ISSN: 0973-7510
E-ISSN: 2581-690X
Thirty one wheat germplasm lines were screened under natural epiphytotic conditions against stripe rust at University Research Farm, Chatha, during Rabi, 2014-15. On the basis of final rust severity (FRS), area under rust progress curve (AURPC) and coefficient of infection (CI), eight lines (Raj 4037, Sonara 64, NP 823, HPW 42, K9351, NIAW 301, PBW 12, and HUW 213) exhibited partial resistance to the disease while as on the basis of infection rate (r) six lines (NP 825, HP 1633, K8434, PBW 12, Ajanta and K9533) showed partial resistance to the disease. Field response of these lines against stripe rust showed that 14 genotypes (Sonara 64, Utkalia, NI 5439, NIAW 301, PBW 12, HUW 213, Ajanta, NP 823, K8434, K9533, Sharbati Sonara, Raj 4037, HP 1633, HPW 42 and K9351) were moderately resistant, 16 (Tawa, KRL, RW 346, HD 2643, HS 1097, NP 825, WH 291, HUW 12, PBW 226, NI 179, NI 5439, K9644, HD1925, PBW 65, PV 18 and GW 503) were moderately susceptible and one genotype (Agra Local) was susceptible. Assessment of losses was also calculated at different growth stages and it was observed that losses at flowering stage and dough stage were highest in one line (Agra Local). Nine wheat germplasm lines (Sonara 64, K9351, HP 1633, Raj 4037, Sharbati Sonara, K9533, K8434, NP 823 and Ajanta) amplifying a band of 523 bp fragment indicating presence of Yr18 gene.
Stripe rust, Severity, Allele specific marker, Yr18, FRS, AURPC, CI, r, Cssfr2.
India is a privileged country to attain and retain the status of being the second largest producer of wheat and registering the historic production of 93.50 million tonnes during 2015-16. Among the three rust, stripe or yellow rust caused by Puccinia striiformis Westend f. sp. tritici is devastating foliar disease and is considered of immense importance in successful cultivation of wheat (Singh et al., 2014). Year after year the susceptible wheat cultivars that suffer from stripe rust disease result in increased inoculum build up thus posing major threat to wheat cultivation. Although remarkable progress has been made in breeding for stripe rust resistant varieties but the subsequent evolution of pathogen races at much greater pace continues to challenge this breeding programme (Singh et al., 2011) and stripe rust remains a threat to wheat cultivation (Sareen et al., 2012). Although the timely application of fungicides against this obligate parasite can manage the disease to some extent but their use add to the production costs. Breeding for resistance remains the most effective and efficient management strategy as it does not add input costs to farmers and is environmentally safe (Yang and Liu, 2004). The identification and knowledge of the resistance genes in commonly used parental germplasm and released cultivars is very important for utilizing the genetic resistance to manage this rust in full potential. The long term and economical strategy could thus be resistance breeding through deployment of effective rust resistance genes over space and time (Pretorius et al., 1997; Zeng et al., 2014). The genes expressing at adult plant stage have special significance because the cultivars having such genes have shown partial resistance that has remained effective for longer durations (Singh and Rajaram, 1991; Khan and Saini, 2009).
The outbreak of stripe rust in temperate areas of Jammu and Kashmir State, India is a matter of great concern owing to the fact that the rust inoculum generated in these areas may act as a reservoir of inoculum for rust initiation in North-West Plain Zones of the country. Therefore there is urgent need to curtail the pathway of rust pathogen in Jammu and Kashmir State on priority basis so that the same may not spread to food bowl of the country thereby causing a great threat to the National Food Security Mission (NFSM). With this background information the present study was undertaken to screen the wheat germplasm for identifying stripe rust resistant genotypes and then validate resistance gene (Yr18) at molecular level.
The present studies were carried out in the Division of Plant Pathology, Faculty of Agriculture, SKUAST Jammu, Chatha, during rabi 2014-2015.
Screening of wheat germplasm for slow rusting traits
Thirty one wheat genotypes viz., Sonara 64, Utkalia, NI 5439, NIAW 301, PBW 12, HUW 213, Ajanta, NP 823, K8434, K9533, Sharbati Sonara, RAJ 4037, HP 1633, HPW 42 , K9351, Tawa, KRL, RW 346, HD 2643, HS 1097, NP 825, WH 291, HUW 12, PBW 226, NI 179, K9644, HD1925, PBW 65, PV 18, GW 5031 and Agra Local) were used for screening for stripe rust disease under natural epiphytotic conditions during rabi 2014-2015 at University Research Farm, Chatha (32o 43’ N, 74o 54’ E). Each collected genotype was grown in three rows 2m apart with spacing of 22.5cm in third week of November, 2014. The entire experimental field was surrounded by the susceptible genotype Agra Local. The observations regarding per cent stripe rust severity were recorded from 29th Jan. 2015, onward till the crop was harvested. The modified Cobb’s scale which gives combined scores for level of severity and infection type was adopted to record the disease severity. Observations for slow rusting traits such as Final rust severity (FRS), Area Under Rust Progress Curve (Milus and Line, 1986), Coefficient of infection and Infection Rate (Brorers, 1989) were recorded.
Molecular validation of stripe rust resistance gene in wheat germplasm
Nineteen wheat genotypes (Sonara 64, K 9351, HP 1633, Raj 4037, Sharbati Sonara, K 9533, K 8434, NP 823, Ajanta, PBW 12, KRL, RW 346, HD 2643, HS 1097, NP 825, PBW 226, NIAW 301, PBW 343, and NI 179) were selected on the basis of low disease severity under field conditions for molecular validation of stripe rust resistance gene Yr18 by using allele-specific markers Cssfr2 (F=TTGATGAAACCAGTTTTTTTTCTA R=TATGCCATTTAACATAATCATGAA). The genomic DNA of selected genotypes was isolated by CTAB (cetyl-trimethyl ammonium bromide) method (Doyle and Doyle, 1987). Polymerase chain reaction (PCR) ampliûcation was conducted by Cssfr2 primer with temperature proûles as described by Laugdah et al. (2009). The ampliûcation products were separated on 3 per cent agarose gels containing ethidium bromide and 1× TBE buûer. The gels were visualized using gel documentation system for documentation of allele type in selected genotypes for resistance gene Yr18 using a standard molecular ladder of 100 bp.
Assessment of losses caused by stripe rust
Assessment of per cent losses due to the stripe rust in case of thirty one test genotypes viz., Tawa, KRL, Raj 4037, RW 346, SONARA 64, Utkalia, HD 2643, HS 1097, NP 825, WH 291, HP 1633, HUW 12, PBW 226, NP 823, PV 18, K 8434, NI 179, HPW 42, NI 5439, K9644, K9351, HD1925, NIAW 301, PBW 12, PBW 65, HUW 213, Ajanta, K9533, GW 503, Sharbati Sonara and Agra Local, were estimated by working out the per cent losses at different phonological stages of wheat crop, such as tillering, flowering and milking Mundy’s equation (1973) as : Loss = (0.44 x disease severity) + 3.15
Screening of wheat germplasm for slow rusting traits
During the present studies thirty-one genotypes of wheat were screened for their resistance and susceptibility response against stripe rust disease under subtropical agro climatic conditions of Jammu. The disease was first observed on 4th Feb., on a susceptible host Agra Local. Subsequently, the disease was recorded in other tested genotypes as well, but it was on 25th Feb. that the disease engulfed all the test genotypes and the disease severity ranged from 5-25 per cent. Thereafter the disease severity increased at a steady pace till 25th Mar. which was considered as the Final Rust Severity (FRS). The FRS ranged from 25 to 95 per cent, with maximum disease severity (95%) in Agra Local and minimum of 25 per cent each in K9351, NIAW301, PBW 12 and HUW213 (Table 1). Bariana et al. (2002) screened 176 doubled haploid lines derived from CD87/Katepwa (CD/K) for stripe rust resistance. Both parental lines, CD87 and Katepwa, showed stripe rust resistance. Both lines were susceptible to 110 E143A+ at seedling stage. The presence of susceptible segregates indicated the genetic independence of resistance in CD87 and Katepwa. Data in Table 1 further exhibited that AURPC value in 9 genotypes (Raj 4037, Sonara 64, HS 1097, NP 823, HPW 42, K 9351, NIAW 301, PBW 12 and HUW 213) ranged from 0-800 and in 21 genotypes (Tawa, KRL, RW 346, Utkalia, HD 2643, NP 825, WH 291, HP 1633, HUW 12, PBW 226, PV 18, K 8434, NI 179, NI 5439, K 9644, HD1925, PBW 65, Ajanta, K 9533, GW 503 and Sharbati Sonara) from >800 to 1600, however, in Agra Local it was highest (2340). While calculating the CI, a value of 6.25 to 8.25 was observed in Sonara 64, NIAW 301, PBW 12, HUW 213, Ajanta, NP 823, K8434, K9533, Sharbati Sonara, HP 1633, HPW 42, K9351 and Raj 4037 and 30 to 33.75 in Tawa, KRL, RW 346, Utkalia, HD2643, HS 1097, NP 825, WH 291, PBW 226, PV 18, NI 179, NI 5439, K9644, HD 1925, PBW 65 and GW 503 , whereas, in Agra Local the CI value (95) was the highest. Similarly, the infection rate (r) varied from 0.014 in case of NP 825 to 0.122 in Agra Local. Among them six germplasm (NP 825, HP 1633, K8434, PBW 12, Ajanta and K9533) exhibited a range of 0.014-0.019, whereas, in eight genotypes (KRL, NP 823, HD1925, NIAW 301, PBW 65, HUW 213, GW 503, and Sharbati Sonara) the infection rate ranged from 0.021 to 0.028. In case of TAWA, RW 346, Sonara 64, Utkalia, HD 2643, WH 291, HUW 12, PBW 226, PV 18, NI 179, HPW 42, NI 5439, K9644, and K9351 the infection rate ranged from 0.036 to 0.049. However, the infection rate in case of HS 1097, Raj 4037 and Agra Local was comparatively high (0.056, 0.084 and 0.122, respectively). The result was in conformity with the findings of other workers who reported that breeding lines with low value of AURPC, CI and r were expected to possessed genes that conferred partial resistance (Ali etal.,2012). Data in Table 2 indicate that genotypes were categorized as moderately resistant (Sonara 64, Utkalia, NI 5439, NIAW 301, PBW 12, HUW 213, Ajanta, NP 823, K8434, K9533, Sharbati Sonara, Raj 4037, HP 1633, HPW 42 and K9351), moderately susceptible (Tawa, KRL, RW 346, HD 2643, HS 1097, NP 825, , WH 291, HUW 12, PBW 226, NI 179, NI 5439, K9644, HD1925, PBW 65, PV 18 and GW 503) and susceptible (Agra Local), whereas, none of the tested genotypes showed resistant response against stripe rust
Molecular validation of stripe rust resistance gene in wheat germplasm:
The Yr18 locus confers partial and durable adult plant resistance against stripe rust fungus. In order to track Yr18, PCR was carried out with an allele specific primer cssfr2 which resulted in the amplification of 523 bp fragment, which exhibited the presence and absence of banding pattern. Nine wheat genotypes (Sonara 64, K9351, HP 1633, Raj 4037, Sharbati Sonara, K9533, K8434, NP 823 and Ajanta) amplified a band of 523 bp fragments which indicated the presence of Yr18 gene (Table 3, Fig., 1). Five allele-specific markers (cssfr1–cssfr5) were developed by Lagudah (2009) based on a 3 bp deletion in exon 11 of the Yr18-gene which were closely linked to the Lr34/Yr18/Ltn1/Pm38 locus and have been shown to provide a much wider diagnostic ability for this multi-pathogen resistance trait in diverse wheat cultivars. The validation of results are in accordance with the findings of Laugdah et al. (2009).
Assessment of losses caused by stripe rust
While assessing the losses while evaluating thirty one genotypes at tillering, flowering and dough phonological stages, though there was no loss at the initial stage of tillering, but the per cent losses at flowering stage ranged from 11.95 to 36.15. In case of Raj 4037, NIAW 301, PBW 12 and HUW 213 the per cent losses estimated were 11.95, whereas, the per cent losses estimated were 20.75 (HD 2643 and NP 825), 22.95 (PBW 65) and 36.15 (Agra Local). Similarly, at the dough stage the per cent losses estimated in K9351, NIAW 301, PBW 12 and HUW 213 was 14.15, whereas, in case of PBW 65 and Agra Local the losses recorded were 25.15 and 44.95 per cent, respectively. Chen (2005) have reported that the disease may cause 40-78 per cent losses under normal conditions, but if the conditions were favorable for disease spread the loss might rise to 84 per cent (Murray et al., 1994). The predicted losses may be less if the wheat varieties were resistant or slow rusting against rust diseases (Salman et al., 2006). Similarly, the wheat varieties which showed less area under disease progress curve normally bear less yield losses (Hussain et al., 1996).
Table (1):
Evaluation of wheat germplasm for slow rusting parameters against Puccinia striiformis.
S. No. |
Germplasm |
Final Rust Severity (FRS) |
Area Under Rust Progress Curve (AURPC) |
Coefficient of Infection (CI) |
Infection rate (r) |
|---|---|---|---|---|---|
01 |
Tawa |
40 |
1040 |
30.00 |
0.036 |
02 |
KRL |
40 |
960 |
30.00 |
0.025 |
03 |
RAJ 4037 |
30 |
604 |
07.50 |
0.084 |
04 |
RW 346 |
40 |
882 |
30.00 |
0.036 |
05 |
Sonara 64 |
30 |
606 |
07.50 |
0.049 |
06 |
Utkalia |
40 |
842 |
30.00 |
0.036 |
07 |
HD 2643 |
45 |
984 |
33.75 |
0.039 |
08 |
HS 1097 |
40 |
644 |
30.00 |
0.056 |
09 |
NP 825 |
45 |
1300 |
33.75 |
0.014 |
10 |
WH 291 |
45 |
1020 |
33.75 |
0.039 |
11 |
HP 1633 |
30 |
1020 |
07.50 |
0.017 |
12 |
HUW 12 |
40 |
922 |
30.00 |
0.036 |
13 |
PBW 226 |
40 |
962 |
30.00 |
0.036 |
14 |
NP 823 |
30 |
726 |
07.50 |
0.028 |
15 |
PV 18 |
40 |
926 |
30.00 |
0.036 |
16 |
K8434 |
30 |
924 |
07.50 |
0.017 |
17 |
NI 179 |
45 |
864 |
33.75 |
0.039 |
18 |
HPW 42 |
30 |
644 |
07.50 |
0.049 |
19 |
NI 5439 |
45 |
902 |
33.75 |
0.039 |
20 |
K9644 |
40 |
886 |
30.00 |
0.036 |
21 |
K9351 |
25 |
626 |
06.25 |
0.044 |
22 |
HD 1925 |
45 |
984 |
33.75 |
0.028 |
23 |
NIAW 301 |
25 |
648 |
06.25 |
0.024 |
24 |
PBW 12 |
25 |
758 |
06.25 |
0.019 |
25 |
PBW 65 |
50 |
1362 |
37.50 |
0.023 |
26 |
HUW 213 |
25 |
666 |
06.25 |
0.024 |
27 |
Ajanta |
30 |
978 |
07.50 |
0.017 |
28 |
K9533 |
30 |
820 |
07.50 |
0.017 |
29 |
GW 503 |
45 |
1022 |
33.75 |
0.028 |
30 |
Sharbati sonara |
35 |
1046 |
08.25 |
0.021 |
31 |
Agra Local |
95 |
2340 |
95.00 |
0.122 |
Table (2):
Response of wheat germ plasm against stripe rust under field conditions.
S.No. |
Disease response |
Genotype |
|---|---|---|
1 |
Immune |
Nil |
2 |
Nearly immune |
Nil |
3 |
Resistant (R) |
Nil |
4 |
Moderately resistant (MR) |
SONARA 64Utkalia NI 5439 NIAW 301 PBW 12 HUW 213 Ajanta NP 823 K8434 K9533 Sharbati Sonara Raj 4037 HP 1633 HPW 42 and K9351 |
5 |
Moderately susceptible (S) |
Tawa KRL RW 346 HD 2643 HS 1097 NP 825 WH 291 HUW 12 PBW 226 NI 179 NI 5439 K9644 HD1925 PBW 65 PV 18 and GW 503 |
6 |
Susceptible |
Agra Local |
Table (3):
Molecular validation of Yr18 for yellow rust.
S. No. |
List of Wheat Germplasm lines |
Yr18 (+/-) |
|---|---|---|
1. |
Sonara 64 |
+ |
2. |
K9351 |
+ |
3. |
HP 1633 |
+ |
4. |
Raj 4037 |
+ |
5. |
Sharbati Sonara |
+ |
6. |
K9533 |
+ |
7. |
K8434 |
+ |
8. |
NP 823 |
+ |
9. |
AJANTA |
+ |
10. |
PBW 12 |
– |
11. |
KRL |
– |
12. |
RW 346 |
– |
13. |
HD 2643 |
– |
14. |
HS 1097 |
– |
15. |
NP 825 |
– |
16. |
PBW 226 |
– |
17. |
NIAW 301 |
– |
18. |
PBW 343 |
– |
19. |
NI 179 |
– |
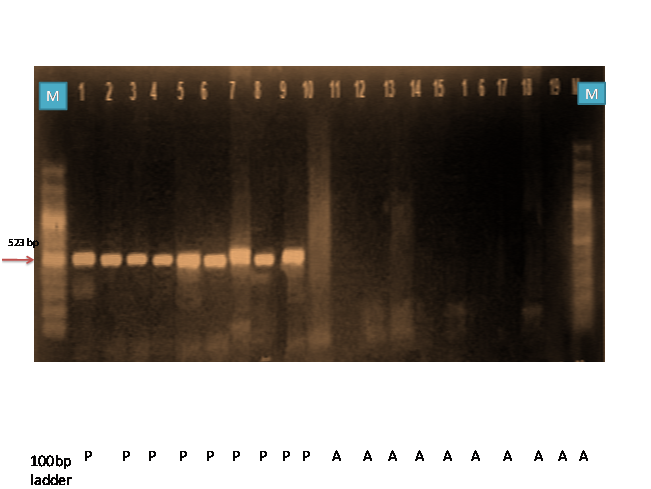
Fig. 1: PCR amplification of wheat germ plasm by cssfr2
M = Ladder Samples – 1 (SONARA 64) 2 (K9351) 3(HP 1633) 4(Raj 4037) 5(Sharbati Sonara) 6(K9533) 7(K8434) 8(NP 823) 9 ( Ajanta) 10( PBW 12) 11(KRL) 12 (RW 346) 13(HD 2643) 14(HS 1097) 15(NP 825) 16( PBW 226) 17(NIAW 301) 18(PBW 343) 19(NI 179)
Table (4):
Assessment of losses in wheat germplasm due to stripe rust (Puccinia striiformis) of at different growth stages.
| S. No. | Genotype | Per cent losses | ||
|---|---|---|---|---|
| Tillering stage | Flowering stage | Dough stage | ||
| 01 | Tawa | 0 | 16.35 | 20.75 |
| 02 | KRL | 0 | 16.35 | 20.75 |
| 03 | RAJ 4037 | 0 | 11.95 | 16.35 |
| 04 | RW 346 | 0 | 16.35 | 20.75 |
| 05 | Sonara 64 | 0 | 14.15 | 16.35 |
| 06 | Utkalia | 0 | 16.35 | 20.75 |
| 07 | HD 2643 | 0 | 20.75 | 22.95 |
| 08 | HS 1097 | 0 | 14.15 | 20.75 |
| 09 | NP 825 | 0 | 20.75 | 22.95 |
| 10 | WH 291 | 0 | 16.35 | 22.95 |
| 11 | HP 1633 | 0 | 16.35 | 16.35 |
| 12 | HUW 12 | 0 | 16.35 | 20.75 |
| 13 | PBW 226 | 0 | 16.35 | 20.75 |
| 14 | NP 823 | 0 | 14.15 | 16.35 |
| 15 | PV 18 | 0 | 16.35 | 20.75 |
| 16 | K8434 | 0 | 16.35 | 16.35 |
| 17 | NI 179 | 0 | 16.35 | 22.95 |
| 18 | HPW42 | 0 | 14.15 | 16.35 |
| 19 | NI 5439 | 0 | 16.35 | 22.95 |
| 20 | K9644 | 0 | 16.35 | 20.75 |
| 21 | K9351 | 0 | 14.15 | 14.15 |
| 22 | HD1925 | 0 | 16.35 | 22.95 |
| 23 | NIAW 301 | 0 | 11.95 | 14.15 |
| 24 | PBW 12 | 0 | 11.95 | 14.15 |
| 25 | PBW 65 | 0 | 22.95 | 25.15 |
| 26 | HUW 213 | 0 | 11.95 | 14.15 |
| 27 | Ajanta | 0 | 14.15 | 16.35 |
| 28 | K9533 | 0 | 16.35 | 16.35 |
| 29 | GW 503 | 0 | 16.35 | 22.95 |
| 30 | Sharbati Sonara | 0 | 16.79 | 18.55 |
| 31 | Agra Local | 0 | 36.15 | 44.95 |
- Ali, S., Leconte, M., Delos, M., Enjalbert, J and de, Vallavieille-Pope. Virulence dynamics and regional structuring of Puccinia striiformis f. sp. tritici in France between 1984 and 2009. Pl. Dis., 2012; 96: 131-140.
- Bariana, H. S., Brown, G. N., Ahmed, N. U., Khatkar, S., Conner, R. L., Wellings, C. R., Haley, S., Sharp, P. J and Laroche, A. Characterization of Triticum vavilovii derived stripe rust resistance using genetic, cytogenetic and molecular analyses and its marker-assisted selection. Theo. App. Gen., 2002; 104: 315-320.
- Broers, L. H. M. Partial resistance to wheat leaf rust in 18 spring wheat cultivars. Euphy., 1989; 44: 247-258.
- Chen, X. M. Epidemiology and control of stripe rust (Puccinia striiformis f. sp. tritici) on wheat. Can. J. Pl. Path., 2005; 27: 3.
- Doyle, J. J and Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull., 1987; 19: 11-15.
- Hussain, M., Mumtaz, H and Shah, J. A. Wheat rust scenario in 1994-1995. J. Phytopatho., 1996; 96-100.
- Khan, M. A and Saini, R. G. Non-hypersensitive leaf rust resistance of bread wheat cultivar PBW 65 conditioned by genes different than Lr34. Czech J. Gen.Pl Breed., 2009; 45: 26-30.
- Lagudah, E. S., Krattinger, S. G., Herrera-Foessel, S., Singh, R. P., Huerta-Espino, J., Spielmeyer, W., Brown-Guedira, G., Selter, L. L and Keller, B. Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theo. App. Gen., 2009; 119: 889-898.
- Milus, E. A and Line, R. F. Gene action for inheritance of durable, high temperature, adult plant resistance to stripe rust in wheat. Phytopatho., 1986; 76: 435-441.
- Munday, E. J. The effect of yellow rust and its control on the yield of Joss Cambier Winter Wheat. Pl. Pathol., 1973; 22: 171-176.
- Murray, G. M., Ellison, P. J., Watson, A and Cullis, B. R.. The relationship between wheat yield and stripe rust as affected by length of epidemic and temperature at grain development stage of crop growth. Pl. Patho., 1994; 43: 397-405.
- Pretorius, Z. A., Boshoff, W.H.P and Kema, G.H.J. First report of Puccinia striiformis f. sp. tritici on wheat in South Africa. Pl. Dis., 1997; 81: 424.
- Salman, A., Khan, M. A and Hussain, M. Prediction of yield losses in wheat varieties/lines due to leaf rust in Faisalabad. Pakistan J. Phytopatho., 2006; 18: 178-182.
- Sareen, P., Sundeep, K., Uttam, K., Lakshman, P,, Amit, K.S., Rakesh, S and Joshi, A.K.. Pathological and molecular characterizations of slow leaf rusting in fifteen wheat (Triticum aestivum L.) genotypes. African J. Biotech., 2012; 11: 14956-14966.
- Singh, R. P and Rajaram, S. Resistance to Puccinia recondita f.sp. tritici in 50 Mexican bread wheat cultivars. Crop Sci., 1991; 31: 1472-1479.
- Singh, R., Huerta-Espino, J., Bhavani, S., Herrera-Foessel, S., Singh, D., Singh, P., Velu, G., Mason, R., Jin, J., Njau, P and Crossa, J.. Race non-specific resistance to rust diseases in CIMMYT spring wheat. Euphy. , 2011; 179: 175-186.
- Singh, D.P., Sharma, A. K., Babu, K. S and Sharma, I. Multiple diseases, insect pests resistant genotypes and their utilization in breeding for resistance in wheat and Triticale. Greener J. Ag. Sc. 2014; 150-165.
- Yang, W. X and Liu, D. Q. Advances in localization and molecular markers of wheat leaf rust resistance genes. Sci. Ag. Sinica., 2004; 3: 770-779.
- Zeng, Q.D., Han, D. J., Wang, Q. L and Kang, Z. S. Stripe rust resistance and genes in Chinese wheat cultivars and breeding line. Euphy., 2014; 196: 271-284.
© The Author(s) 2017. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


