ISSN: 0973-7510
E-ISSN: 2581-690X
The microbiota of fish intestine is the most complex of all organs that interact closely with the immune system. This can be used as biological control agents that can control the disease caused by pathogenic bacteria. Thirty isolates were identified based on 16s rRNA gene sequencing. These bacteria were then tested for their pathogenicity and antagonistic activity. The bioassay was conducted by administering selected bacteria into the fish and inoculating with Aeromonas. Total 30 bacterial isolates were successfully isolated from the carp intestine. Morphology and DNA marker analysis shows wide diversity of bacterial consortium within the fish intestine. These bacterial community are dominated by Proteobacteria and Firmicutes. The pathogenicity test showed that 20 isolates were non-pathogenic at a density of 106 cfu/mL, while the rest were pathogenic to the fish. The antagonistic test showed that some isolates strongly or mildly inhibit Aeromonas and Vibrio and the rest weakly or do not inhibit the assayed bacteria. Two isolates (CgM8=Bacillus sp. and CgM37=Bacillus subtilis) are significantly better than control to protect the fish from Aeromonas infection. Two species of commensal bacteria originated from fish intestine are potential to be used as biological control agents against Aeromonas for common carp.
Aeromonas; Biological Control; Cyprinus carpio; Intestinal Bacteria, Bacillus subtilis
Common carp (Cyprinus carpio L.) is a freshwater fish that is widely cultivated in Indonesia. This fish has a rapid growth and high fecundity, so it has a high potential to become a source of protein. Cultivation of carp has quite a rapid development, but there are limitations in the cultivation of this fish. Like some other fish species, in the aquaculture sector, one of the limiting factors for its development is disease control1. Aeromonas is the causative agent of MAS (Motile Aeromonas Septicemia). It is either acute or subacute or even a chronic infectious disease of all freshwater fishes, characterized by rapidly fatal septicemia with a few gross signs, exophthalmia, ascitis and ulcer formation. The bacteria Aeromonas is a widely distributed pathogenic bacteria, and killed the fish up to 80-100% within 1-2 weeks2. Most cultured and wild fish are susceptible to infection with Aeromonas such as carp, channel catfish, eel, goldfish, snakehead fish, rainbow trout, brown trout and tilapia.
Control of bacterial diseases that have been done is to use antibiotics. But the use of this material creates a dependence on the provision of antibiotics, as well as increase production costs because the price is relatively high. The presence of antibiotics as one of the efforts to overcome diseases has many negative impacts, such as causing microbial resistance and producing residues that are harmful to humans who consume them. Therefore, it is important to search for other alternatives in controlling fish diseases more safely and effectively. The use of biological control agents, such as bacteria can be one of the alternatives. Bacteria that have been widely used as biological control agents in the field of aquaculture usually come from Lactic Acid Bacteria (LAB) or from the genus Bacillus, but these bacteria are mostly isolated from mammals or from terrestrial areas, not aquatic environments, therefore it renders these probiotics less effective when used for aquaculture2-4.
A fish’s intestine plays an important role on its health status, because the organ provides a micro-environment where there are numerous microbiota that interact with each other, and also interact with the host. The biological control agents which originate from the commensal bacteria within the fish itself are more advantageous than the bacteria from the outside because the bacteria have been proven to colonize and adapt to their host environment. Therefore, microbial communities of each aquatic species need to be identified to provide a more effective chance of developing new probiotics or new immunostimulants5
For different experimental purposes, the intestinal microbial flora of fish has been studied by several workers. These include microbial flora as food of fish6-7, microbial flora‘s role in production of enzymes8 and antibiotic resistance profile of indigenous flora9-10. The micro flora of reared fish has also been studied as a source of protection against diseases11-13 For all these reasons, study of intestinal bacterial flora is important.
Microbiota in fish intestines have been extensively studied, such as Oncorhynchus mykiss14, Garra mullya15, Ictalurus punctatus, Micropterus salmoides, and Lepomis macrochirus5. Bacterial communities of several carp species have also been studied, namely grass carp (Ctenopharyngodon idellus), crucian carp (Carassius cuvieri), and bighead carp (Hypophthalmichthys nobilis)16. While the potentials of bacteria as probiotics, immunostimulants and / or biological control agents have been investigated, such as Bacillus subtilis on Koi and Nila17-18, Lactobacillus on Flounder and Grouper19, Enterococcus faecalis for Snakehead fish20. In this study we have analyzed the intestinal bacterial of a Common carp. There are few studies that discuss intestinal bacteria, but studies exploring Common carp intestinal bacteria as a biological control agent for Aeromonas are even rarer.
The aim of this study was to isolate the Common carp’s intestinal bacteria, to study the morphological characterization and molecular identification of the isolated bacteria, to analyze the diversity of the bacteria based on the 16s rRNA gene sequence, and to explore its potential as a biological control agent for Aeromonas.
Collection of fish and Isolation of intestinal bacteria
Common carp was collected from Floating nets cage in Cirata reservoir, West Java, Indonesia. The fresh fish was rinsed with sterile aquadest and surface sterilized with ethanol (70%). The intestine were removed by dissection in sterile condition, then scraped the inner part and suspended in sterile saline and serially diluted in test tube. The presence of bacteria were checked by performing wet mount of the saline suspension. Bacterial suspension was then streaked on NA (Nutrient Agar) and MRS (de Man Rogosa Sharpe) agar media. The plates were incubated at 37oC for 24 h.
Molecular identification
Bacterial isolates were identified by 16s rRNA gene sequencing (Macrogen, Korea). All sequences were edited in BIOEDIT. Sequencing data were compared with DNA databases using the BLAST in EzTaxon (www.EzBioCloud.net). Neighbor-joining analysis was performed using MEGA7 software package. The confidence limits for the tree topology were estimated using 1000 x bootstrap analysis.
Pathogenicity and Antagonistic Test
Thirty bacterial isolates were tested to their pathogenicity test to fish survival. Antagonistic activity of the isolates were tested in-vitro against three bacterial pathogens (Aeromonas hydrophila, Staphylococcus epidermidis and Vibrio sp.)
Testing bacteria in fish (Induction and Challenge experiment)
A direct immersion method was done to deliver the bacteria as a biological control agent into the fish. The selected 3 isolates (8, 36 and 37) were administered by soaking the fish into the bacterial suspension over 30 minutes separately. Bacterial density was set at up at 106 cfu/ml before admistering into the fish and then maintaining the fish for 12 days to induce the immune system. During maintanance, the fish were feed twice daily with a standard fish pellet. Challenge test to see the effects of selected bacteria on the fish survival, a pathogenic strain of Aeromonas was inoculated into the fish tank at a density of 106 cfu/ml. The challenge test was performed over 4 days before counting the fish survival for each treatment.
Data Analysis
The challenge experiment data were analyzed using variance analysis (ANOVA) with Complete Randomized Design (Four treatments with three replicates). If the result of ANOVA showed a significant effect (p < 0.05), further test was carried out using Duncan’s Multiple Range Test (DMRT) with a confidence significance level of 5%.
Bacterial Characterization and Identification
Thirty bacterial strains were isolated from the intestine of Common Carp. Bacteria that grew on NA medium were dominated by Gram negative bacilli, while bacteria that grew on MRS medium were mostly Gram positive bacilli or cocci. Molecular identification of the intestinal bacteria based on 16s rRNA gene was performed. Phylogenetic tree was constructed from the 30 isolates. The phylogenetic analysis suggested that CgM36 isolate was closely related to Enterococcus faecalis (Fig. 1). The same analysis was also used to identify the other isolates (Table 1).
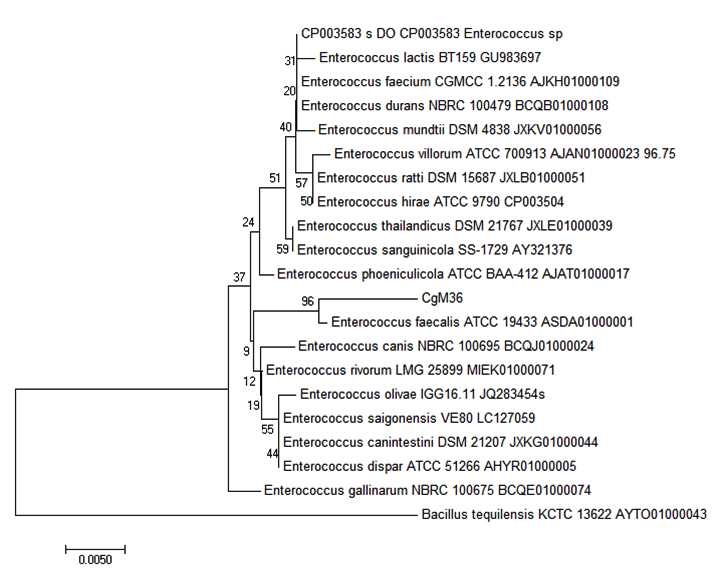
Fig. 1. Phylogenetic analysis of CgM36 isolate based on 16s rRNA gene sequence with MEGA 7
Table (1):
Molecular identify of intestinal bacteria of Common Carp based on 16s rRNA gene
No. |
Isolate |
Top-hit taxon |
Top-hit strain |
Similarity (%) |
|---|---|---|---|---|
1 |
CgN1 |
Enterobacter cloacae |
LMG 2683(T) |
92.48 |
2 |
CgN2 |
Bacillus flexus |
NBRC 15715(T) |
95.80 |
3 |
CgN3 |
Bacillus flexus |
NBRC 15715(T) |
95.50 |
4 |
CgN4 |
Bacillus cereus |
ATCC 14579(T) |
95.59 |
5 |
CgN5 |
Proteus mirabilis |
ATCC 29906(T) |
88.50 |
6 |
CgN6 |
Proteus mirabilis |
ATCC 29906(T) |
80.87 |
7 |
CgN9 |
Bacillus carboniphilus |
JCM 9731(T) |
68.48 |
8 |
CgN10 |
AF426002_s Niveispirillum sp. |
UNSW7 |
59.79 |
9 |
CgN12 |
FJ369991_s Blautia sp. |
TS55_a03b11 |
55.44 |
10 |
CgN13 |
Proteus mirabilis |
ATCC 29906(T) |
85.36 |
11 |
CgN14 |
Proteus mirabilis |
ATCC 29906(T) |
85.13 |
12 |
CgN15 |
Proteus mirabilis |
ATCC 29906(T) |
88.44 |
13 |
CgN16 |
Proteus mirabilis |
ATCC 29906(T) |
80.82 |
14 |
CgN17 |
Proteus mirabilis |
ATCC 29906(T) |
87.14 |
15 |
CgN18 |
Proteus mirabilis |
ATCC 29906(T) |
77.03 |
16 |
CgM1 |
Bacillus haynesii |
NRRL B-41327(T) |
99.79 |
17 |
CgM5 |
Bacillus licheniformis |
ATCC 14580(T) |
98.30 |
18 |
CgM6 |
Staphylococcus gallinarum |
ATCC 35539(T) |
99.41 |
19 |
CgM8 |
CP013984_s Bacillus sp. |
DE111 |
99.54 |
20 |
CgM15 |
Staphylococcus gallinarum |
ATCC 35539(T) |
99.55 |
21 |
CgM16 |
Lactococcus garvieae |
ATCC 49156(T) |
99.12 |
22 |
CgM18 |
Bacillus zhangzhouensis |
DW5-4(T) |
100.00 |
23 |
CgM19 |
Rummeliibacillus stabekisii |
KSC-SF6g(T) |
99.87 |
24 |
CgM22 |
CP013984_s Bacillus sp. |
DE111 |
98.57 |
25 |
CgM34 |
Bacillus tequilensis |
KCTC 13622(T) |
98.29 |
26 |
CgM36 |
Enterococcus faecalis |
ATCC 19433(T) |
98.22 |
27 |
CgM37 |
Bacillus subtilis |
KCTC 13429(T) |
99.34 |
28 |
CgM38 |
CP013984_s Bacillus sp. |
DE111 |
99.56 |
29 |
CgM19a |
Bacillus albus |
N35-10-2(T) |
96.70 |
30 |
CgM20a |
Bacillus megaterium |
NBRC 15308(T) |
95.21 |
Table 1 indicates that Firmicutes and Proteobacteria are dominantly found in the fish intestine. In this study, the NA culture medium (CgN) was dominated by Proteus. Molecular data showed, their proximity is close to Proteus mirabilis species. In addition, other genus were also found such as Enterobacter , Bacillus, as well as Niveispirillum and Blautia. In the MRS agar medium, cultured bacteria (CgM) which belong to the Lactic Acid Bacteria group were also obtained from Lactococcus and Enterococcus genera. Other generas were also recovered from MRS medium including Staphylococcus and Rumeliibacillus.
Pathogenicity Test
Before choosing as a biological control agent, the bacterial isolates were confirmed first for their pathogenic effects to the host. The fish model should be challenged with the bacterial isolates, under normal or stress conditions3. In this research, the fish was immersed in a suspension of the candidate bacteria. From the pathogenicity test, thirty isolates , show the varied survival rate as shown in figure 2.
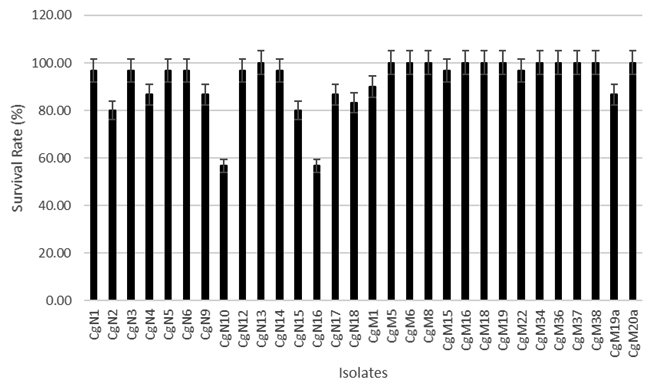
Fig. 2. The Survival Rate of Common carp after infected with bacterial isolates at a density of 106 cfu/mL
There were 20 non-pathogenic isolates at density of 106 cfu/mL, while the other 10 were pathogenic. Eight pathogenic isolates were obtained from NA medium (2, 4, 9,10, 15, 16,17, 18) and 2 isolates were obtained from MRS agar medium (1 and 19a).
Antagonistic Test
Bacterial antagonism occure in nature, therefore microbial interactions play a major role in the equilibrium between beneficial and potentially pathogenic microorganisms. However, the composition of microbial communities can be altered by any human activities including animal husbandry practices and environmental conditions that stimulate the proliferation of certain bacterial species. The microbiota in the gastrointestinal tract of aquatic animals are also influenced by aquatic microbes. Intestinal microbial manipulation constitutes a viable tool to reduce or eliminate the incidence of opportunist pathogens21.
In this study, thirty bacterial isolates were tested for their antagonisity against three pathogenic bacteria, Aeromonas and Vibrio (representative from pathogenic Gram negative bacteria group), and Staphylococcus (representative from pathoegenic Gram positive bacterial group). The results of the antagonistic test are shown in Figure 3.
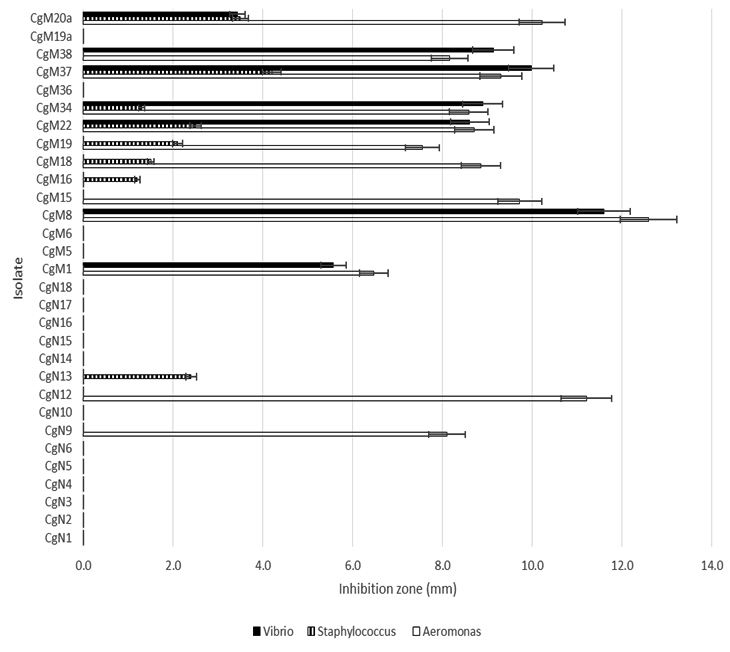
Fig. 3. The result of in vitro antagonism test of intestinal bacterial isolates against Aeromonas, Vibrio and Staphylococcus
Figure. 3 shows that there are 12 isolates of intestinal bacteria can inhibit Aeromonas and 7 bacterial isolates can inhibit Aeromonas and Vibrio. There is no isolate that only inhibits Vibrio without inhibiting Aeromonas. The data also show that 8 isolates can inhibit Staphylococcus and 4 isolates can inhibit all three bacteria.
Bioassay
From the results of bacterial identification, pathogenic and antagonistic tests, three of bacterial species were selected for further testing. The selected bacterial isolates were obtained from isolate CgM8 (Bacillus sp.), CgM36 (Enterococcus faecalis) and CgM37 (Bacillus subtilis). All three bacterial species were introduced to the fish, and incubated first for 12 days for the induction of the immune system. Afterwards, they were then challenged with Aeromonas. The highest percentage of survival rate of the fish was obtained from the treatment with isolate CgM37 (Figure. 4).
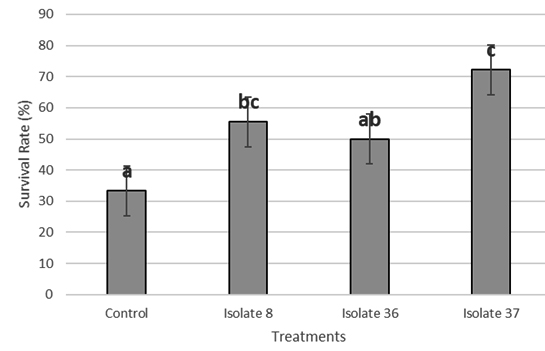
Fig. 4. The survival rate of Common carp treated with bacterial isolates CgM 8, 36, and 37
Media for the cultivation of microorganisms contain the substances necessary to support the growth of microorganisms. Due to the diversity of microorganisms and their diverse metabolic pathways, there are numerous media22. Nutrient agar (NA) is a medium for the isolation, cultivation and maintenance of a wide variety of microorganisms. NA is one of the most commonly used media in bacteriological procedures such as water testing, procedures run on food products or culture stock, for growth of samples in bacterial tests, and for isolating organisms in pure cultures. In this study, the NA culture (CgN) obtained was dominated by Proteus, although morphologically seen differently, but after molecular analysis, its proximity was similar to Proteus mirabilis species. Proteus is widely distributed in animals, polluted water, and is also found in the intestines of humans. In this research, this genus dominated intestinal microbial communities cultured with NA medium. The dominance of this species is in accordance with the opinion of Schaffer and Pearson23 which states that Proteus mirabilis is well-known in clinical laboratories and microbiology survey as the species that swarms across agar surfaces, overtaking any other species present in the process. P. mirabilis is often isolated from the gastrointestinal tract, although whether it is a commensal, a pathogen, or a transient organism, is somewhat controversial.
MRS agar is used for the isolation, cultivation, and maintenance of Lactobacillus species from clinical specimens, foods, and dairy products. It is a good medium for the cultivation of lactic acid bacteria. MRS agar contains polysorbate, acetate, magnesium, and manganese that act as growth factors for Lactobacillus. In order to act as medium, MRS is enriched by various nutrients to support the growth of lactic acid bacteria, but not very selectively, so it still allows other types of bacteria to grow.
Table 1 indicates that Proteobacteria and Firmicutes are dominantly found as the common phyla. The results of this study are similar to those found by other studies. Proteobacteria, Firmicutes and Actinobacteria were the dominant allochthonous microbiota in the gut content of grass carp cultured in pond24 while Luo et al. 25 identicated Proteobacteria, Firmicutes, Bacteroides and Actinobacteria as the dominant allochthonous bacteria in the intestine of transgenic carp. Al Harbi and Udin26 studied that many of the bacteria found in Tilapia intestine were predominantly from proteobacteria and firmicutes. In the rainbow trout intestine, Kim and Austin14 found that Aeromonadaceae, Enterobacteriaceae and Pseudomonadaceae representatives were the dominant cultured bacteria.
In zebrafish, the morphologically different bacteria found were Aeromonas, Vibrio, Photobacterium, Pseudomonas, Comamonas, Ochrobactrum, and Staphylococcus27. One prominent difference found in the present study compared to the study of Roeselers et al.28 was that Firmicutes represented the second most abundant phylum. Firmicutes represents one of the most abundant phyla in the mammalian intestine, and members of Clostridia within Firmicutes are obligate anaerobes29.
Perez et al.30 stated that the intestinal microbiota of freshwater species tend to be dominated by members of the genera Acinetobacter, Aeromonas, Flavobacterium, Lactococcus and Pseudomonas, representatives of the family Enterobacteriaceae, and obligate anaerobic bacteria of the genera Bacteroides, Clostridium, and Fusobacterium. The intestinal content in grass carp, crucian carp, and bighead carp were dominated by four major phyla, including Fusobacteria, Firmicutes, Proteobacteria and Bacteroidetes16.
The result on the pathogenicity test shows that there are 20 non-pathogenic isolates at the density of 106 cfu/mL, while the other isolates are pathogenic. In this study, there are several isolates that are pathogenic when tested. The type of bacteria that are pathogenic include Bacillus and Proteus. In general, bacterial pathogens are caused by their ability to produce endotoxins and exotoxins, such as LPS, and their ability to produce certain enzymes or proteins that can damage the immune system in fish. In addition, some bacteria have the ability to lyse cells or blood hosts, so that the tissue or organ hosts become damaged31.
The antibacterial effect of bacteria is due to the production of antibiotics, bacteriocins, siderophores, enzyme and/or hydrogen peroxide and the alteration of pH values by the production of organic acid2. The inhibition of pathogenic bacteria occurs by destroying the cell wall causing lysis or inhibiting cell wall growth in the growing bacterial cell, altering the permeability of the cytoplasmic membrane, causing leakage and resulting in nutrients exiting the cell. In addition, antagonistic bacteria can also inhibit the synthesis of proteins and nucleic acids by denaturing proteins and destroying nucleic acids so that their function as genetic material is lost. Then it also inhibits intracellular enzyme activity that disrupts cell metabolism32. Antibacterial compounds can also increase lysozyme activity, which can be used as immunogenic parameters. The lysozyme enzyme works by lysis of bacterial cell walls such as hydrolyzing N-acetylglucosamine and N-acetylmuramic acids in peptidoglycan, so that with the loss of cell walls, the bacteria die33. Lyszozymes are more active in Gram-positive bacteria than in Gram-negative bacteria, because the peptidoglycan content in its cells more abundant34.
Based on this study, cultured bacterial isolates have varying abilities in inhibiting pathogenic bacteria, (Aeromonas, Vibrio and Staphylococcus), seen from clear zones formed around disc paper (figure 3). The clear zone is caused by bacterial isolates to produce metabolites which are antibacterial compounds that can inhibit the development of pathogenic bacteria. Antibacterial compounds work by disrupting components of the cell wall, causing plasmolysis resulting in inhibition of growth or death of pathogenic bacteria35. The ability of bacterial isolates in inhibiting the growth of pathogenic microbes is one indicator that the isolate has potential as a biological control agent. From the results of antagonistic test of cultured bacteria showed that not all isolates have the ability to inhibit three pathogenic bacteria (Aeromonas, Vibrio, Staphylococcus).
The results of the antagonistic test of intestinal bacteria cultured on Aeromonas showed that the isolates grown on NA medium which had antagonistic activity in this study were isolates CgN 9 and 12, while isolates grown on MRS medium which had antagonistic activity, were isolates CgM 1, 8, 15, 18, 19, 22, 34, 37, 38, and 20a. The isolates produced clear zones ranging from 6.5 to 12.6 mm. Pan et al.36 states that the clear zone diameter 0-3 mm has a weak inhibition, the 3-6 mm clear zone diameter has good inhibition, and the clear zone diameter of > 6 mm has strong inhibition. Therefore, all clear zones formed in inhibition of Aeromonas are included in the strong inhibition.
The result of the antagonistic test of intestinal bacteria on Vibrio pathogen showed that the bacterial isolates had a moderate to strong inhibition. The isolates of CgM 1 and 20a resulted in moderate inhibition, with clear zones ranging from 3.4 to 5.6, whereas CgM 8, 22, 34, 37 and 38 isolates had strong inhibitory powers, resulting in clear zones ranging from 8.6 – 11.6 mm.
The results of the antagonistic test of intestinal bacteria against Staphylococcus showed that the bacterial isolates had a weak to moderate inhibition. Isolate CgN 13 produced a weak inhibition, with a clear zone of 2.4 mm. Isolates CgM 16, 18, 19, 22, and 34 also had weak inhibition, ranging from 1.2 to 2.5. While the isolates CgM 37 and 20a had a moderate inhibition, which produced a clear zone of 4.2 and 3.5 mm.
Differences in intestinal bacterial inhibition to the pathogenic bacteria tested were influenced by the differences in the ability of these bacteria to produce antibacterial compounds, as well as the thickness and composition of pathogenic bacterias’cell wall.
Bioassay was performed using three selected intestinal bacteria, which were isolate CgM8 (Bacillus sp.), CgM36 (Enterococcus faecalis) and CgM37 (Bacillus subtilis). Isolate CgM8 (Bacillus sp.) produced the highest inhibition against Aeromonas and Vibrio pathogens, then CgM36 (Enterococcus faecalis), which gave the best inhibitory ability against three pathogenic bacteria and CgM37 (Bacillus subtilis) produced very weak clear zones in antagonistic tests. The statistical test of the survival of fish on bioassay proved that the two isolates gave significantly different results than controls, proving that these isolates could act as biological control agents, although further studies are still needed.
These bacteria are normal intestinal microflora in common carp. If these species are common in this fish, this findingis not in line with the study of Wu et al.37 who states that lactic acid bacteria can not build large populations in the intestine of grass carp. The results of this study are in line with the statement of Ghosh et al.38 that LAB isolated from the gut of M. cephalus provides an opportunity to develop a sustainable and organic means of combating the aquaculture pathogens. It has been demonstrated by Balcázar39 that a mixture of bacterial strains (Bacillus and Vibrio sp.) had a beneficial effect on the growth and survival of juveniles of white shrimp besides improving their immunity against Vibrio harveyi and white spot syndrome virus. Most probiotics proposed as biological control agents in aquaculture belong to lactic acid bacteria and the genus Bacillus, although other genera can also be included2.
Probiotics or immunostimulant bacteria as biological control agents in aquaculture have several possible modes of action. Those modes are: production of inhibitory compounds; competition for chemicals or available energy; competition for adhesion sites; enhancement of the immune response; improvement of water quality; interaction with phytoplankton; source of macro and micronutrients; and enzymatic contribution to digestion2.
Some bacteria have been used as immunostimulants, they are susceptible to phagocytosis by macrophages and they stimulate cytokine synthesis. So their immunostimulant effect is due to the release of a mixture of cytokines40. Among the cytokine synthesis enhancers, the most effective is bacille Calmette-Guerin (BCG). A live and attenuated vaccine from Mycobacterium bovis BCG strain results in elevated B-cell and T-cell mediated response, enhancing phagocytosis and providing resistance to infection41.
Immunostimulatory bacteria have a major role in inducing innate immune cells to stimulate and modulate the mucosal immune system by decreasing the production of proinflamatory cytokines through NFºB pathways, increasing the production of anti-inflammatory cytokines (IL-10), increasing IgA defenses and affecting the maturation of dendritic cells42 . Some bacterial products with immunomodulatory properties such as lipopolysaccharide (LPS), peptidoglycan and lipoteichoic acid (LTA) from Bifidobacteria may activate macrophages to evoke an immune response. The bacteria can also increase the production of anti-inflammatory cytokines and reduce the production of proinflammatory cytokines thereby reinforcing the intestinal mucosal barrier43.
ACKNOWLEDGMENTS
Acknowledgments are addressed to the Ministry of Research, Technology and Higher Education of the Republic of Indonesia through the Post-Graduate Scholarship to provide funding and support for this research.
- FDA. 2012. https://www.centerwatch.com/drug-information/fda-approved-drugs/year/2012
- Cipriano, R.C., G.L. Bullock, and S.W. Pyle. 1984. Aeromonas hydrophila and Motile Aeromonad Septicemias of Fish. US Fish & Wildlife Publications. 134. http://digitalcommons.unl.edu/usfwspubs/134
- Verschuere, L. G. Rombaut, P. Surgeloos and W. Verstraete. Probiotic Bacteria as Biological Control Agents in Aquaculture. Microbiology and Molecular Biology Reviews. 2002; 64(4): 655-671.
- Ran, C., A. Carrias, M.A. Williams, N. Capps, C.T. Bui, J.C Newton, E.L.Oi, C.L.Browdy, J.S.Terhune, and M.R.Liles. Identification of Bacillus Strains for Biological Control of Catfish Pathogens. Journal Plos ONE. 2012; 7(9): e45793.
- Wang YB, Li JR, Lin J. Probiotics in aquaculture: challenges and outlook. Aquaculture, 2008; 281: 1–4
- Kamjunke, N., R. Mendonca, I. Hardewing and T. Mehner. Assimilation of different cyanobacteria as food and the consequences for internal energy stores of juvenile roach. J. Fish Biology. 2002; 60(3): 731-738pp
- Askarian, F., Z. Zhou., R. E. Olsen, S. Sperstad, and E. Ringø. Culturable Autochthonous Gut Bacteria in Atlantic Salmon (Salmo salar 3 L.) Fed Diets with or without Chitin. Characterization by 16S rRNA Gene Sequencing, Ability to Produce Enzymes and in vitro Growth Inhibition of Four Fish Pathogens. Journal Aquaculture 2009.
- Maslowski, K. M dan C.R. Mackay. Diet, Gut Microbiota and Immune Responses. Nature Immunology. 2011; 12(1)
- Hooper, L.V., D. R. Littman and A. J. Macpherson. Interactions Beetwen the Microbiota and the Immune System. The Gut Microbiota., 2012; 336: 1268-1273.
- Galdeano, M dan G. Perdigo. The Probiotic Bacterium Lactobacillus casei Induces Activation of the Gut Mucosal Immune System through Innate Immunity. American Society for Microbiology. 2006; 13(2): 219-226.
- Gaskins, H. R., J. A. Croix, N. Nakamura dan G. M. Nava. Impact of the Intestinal Microbiota on the Development of Mucosal Defense. Clinical Infectious Diseases. 2008; 46(2): 80-86.
- Guarner, F and J. Malagelada. Gut Flora in Health and Disease. The Lancet 2003; 361 no 9356: 512–519.
- Sissons, J.W. Potential of Probiotic Organism to Prevent and Promote Digestion in Farm Animals. Journal of The Science of Food and Agriculture. 1989; 49(1): 1-13pp.
- Kim, D.H., J.Brunt, B.Austin. Microbial Diversity of Intestinal Contents and Mucus in Rainbow Trout (Oncorhynchus mykiss). Journal of Applied Microbiology. 2006; 102: 1654-1664.
- Rudresh, B.S., N. Dahanukar, G.M Watve, and N.S. renukaswamy. Microbial Gut Flora Of A Freshwater Fish Garra Mullya (Sykes) From Mutha River, Northern Western Ghats, India. Ecological Society., 2010; 17: 53-57.
- Li, T., M.Long, F-J. Gatesoupe, O. Zhang, A. Li, X. Gong. Comparative Analysis of the Intestinal Bacterial Communities in Different Species of Carp by Pyrosequencing. Mocrobial Ecology. 2014; 69(1): 25-36.
- He, S., W. Liu, Z. Zhou, W. Mao, P. Ren, et al. Evaluation of probiotic strain Bacillus subtilis C-3102 as a feed supplement for koi carp (Cyprinus carpio). Journal of Aquaculture Research and Development. 2011; 5(1): 1-7pp.
- Elkamel, A.A and G.M. Mosaad. Immunumodulation of Nile Tilapia, Orechromis niloticus by Nigella sativa and Bacillus subtilis. Journal Aquaculture. 2012; 3(6).
- Cordero, H., M.A. Esteban, A. Cuesta. Use of Probiotic Bacteria against Bacterial and Viral Infections in Shellfish and Fish Aquaculture. Agricultural and Biological Science. 2014; 1224. No 51: 953-978.
- Allameh, S.K., E. Ringo, F.M. Yusoff, H. M. Daud, A.Ideris. Properties of Enterococcus faecalis, a new probiotic bacterium isolated from the intestine of snakehead fish (Channa striatus Bloch). Afr. Journal of Microbiology Research. 2014; 8(22): 2215-2222.
- Balcazar, J.L., I. de Blas, I. Ruiz-Zarzuela, D. Cunningham, D. Vendrell, J.L. Muzquiz. 2006. The Role of Probiotics in Aquaculture, Veterinary Microbiology. Elsevier. Vol. 114 : 172-286pp.
- Atlas, R.M. Handbook of Microbiological Media. 4 ed. CRC Press, Boca Raton, FL 2010.
- Schaffer, J. N and M. M. Pearson. Proteus mirabilis and Urinary Tract Infections. Microbiology Spectrum. 2015; 3(5):
- Han, S., Y. Liu, Z. Zhou, S. He, Y. Cao, P. Shi, B. Yao and E. Ring. Analysis of Bacterial Diversity in the Intestine of Grass Carp (Ctenopharyngodon idellus) Based on 16S rDNA Gene Sequences. Journal of Aquaculture Research and Development. 2010; 42: 47-56pp.
- Luo, X., Q. Yan, S. Xie, W. Hu, Y. Yu, Z. Hu. Gut Microbiota Contributes to the Growth of Fast-Growing Transgenic Common Carp (Cyprinus carpio L.). 2001; 8(5): e64577
- Al Harbi, A and N. Udin. Bacterial Diversity of Tilapia (Oreochromis niloticus) Cultured in Brakish Water in Saudi Arabia. Journal Aquaculture. 2005; 566-572
- Cantas L, Sørby JRT, Aleström P, Sørum H. Culturable gut microbiota diversity in Zebrafish. Zebrafish. 2012; 9(1):26–37.
- Roeselers, G., E. K. Mittge, W. Z. Stephens, D. M. Parichy, C. M. Cavanaugh, K. Guillemin dan J. F. Rawls. Evidence for a Core Gut microbiota in the zebrafish. The ISME Journal 2011; 5: 1595–1608pp.
- Lan, C.C. and D. R. Love. Molecular Characterization of Bacterial Community Structure along the Intestinal Tract of Zebrafish (Danio rerio). International Scholarly Research Network Microbiology. 2011; 2012: 10pp
- Perez, T., Jl. Balcazar, I. Ruiz-Zarzauela, N. Halaihel, D. Vendrell, I. de Blas and J.L. Muzquiz. Host Microbiota Interactions within the Fish. Mucosal Immunology 2010.
- Sarkar, M.J.A and M.M. Rashid. 3030 Pathogenicity of the bacterial isolate Aeromonas hydrophila to catfishes, carps and perch. J. Bangladesh Agril. 2012; 10(1): 157–161
- Pelczar, J. M and E. C. S. Chan. Elements of Microbiology. McGraw-Hill Inc., US 1981.
- Paulsen S.M., H. Lunde , R.E Engstad and B. Robertsen. In vivo effects of ²-glucan and LPS on regulation of lysozyme activity and MRNA expression in atlantic salmon (Salmo salar L.). Fish And Shellfish Immunology. 2003; 14(1): 39-54pp
- Ibrahim, H.R., T. Matsuzaki and T. Aoki. Genetic evidence that antibacterial activity of lysozyme is independent of its catalytic function. FEBS Lett. 2001; 506: 27–32pp
- Brooks, G. F.., K. C. Caroll, J. S. Butel, S. A. Morse, T. A. Mietzner. Jawetz, Melnick, & Adelberg’s Medical Microbiology. 26th Edition. The McGraw-Hill Companies, Inc 2013.
- Pan, X., F. Chen, T. Wu, H. Tang, and Z. Zhao. The acid, Bile Tolerance and Antimicrobial property of Lactobacillus acidophilus NIT. J. Food Control. 2009; 20: 598- 602.pp
- Wu, S., G. Wang, E. R. Angert, W. Wang, W. Li dan H. Zou. Composition, Diversity, and Origin of the Bacterial Community in Grass Carp Intestine. 2012; 7(2): 230-440
- Ghosh, S., RingØ, E., Selvam, D.G., Rahiman, K.M., Sathyan, N., John, N., Hatha, A.A.M., Gut aassociated lactic acid bacteria isolated from the estuarine Fish Mugil cephalus: molecular diversity and antibacterial activities against pathogens. Int. J.Aquacult. 2014; 4: 1-11.
- Balcázar J.L., Evaluation of probiotic bacterial strains in Litopenaeus vannamei. Final report, National Center for Marine and Aquaculture Research, Guayaquil, Ecuador 2003.
- Galeotti M. Some aspects of the application of immunostimulants and a critical review of methods for their evaluation. J ApplIchthyol, 1998; 14: 189-199.
- Tizard I., Veterinary immunology: an introduction. W.B. Saunders, Philadelphia. 1996; Chapter 35, pp. 475-478.
- Sahalan, A. Z..H. A.A. Abdul, H.L. Hing, and A.G M. Kamel. Divalent cations (Mg2+, Ca2+) protect bacterial outer membrane damage by polymyxin. Sains Malaysiana. 2013; 42(3) : 301–306pp
- Rao, R.K and G. Samak. Protection and restitution of gut barrier by probiotics nutritional and clinical implications. National Institutes of Health. 2013; 9(2) : 99–107.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


