal ETA. Genomic analysis and potential polyhydroxybutyrate (PHB) production from Bacillus strains isolated from extreme environments in Mexico. BMC Microbiol. 2025;25(1):15.
Crossref - Zhao J, Cui YW, Zhang HY, Gao ZL. Carbon Source Applied in Enrichment Stage of Mixed Microbial Cultures Limits the Substrate Adaptability for PHA Fermentation Using the Renewable Carbon. Appl Biochem Biotechnol. 2021;193(10):3253-3270.
Crossref - Khan FI, Aktar L, Islam T, Saha ML. Isolation and Identification of Indigenous Poly-β-Hydroxybutyrate (PHB) Producing Bacteria from Different Waste Materials. Plant Tissue Cult Biotechnol. 2019;29(1):15-24.
Crossref - Sanhueza C, Diaz-Rodriguez P, Villegas P, et al. Influence of the carbon source on the properties of poly-(3)-hydroxybutyrate produced by Paraburkholderia xenovorans LB400 and its electrospun fibers. Int J Biol Macromol. 2020;152:11-20.
Crossref - Kominek LA, Halvorson HO. Metabolism of poly-beta-hydroxybutyrate and acetoin in Bacillus cereus. J Bacteriol. 1965;90(5):1251-1259.
Crossref - Shayesteh H, Laird DW, Hughes LJ, Nematollahi MA, Kakhki AM, Moheimani NR. Co-Producing Phycocyanin and Bioplastic in Arthrospira platensis Using Carbon-Rich Wastewater. BioTech. 2023;12(3):49.
Crossref - Wang H, Guo J, Chen X, He H. The Metabolomics Changes in Luria–Bertani Broth Medium under Different Sterilization Methods and Their Effects on Bacillus Growth. Metabolites. 2023;13(8):958.
Crossref - Lathwal P, Nehra K, Singh M, Jamdagani P, Rana JS. Optimization of Culture Parameters for Maximum Polyhydroxybutyrate Production by Selected Bacterial Strains Isolated from Rhizospheric Soils. Pol J Microbiol. 2015;64(3):227-239.
Crossref - Lee SM, Lee HJ, Kim SH, et al. Screening of the strictly xylose-utilizing Bacillus sp. SM01 for polyhydroxybutyrate and its co-culture with Cupriavidus necator NCIMB 11599 for enhanced production of PHB. Int J Biol Macromol. 2021;181:410-417.
Crossref - Amiri Kojuri S, Issazadeh K, Heshmatipour Z, Mirpour M, Zarrabi S. Production of Bioplastic Polyhydroxybutyrate) with Local Bacillus megaterium Isolated from Petrochemical Wastewater. Iran J Biotechnol. 2021;19(3).
Crossref - Lee SH, Oh YT, Lee DY, Cho E, Hwang BS, Jeon J. Large-Scale Screening of the Plant Extracts for Antifungal Activity against the Plant Pathogenic Fungi. Plant Pathol J. 2022;38(6):685-691.
Crossref - Cavaille L, Albuquerque M, Grousseau E, et al. Understanding of polyhydroxybutyrate production under carbon and phosphorus-limited growth conditions in non-axenic continuous culture. Bioresour Technol. 2016;201:65-73.
Crossref - Getachew A, Woldesenbet F. Production of biodegradable plastic by polyhydroxybutyrate (PHB) accumulating bacteria using low cost agricultural waste material. BMC Res Notes. 2016;9(1):509.
Crossref - Hasanin MS, Hashem AH. Eco-friendly, economic fungal universal medium from watermelon peel waste. J Microbiol Methods. 2020;168:105802.
Crossref - Chaijamrus S, Udpuay N. Production and Characterization of Polyhydroxybutyrate from Molasses and Corn Steep Liquor produced by Bacillus megaterium ATCC 6748. Agric Eng Int: CIGR Journal. 2008;X
- Balakrishnan S, Prabakaran G. Production of PHB (Bio-plastics) using bio – effluent as substrate by Alcaligens eutrophus. Indian J Biotechnol. 2006;5(1):76-79.
- Patel SKS, Kumar P, Singh M, Lee JK, Kalia VC. Integrative approach to produce hydrogen and polyhydroxybutyrate from biowaste using defined bacterial cultures. Bioresour Technol. 2015;176:136-141.
Crossref - Gonzalez A, Irusta L, Fernandez-Berridi MJ, Iriarte M, Iruin JJ. Application of pyrolysis/gas chromatography/Fourier transform infrared spectroscopy and TGA techniques in the study of thermal degradation of poly (3-hydroxybutyrate). Polym Degrad Stab. 2005;87(2):347-354.
Crossref - Gonzalez M, Rapado M, Gonzalez AP, Perez MG. Radiation-Induced Graft Copolymerization of Metacrylic Acid and Butyl Methacrylate onto Poly(3-hydroxybutyrate). Int J Polym Anal Charact. 2009;14(3):179-195.
Crossref - Mesquita DP, Amaral AL, Leal C, Oehmen A, Reis MAM, Ferreira EC. Polyhydroxyalkanoate granules quantification in mixed microbial cultures using image analysis: Sudan Black B versus Nile Blue A staining. Anal Chim Acta. 2015;865:8-15.
Crossref - Susianingsih E, Kadriah IAK, Nurhidayah. Screening and isolation of PHB (Poly-β-hydroxybutyrate) producing bacteria as an alternative material for disease prevention on the shrimp culture. IOP Conf Ser Earth Environ Sci. 2020;564(1):012053.
Crossref - Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4(4):406-425.
Crossref - Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evol Int J Org Evol. 1985;39(4):783-791.
Crossref - Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci. 2004;101(30):11030-11035.
Crossref - Tamura K, Stecher G, Kumar S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol. 2021;38(7):3022-3027.
Crossref - Hamdy SM, Danial AW, Gad El-Rab SMF, Shoreit AAM, Hesham AEL. Production and optimization of bioplastic (Polyhydroxybutyrate) from Bacillus cereus strain SH-02 using response surface methodology. BMC Microbiol. 2022;22(1):183.
Crossref - Singh G, Mittal A, Kumari A, Goel V, Aggarwal NK, Yadav A. Optimization of Poly-β-Hydroxybutyrate Production from Bacillus species. Europ J Biol Sci. 2011;3(4):112-116
- Rivas-Castillo AM, Valdez-Calderon A, Angeles-Padilla AF, et al. PHB production by Bacillus megaterium strain MNSH1-9K-1 using low-cost media. Braz J Microbiol. 2024;55(1):245-254.
Crossref - Musa B, Ado SA, Joseph GL, et al. Production of Bioplastic by Local Strains of Bacillus subtilis using Watermelon Peels as Substrate. UMYU J Microbiol Res UJMR. 2022;7(2):43-48.
Crossref - Khairul Anuar SZ, Nordin AH, Nur Husna SM, et al. Recent advances in recycling and upcycling of hazardous plastic waste: A review. J Environ Manage. 2025;380:124867.
Crossref - Nasharudin MIH, Mahmud N, Rahim MHA. Watermelon waste as a growth media substitute for bacterial cellulose production. AIP Conf Proc. 2024;3023(1):020001.
Crossref - Ansari S, Sami N, Yasin D, Ahmad N, Fatma T. Biomedical applications of environmental friendly poly-hydroxyalkanoates. Int J Biol Macromol. 2021;183:549-563.
Crossref - Atalayin Ozkaya C, Ertan B, Kaftan Ocal G, et al. Polyhydroxybutyrate as a Novel Biopolymer for Dental Restorative Materials: Biological and Morphological Analysis. Polymers. 2025;17(3):313.
Crossref - Adnan M, Siddiqui AJ, Ashraf SA, et al. Polyhydroxybutyrate (PHB)-Based Biodegradable Polymer from Agromyces indicus: Enhanced Production, Characterization, and Optimization. Polymers. 2022;14(19):3982.
Crossref - Arrieta MP, Samper MD, Aldas M, López J. On the Use of PLA-PHB Blends for Sustainable Food Packaging Applications. Materials. 2017;10(9):1008.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.