ISSN: 0973-7510
E-ISSN: 2581-690X
Staphylococcus aureus is a known enterotoxin-producing foodborne pathogen; however, the invasion mechanism of the bacterium into intestinal cells remains unclear. The aim of this study was to determine whether S. aureus can invade Caco-2 cells, and to elucidate the gene responsible for this invasion. Caco-2 cells were infected with S. aureus strains NCCP10862, KACC13236, KACC10768 and KACC11596, and their invasion efficiencies were evaluated. Proteins found in the invasive and noninvasive S. aureus strains were labelled with isobaric tags for relative and absolute quantification (iTRAQ), and the gene encoding the protein responsible for S. aureus invasion was deleted using a temperature-sensitive plasmid, pIMAY. The Caco-2 cell invasion efficiencies of the wild type and mutant S. aureus were then compared. Among the S. aureus strains, only NCCP10862 and KACC10768 were able to invade Caco-2 cells, and these strains had a higher level of pyruvate formate lyase (Pfl) protein expression than that of the noninvasive strains. Therefore, a pflB-deletion mutant of KACC10768 was prepared, which revealed a 60% decrease in invasion efficiency when compared to the wild type. These results indicate that certain S. aureus strains can invade intestinal cells, and the protein encoded by the pfl gene is involved in this invasion.
Staphylococcus aureus, invasion, pyruvate formate lyase, foodborne pathogen.
Staphylococcus aureus, a gram-positive and facultative anaerobic bacterium, is a foodborne pathogen that is a global health threat1. S. aureus is generally found on human skin, and when food is exposed to S. aureus-laden skin, it can result in food contamination and disease. S. aureus can grow and multiply up to 106 CFU/g under optimal growth conditions2, 3, and produces endotoxins in the food4,5. Staphylococcal foodborne illness is caused by consuming food contaminated with S. aureus enterotoxins (SEs), and symptoms of the illness include vomiting and diarrhea6. SEs (SEA-SEE, SEG-SEJ, and SER-SET) are heat stable and resistant to low pH. SEA is the most common enterotoxin recovered from foodborne illness outbreaks followed by SED and SEB7,8.
Therefore, this bacterium is considered as an enterotoxigenic rather than an invasive infection-causing bacterium. The possibility of S. aureus intestinal colonization and its implications on host health were previously reported by Gries et al.9, wherein they suggested that the cecal mucus layer facilitates the intestinal colonization of methicillin-resistant S. aureus. Furthermore, Vriens et al.10 reported the presence and interaction of S. aureus in the rectum of patients in surgical intensive care and liver transplant units. However, the mechanism underlying the intestinal invasion of S. aureus remains unclear.
Therefore, this study aimed to determine whether S. aureus can invade human intestinal cells, and to elucidate gene for the invasion.
Inoculum preparation
Bacterial strains were purchased from National Culture Collection for Pathogens (NCCP; Seoul, Korea) and Korean Agricultural Culture Collection (KACC). Four S. aureus strains, NCCP10862, KACC13236, KACC10768, and KACC11596, were cultured on mannitol salt agar (MSA; Becton, Dickinson and Company, Sparks, MD, USA) and then inoculated into tryptic soy broth (TSB; Becton, Dickinson and Company). They were incubated at 35°C for 24 h. Hundred microliters of the culture was sub-cultured into fresh TSB, followed by incubation at 35°C for 24 h. The subcultures were centrifuged at 1912 xg and incubated at 4°C for 15 min. Thereafter, the cell pellets were washed twice with phosphate buffered saline (PBS; pH 7.4; 0.2 g KH2PO4, 1.5 g Na2HPO4·7H2O, 8.0 g NaCl, and 0.2 g KCl in 1 L of distilled water), and eventually suspended in 10 mL of PBS. The suspensions to be inoculated were diluted with PBS to OD600 = 0.01.
Intestinal cell invasion assay
The intestinal cell line, Caco-2, was purchased from the Korean Cell Line Bank (Seoul, Korea). The cells were cultured for 72 h at 37°C and 5% CO2 in Eagle’s minimum essential medium (MEM; Gibco, Penrose, Auckland, New Zealand), supplemented with 20% fetal bovine serum (FBS; Gibco) and 1% penicillin–streptomycin (PS; Gibco), until the Caco-2 cells formed a monolayer, which was then detached by trypsin. Thereafter, the cells were washed twice with Dulbecco’s phosphate buffered saline (DPBS; Welgene, Daegu, Korea), and suspended in MEM supplemented with 20% FBS. For the invasion assay, diluted Caco-2 cells, at a final concentration of 5.0 x 104 cells mL-1, were placed in 24-well plates and incubated at 37°C and 5% CO2 for 48 h. One milliliter of each S. aureus culture was inoculated into the plate and incubated for 2 h at 37°C and CO2 to allow S. aureus invasion into Caco-2 cells. Aliquot of the inoculum was properly diluted in BPW and 100 µL of the diluents was spread-plated on TSA plates. Colonies grown on TSA plates were counted after 24 h of incubation, and the bacterial cell counts were used as initial bacterial cell counts. Planktonic S. aureus cells in the supernatant were discarded, and the attached S. aureus cells were treated with gentamicin (50µg mL-1) or PBS for 2 h. The Caco-2 cell membrane was then lysed using 0.1% TritonX-100 for 20 min. About 100-µL aliquots of the suspensions were spread-plated on TSA, and incubated at 37°C for 24 h. The colonies were manually counted to enumerate the invasive S. aureus. Invasion efficiency (%) was calculated as follows: (invaded bacterial cell counts/initial bacterial cell counts) x 100.
Isobaric tags for relative and absolute quantification (iTRAQ) screening for invasion-related protein
To identify the protein responsible for S. aureus invasion of Caco-2 cells, bacterial cells harvested from 1–1.5 mL of S. aureus culture were lysed by suspension in lysis buffer (iNtRON, Seongnam, Korea) and lysostaphin (0.5 mg mL-1; Sigma Aldrich, St. Louis, MO, USA), and incubation at 37°C for 2 h. The lysates were then treated with SMART bacterial solution (iNtRON), and centrifuged (8,000 xg and 5 min). Proteins in the supernatants were stored at -80°C until used. The proteins were precipitated using acetone at -20°C. After acetone was then removed by vaporization, the precipitated proteins were resuspended in urea buffer (50 mmol l-1 Tris pH 8.3, 3 M urea), and digested by trypsin (1:10) at 37°C for 16 h. The digested proteins were labeled with the iTRAQ reagent suspended in isopropanol (Global standard; GS-113, 10862-116, 13236-117, 10768-119, and 11596-121) for 1 h at 25°C, and then, completely dried in a SpeedVac centrifuge. The labeled samples were fractionated using cation exchange chromatography on Agilent 1100 system (Agilent Technologies, Santa Clara, CA, USA), and analyzed using LC-MS/MS with a hybrid quadrupole-TOF LC-MS/MS spectrometer (Applied Biosystems, Foster City, CA, USA). Peptides and proteins were then identified and quantified using the ProteinPilot Software 2.0.1 (Applied Biosystems), and differences in protein expression among the S. aureus strains were analyzed.
Gene mutation
iTRAQ data revealed that the invasive S. aureus had a higher level of pyruvate formate lyase expression (Pfl) than that of the noninvasive strain. Thus, a pflB deletion mutant was constructed as described by Lee et al.11. Table 1 lists the bacterial strains, primers, and plasmids used in this study.
Construction of pIMAY-pflBtrunc
Chromosomal DNA of wild type invasive strain was extracted using DNeasy Blood and Tissue Kit (Qiagen, Hilden, Germany), and was used as template for following PCR amplification. The pflB gene was amplified using pflB-F (PstI) and pflB-R (SalI) primers with PCR conditions as follow; 95oC-5 min, 35 cycles of 95oC-30 sec, 43oC-30 sec and 72oC-1 min, and 72oC-5 min. After the amplified pflB was purified with QIAquick gel extraction kit (Qiagen), the gene was digested with EcoRV (Elpisbio, Daegen, Korea), which targets two sites (900 bp distance) of the pflB gene. The digested pflB fragments were self-ligated with DNA ligation kit (Takara, Shiga, Japan) to form truncated pflB. The truncated pflB and pIMAY that was a shuttle vector were then digested by restriction enzymes, PstI and SalI, and the two DNA fragments were ligated by the DNA ligation kit to produce pIMAY-pflBtrunc.
Transformation and cloning in E. coli DC10B
pIMAY-pflBtrunc was transformed into high-efficiency cloning strain, E. coli DC10B as described by Monk et al.12. Briefly, competent E. coli DC10B cells were prepared using a chemical method13, and stored in -80oC deep freezer until used. The competent cells were mixed with pIMAY-pflBtrunc in microcentrifuge tube on ice, and the transformation was induced by heat-shock at 42oC for 30 sec. The transformants were spread-plated on LB agar containing 15 µg mL-1 of chloramphenicol (Cm). The plates were then incubated at 37°C for 24 h to obtain E. coli DC10B containing pIMAY- pflBtrunc.
Electroporation and allelic exchange
pIMAY-pflBtrunc was extracted from the E. coli DC10B with DNA Plasmid SV kit (GeneAll, Seoul, Korea) according to the manufacturer’s protocol. pIMAY-pflBtrunc was precipitated using ethanol, and the plasmid DNA was then transformed into S. aureus by electroporation. To induce single-crossover integration, the S. aureus strain containing pIMAY-pflBtrunc was grown at 37oC for 24 h in LB broth containing Cm (15 µg mL-1). The culture was streaked on LB agar + Cm, and incubated at 37oC for 24 h. One S. aureus colony from the agar was then cultured in LB broth without Cm at 28oC, followed by streaking on LB agar containing 1 µg mL-1 of anhydrotetracycline (ATc) to induce excitation of remained plasmid fragment. Eventually, the colony was dually streaked on LB agar containing ATc and Cm. The object grown only on LB agar+ATc not on LB agar+Cm was considered as a mutant S. aureus strain which contained truncated pflB instead of original pflB11. Existence of pflBtrunc on S. aureus genome was eventually confirmed by DNA sequencing and PCR analysis using pflB’-F and pflB’-R primers, which were created from sequences lying outside the truncated pflB (Cosmogenetech, Seoul, Korea). DNA sequencing was analyzed using BigDyeTM Terminator v3.1 Cycle Sequencing kit and 373 DNA Analyzer (Applied Biosystems). The lengths of pflB in wild- (2,250 bp) and mutant-type (1,350 bp) strains were compared.
Statistical analysis
The experiments were repeated twice, and two samples were used in each replication (n = 4). Because the replication had random effects on a dependent variable (S. aureus invasion efficiency), a mixed procedure of SAS® (Version 9.2, SAS Institute, Cary, NC, USA) was used to analyze the results, and the least squares means of the invasion efficiency among S. aureus strains were then compared, using pairwise t-test at α = 0.05.
Table (1):
Bacterial strains, plasmids, and primers used in this study
Strains |
Description |
Reference |
|---|---|---|
E. coli DC10B
|
High efficiency cloning strain
E. coli DH10B(dcm-) |
Monk et al.12
|
Plasmid |
||
pIMAY |
E. coli–S. aureus shuttle vector, temperature- sensitive |
Monk et al.12
|
pIMAY-pflBtrunc |
pIMAY ligated with truncated pflB |
This study |
Primer |
Sequence (5’→3′) |
|
pflB-F(PstI) |
GGA ACA AGT ACT GCA GTT ATC GAA |
This study |
pflB-R(SalI) |
TGT TAA CTG TCG ACA TTC TTC TGG |
This study |
pflB’-F |
TAA GCC GTC CAG CAG AAA |
This study |
pflB’-R |
CCG TAT ACA ACG TTT GAA G |
This study |
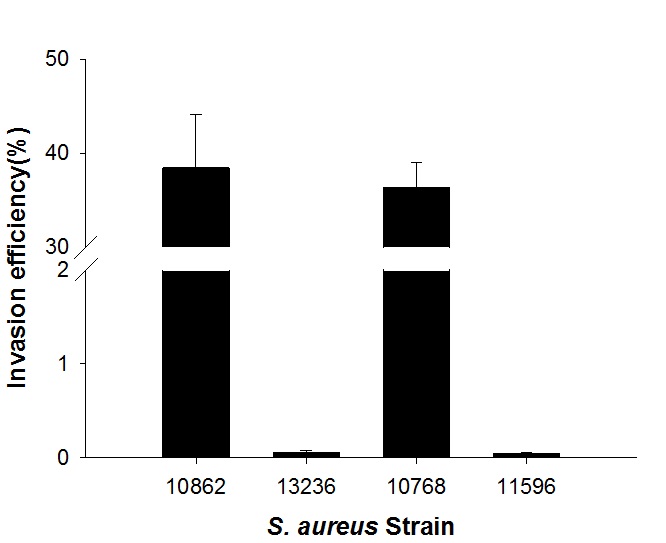
The invasion efficiencies of the S. aureus strains were compared. Among the afore-mentioned strains, S. aureus strains NCCP10862 and KACC10768 showed Caco-2 cell invasion (36.3%–38.4%), but the other strains showed no cellular invasion (Fig. 1). These two strains were clinical isolates, and S. aureus KACC10768 was identified as a methicillin-susceptible strain. These strain-variation was also shown in other studies. Blevins et al.14 showed that regulatory roles of sarA and agr, which control virulence factors in S. aureus, were strain-dependent. Belkum and Melles15 also presented the strain variation of S. aureus strains in pathogenicity. To elucidate the strain variation of S. aureus in Caco-2 cell invasion, the proteins profiles of S. aureus strains NCCP10862 and KACC10768 were screened using iTRAQ. The results indicated that Pfl protein expression was upregulated in the invasive S. aureus strains, followed by an increase in the levels of elongation factor Tu and phosphopyruvate hydratase in the strains (Table 2). The elongation factor Tu and phosphopyruvate hydratase genes were not subjected to mutation because they are essential for S. aureus survival. Thus, only pflB gene was subjected to the mutation.
Table (2):
Relative protein quantification of Staphylococcus aureus strains. 113-Global Standard (GS), 116: S. aureus NCCP10862, 117: S. aureus KACC13236, 119: S. aureus KACC10768, 121: S. aureus KACC11596.
Name |
116:113 |
117:113 |
119:113 |
121:113 |
|---|---|---|---|---|
Elongation factor Tu |
0.4105 |
0.8765 |
5.2007 |
0.7146 |
Phosphopyruvate hydratase |
0.8292 |
0.7503 |
2.7568 |
0.7573 |
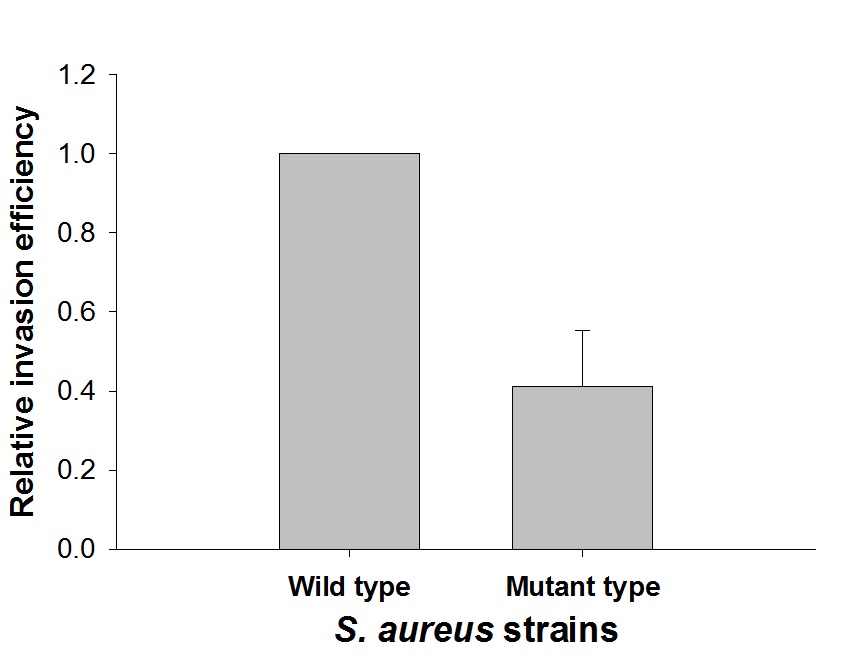
To elucidate the role of Pfl in S. aureus invasion, the pflB mutant of S. aureus was prepared. The truncated pflB (Ca. 900 bp) was constructed by EcoRV. The truncated construct and pIMAY were cut with the same restriction enzymes, ligated, and transformed into E. coli DC10B. The pIMAY-pflBtrunc was then transformed into S. aureus through electroporation, integrated into the chromosome, and the S. aureus pflB mutation was confirmed through PCR. Compared to the wild-type, the invasion efficiency of the mutant strain into the Caco-2 cells decreased by 60% (p < 0.05) (Fig. 2). This result indicates that, although Pfl, known as an oxygen sensitive enzyme, was involved in the conversion of pyruvate to formate, it is crucial for S. aureus invasion into Caco-2 cells (Fig. 2). In a previous study, it was proved that pflB had important role in deeper layer of biofilm formation as Pfl supplies formate on such anaerobic and nutrient-deficient conditions16. Biofilm formation and adhesion are usually first step of S. aureus infection for persistence of bacteria in host, and the colonization and persistence on around of the invaded cells is probably important in the invasion procedure of bacterial cells. Taken together, it can be suggested that pflB may have a role in the intestinal cell invasion of S. aureus in the process of biofilm formation.
In conclusion, although S. aureus is known to cause foodborne illness by release of toxins, and the mechanism of intestinal cell invasion remains unclear, certain S. aureus strains can invade the intestinal cells due to the expression of the pfl gene. Thus, the chronic effect of invasive S. aureus in the intestine should be further researched.
Acknowledgments
We would like to express our heartfelt thanks to Dr. Omar Zayed for English editing of the manuscript and submission process .
Conflicts Of Interest
The authors declare that there are no conflicts of interest.
Authors’ Contribution
AZ, SM NM, LM and AA designed the experiments. AZ and SM performed the experiments. NM and LM analyzed the data and wrote the manuscript. AA supervised and reviewed the manuscript. AZ, SM NM, LM and AA read and approved the manuscript.
Funding
None.
Data Availability
All datasets generated or analyzed during this study are included in the manuscript.
Ethics Statement
Informal written and oral approval was considered before the study from each patient. Study approval was issued and maintained by the local ethical committee at the National Hepatology and Tropical Medicine Research Institute.
- EFSA. Assessment of the public health significance of methicillin resistant Staphylococcus aureus (MRSA) in animals and foods – scientific opinion of the panel on biological hazards. EFSA Journal, 2009; 993: 1e73.
- Foster T. Staphylococcus. In Baron S. Galveston (eds.), Medical Microbiology, 4th Ed. University of Texas Medical Branch at Galveston, 1996.
- Lowy F.D. Staphylococcus aureus infections. New Engl. J. Med., 1998; 339: 520-532.
- Novick R.P. Pathogenicity factors and their regulation. ASM Press, Washington, D.C., 2000.
- Kim J.H., Yoo J.G., Ham J-S., Oh M-H. Direct Detection of Escherichia coli, Staphylococcus aureus, and Salmonella spp. In animal-derived foods using a magnetic bead-based immuno-assay. Korean J. Food Sci. An., 2018; 28: 727-736.
- Argudin M.A., Mendoza M.C., Rodicio M.R. Food poisoning and Staphylococcus aureus entero-toxins. Toxins, 2010; 2: 1751-1773.
- Schmid D., Fretz R., Winter P., Mann M., Hצger G., Styger A., Ruppitsch W., Ladstהtter J., Mayer N., de Martin A., Allerberger F. Outbreak of staphylococcal food intoxication after consumption of pasteurized milk products, June 2007, Austria. Wien Klin Wochenschr., 2009; 121:125–131.
- Veras J.F., do Carmo L.S., Tong L.C., Shupp J.W., Cummings C., Dos Santos D.A., Cerqueira M.M., Cantini A., Nicoli J.R., Jett M. A study of the enterotoxigenicity of coagulase-negative and coagulase-positive staphylococcal isolates from food poisoning outbreaks in Minas Gerais, Brazil. Int. J. Infect. Dis.,2008; 12:410–415.
- Gries D.M., Pultz N.J., Donskey C.J. Growth in cecal mucus facilitates colonization of the mouse intestinal tract by methicillin-resistant Staphy-lococcus aureus. J. Infect. Dis., 2005; 192: 1621-1627.
- Vriens M.R., Fluit A.C., Troelstra A., Verhoet J., Werken C. Staphylococcus aureus rectal carriage and its association with infections in patients in a surgical intensive care unit and a liver transplant unit. Infect. Control Hosp. Epidemiol., 2014; 23: 495-501.
- Lee S., Kim S., Lee H., Ha J., Lee J., Choi Y., Oh H., Yoon Y., Choi K.H. icaA gene of Staphylococcus aureus responds to NaCl, leading to increased biofilm formation. J. Food Prot., 2018; 81: 412-416.
- Monk I.R., Shah I.M., Xu M., Tan M-W., Foster T.J. Transforming the untransformable: application of direct transformation to manipulate genetically Staphylococcus aureus and Staphy-lococcus epidermidis. mBio, 2012; 3: e00277-11.
- Tu Z., He G., Li K.X., Chen M.J., Chang J., Chen L. et al. An improved system for competent cell preparation and high efficiency plasmid transformation using different Escherichia coli strains. J. Biotechnol., 2005; 8: 113-120.
- Blevins J.S., Beenken K.E., Elasri M.O., Hurlburt B.K., Smeltzer M.S. Strain-dependent differences in the regulatory roles of sarA and agr in Staphylococcus aureus. Infect. Immun., 2002; 70: 470-480.
- Van Belkum A., Melles D.C. Not All Staphylococcus aureus strains are equally pathogenic. Discov. Med., 2009; 5: 148-152.
- Leibig M., Liebeke M., Mader D., Lalk M., Peschel A., Gotz F. Pyruvate formate lyase acts as a formate supplier for metabolic processes during anaerobiosis in Staphylococcus aureus. J. Bacteriol., 2011; 193: 952-962.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.