ISSN: 0973-7510
E-ISSN: 2581-690X
Cholera is one of the most important epidemic diseases globally. It is causing of morbidity and mortality in the world. Severe watery diarrhea causes a composite process involving several component that help them reach the lining of the small intestine, form colonies and generate bacterial toxins. This survey was carried out to discover the genetic patterns of clinical isolates according to the presence or absence of toxic genes for the city of Diwaniyah in southern Iraq. Sixty isolates were isolated from patients with cholera. The isolates included the center of the city and the surrounding rural areas. Biochemical and serological diagnosis. All the isolates were V. cholera serogroup O 1 of the serotyping Ogawa in biotype El Tor. Genetic testing was carried out using PCR technique and base on the presence or absence of toxin genes. Three genotypes were identified for the region.
Vibrio cholera, Biochemical and Surgical Diagnosis.
One of the epidemic disease is cholera. Acute watery diarrhea disease is causing common in Iraq. There are several region of Africa , Latin America and south of Asia where seasonal outbreaks happen (Neelam, et al.2012) And It was public health problem ( Shears ,2001).
V. cholera belong to the family Vibrionaceae which have 104 species (DSMZ. Leibniz) . The strains of genus V. cholera classified to serogroups according to epitopic different for lipopolysacharide for surface of cell (Yamai et al. 1997) The serogroup O 1 and O 139 among 210 that causal agent for cholera (Revera, et al.2002).
There are two distinct biotypes for O1 are classical and E1 Tor, every one of biotype has three serotypes are Ogawa , Inaba and Hokojima. (Nair, 2001).
Globally , the V. cholera have seven pandemics . Indian was originated for the first six pandemics of cholera via classical type for v. cholera O1.The seventh pandemic was E1 Tor that spread to the world which originated from Sulawesi in Indonesia. ( Carlos and Eduardo, 1996)
Iraq is confronting tremendous calamities of obliteration of the foundations with lack of sterilized water of drinking particularly in poor areas and exile camp that aide for the presence of contaminate water with cholera. (Al-Abbasi, et al., 2005)
In Iraq, 1999 reported the first occurrence of O 139 in Baghdad although few cases.(Al-Abbassi et al. 2005).After war of Iraq in 2003 with program of the communicable disease reported cholera epidemics in many region of Basrah (Valencino et al., 2003). Biotype El Tor of serogroup O1 include serotype Inaba isolated from 4,667 cases in Iraq when outbreak epidemic cholera in 2007.( Khwaif et al. 2010 ) . As well as, Ogawa serotype reported in this epidemic (Saleh, et al. 2011)
All governorates in Iraq isolate Vibrio cholera O1 serotype Inaba with few cases only of Ogawa serotype form 2013 to 2015. (Al-Abbassi et al.2015)
The enterotoxin (CT) of cholera which codes by ctxAB gene is causing the severity of disease (Joseph, et al.2015). Strains O1 and O139 of V. cholera possess the ctxAB gene on the 6.9-kb CTX prophage integrated within chromosome which responsible for encoding enterotoxins . These Enterotoxins in V. cholera cause diarrhea . And ace gene coding for accessory cholera enterotoxin as well as zot coding for zonula occludens toxin.
The aim of present study to determined genotypes of clinical isolates of V. cholera according to presence and absence of enterotoxin genes are ompW, ctxA, ace , zot by using PCR technique for outbreak and sporadic cholera in Diwaniyah city.
Specimen collection
In all, 60 isolates of V. cholera O1 were obtained in 2016 from patients having acute watery diarrhea .The samples were collected from Al Diwaniyah hospital in Diwaniyah city of Iraq.
These isolates were identified V. cholera O1 after culture and isolation (following World Health Organization 2014 guidelines) on TCBS agar (BIOMARK, Laboratories, India) ‘media via usual biochemical tests that included those for the fermentation of sugars in triple sugar iron agar medium, H2S and gas, indole formation, mannitol motility, oxidase, urease production , catalase production , nitrate reductase . Then isolates were confirmed by serology with commercially available antisera (Denka Seiken, Gosen-shi, Nilgata, Japan). Culture (18–24 h old) for these strains with specific density (0.5–0.63 McFarland standard, measured with colorimeter supplied with the system) and inoculate into sterile normal saline (supplied by BioMerieux, Lyon, France) then load on to Vitek 2 Compact system has software version 4.01 (BioMerieux, Durham, NC) following manufacturer’s instructions for identification using GN (BioMerieux) cards. The isolates were also sent to the Central Health Laboratory of Cholera and Enteric Diseases, Bagdad , which be the WHO Collaborating Centre for research on diarrheal diseases to identity for these isolates.
PCR test
Genomic DNA for V. coleara extracted carried out by geniod kit (Geniad ).Then amplification the target DNA by PCR in a thermal cycler (Techne TC-3000. USA) was performed in a reaction mixture tube which add to it 1µl for each” primer “(forward and reverse),5 µl from gDNA extract and add dionized water to complete the final volume 20 µl.”
The primers were used for cholera toxin in this study(ompW) (ctxA), (ace), (zot) . All primers weres provided by (Bioneer company. Korea) showing in (Table 1). The program of thermocycler was as follows: preincubation at 95 C for 2 min , 30 cycles for 30 sec. at 95C for denaturation, 1 min at 64.8C for annealing, 30 sec. at ( 57.8 ,62.9 and 59.9 for ompW, ctxA and zot respectively)C for elongation and incubation at 72C for 5 min for final elongation. This protocol for each ompW, ctxA and zot genes. While the protocol for ace gene was touchdown PCR protocol. It was performed on 45 cycles of predenaturation at 95°C for 2 min, followed by the 15 cycles with successive annealing temperature decrements that 0.5°C changed from 63°C to 56.5°C in every cycle versus time increments (30 sec in the first 15 cycles and 30 sec in the last 20 cycles). The reaction was denatured at 95°C for 30 min, followed by annealing at this temperature algorithm and polymerization at 72°C for 40 sec. Subsequently, the 20 cycles of denaturation at 95°C for 30 sec, annealing at 56°C for 30 sec, and elongation at 72°C for 40 sec were employed. The last primer extension step was done at 72°C for 5 min. Amplicons were made electrophoresis in agarose 1.5% gel . (Appligene, Illkirch, France) then stained with ethidium bromide (5mg/ml) for visualized.
Table (1):
Primers used in PCR*
Gene |
Sequence of nucleotides (5’-3’) |
PCR product Size (bp |
|---|---|---|
ompW- F ompW – R |
5’- CACCAAGAAGGTGACTTTATTGTG-3’ 5’- GAACTTATAACCACCCGCG3’ |
588 |
ctxA – F ctx – R |
5’- CGGGCAG ATTCTAGACCTCCTG-3’ 5’- CGATGATCTTGGAGCATTCC CAC-3’ |
564 |
ace – F ace – R |
5’- TAAGGATGTGCTTATGATGGACACCC-3’ 5’- CGTGATGAATAAAGATACTCATAGG-3’ |
316 |
zot – A zot – R |
5’- TCGCTTAACGATGGCGCGTTTT-3’ 5’- AACCCCGTTTCACTTCTACCCA-3’ |
947 |
Biotyping and serotyping
A sixty isolates of V. cholera were examined by serogroup, serotyping and biotyping. All isolates were diagnosed V. cholera serogroup O1 of serotyping Ogawa in biotypeb El Tor .
Genotyping
Genotypically, the description of isolates were shown in table(2). All V. cholera O1 isolates were positive for the species specific gene ompW. The gene of toxin ctxA was found in all isolates. And ace gene showed 36(60% ) of the isolate while 24(40%) was negative for this gene. Fifty seven of isolates (95%) was positive for Zot gene and 3 (5%) was negative . For study genotypes for isolates grouped isolate to three isolate based on presence and absence toxins of cholera table – 3.
Table (2):
Results of vibrio choleara genes
Genes |
No.teseted |
No. positive |
Positive % |
|---|---|---|---|
ompW gene |
60 |
60 |
100 |
ctxA gene |
60 |
60 |
100 |
Ace gene |
60 |
36 |
60 |
Zot gene |
60 |
57 |
95 |
Table (3):
Genotypes of the Isolates
Grouping of genoype |
CtxA |
Ace |
Zot |
No.isolates |
% |
|---|---|---|---|---|---|
Genotype 1 |
+ |
+ |
+ |
36 |
60% |
Genotype 2 |
+ |
– |
+ |
21 |
35% |
Genotype 3 |
+ |
+ |
– |
3 |
5% |
Total |
60 |
100% |
The larger part of V. cholerae strain in nature are thought its innocuous estuarine microbes. Nonetheless, particular strains seem to be advanced that reason sickness for people by successfully colonize in small digestive system and discharging a powerful enterotoxin . Four genes in chromosomal DNA codes for destructiveness related factors in V. cholerae (OmpW, ctxA, ace and zot) were incorporated into this investigation.
It found all isolate positive for Omp gene. So that indicate all samples were V. cholera. The strains isolated from the common source are important for epidemiology, through which the genetic relationship of the isolates is determined and determines the source of transmission, spread and origin of cholera infection. In previous studies demonstrated that omp W gene represented as inner control for testing V. cholera Goel, et al. 2007; Goel, et al. 2010; Izumiya, et al. 2011; Jain, et al. 2011. Our results agreement with Menezes et al. l concerned that identification of V. choleara via ompW gene give result 100% (Menezes et al. 2014).

Fig. 1. Agarose (1.5%) gel electrophoresis of ompW gene with 588 bp in lanes (1-20 ). M was DNA marker
The genetic elements of CTX has a compound transposon with 4.5 kb central core region (ctx AB ,zot , ace ,orfU, and cep)which encodes for both CT and functions for virion and flanked with one or more copies of a 2.7Kb repetitive sequences that responsible for regulation, replication and integration of ctxö “(Rivera et al.2001). we tested three genes in this region (ctxA, ace, and zot) in our isolates.
Enterotoxin gene ctx A found in all isolates. This gene is caustive agent to sever diarrhea. All our isolates are Toxogenic strains because possessing ctx elements (Julianai et al., 2000; Chen et al., 2004). And ctxA gene located in this element (Waldor et al., 1996) .This result agree with parrick et al.,2012 and Riveraa et al.,2001that their result found ctx A gene was 100% in your toxigenic isolates. Ratnam et al. show 98% for their isolation of V. cholera was positive for ctx gene in their clinical isolate (Ratnam et al. 2015). However, Menezes et al. Show negative results when tested environmental samples via ctxAB gene .other studies not found positive tested for ctxA gene in their isolates Gonçalves, et al. 2004. While THEOPHILO et al found one positive isolate for virulence factor ctx A gene in their environmental strains of non O1 and O139 Theophilo, et al. 2006.
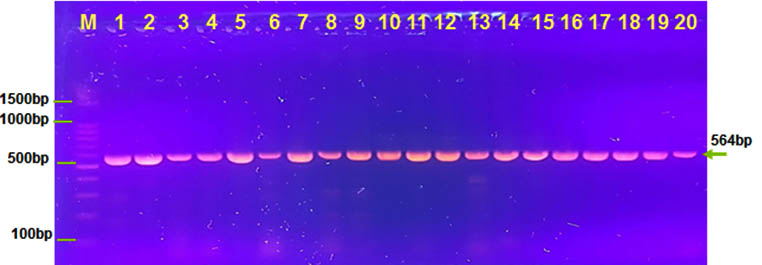
Fig. 2. Agarose (1.5%) gel electrophoresis of ctxA gene with 564 bp in lanes (1-20 ). M was DNA marker
Accessory cholera enterotoxin (ace) gene and (zot) gene show high percentage which was 60% and 95% in all our isolates .In contrast Pourshafie et al found lower percentage for virulence cassette genes (Pourshafie et al. 2002). The zot toxin is increases permeability of the gut via intercellular tight junction or (zonula occludens) (fasno et al.,1999).
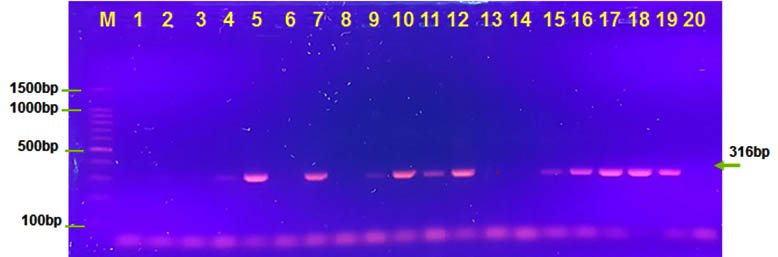
Fig. 3. Agarose (1.5%) gel electrophoresis of ace gene with 316 bp in lanes (5,7,9,10,11,12,15,16,17,18and 19). M was DNA marker
However, in all our result show 60% of V. cholera has three genes ctx, ace and zot in genotype 1. In genotype 2 show presence ctx, zot genes and absence ace gene was 35% for all isolates of V. cholera. However, Rivera et al. found that ctxA and zot genes in their toxigenic isolates was 100%. (Revira et al. 2001). Third genotype show presence ctx ,ace genes and absence zot gene was 5% for all isolates of V. cholera that result agree with many studies (Chowdhury et al., 1995 ; Ghosh et al., 1997; Karasawa et al., 1993).
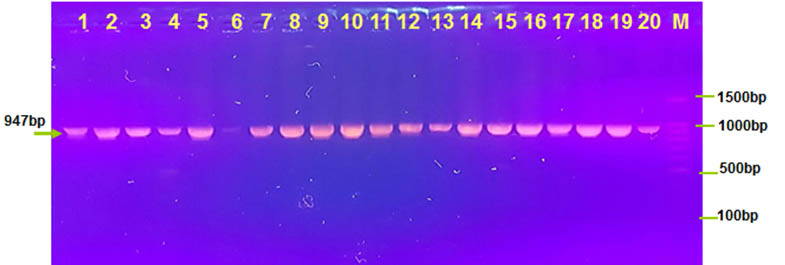
Fig. 4. Agarose (1.5%) gel electrophoresis of zot gene with 947 bp in lanes (1-20 except lane 6 ). M was DNA marker
Cholera is still a major epidemic disease that must be investigated and deepened. The continuous monitoring of cholera strains should be carried out through genomic research and the investigation of new strains. Despite the development of medical services. Precautions must be taken to prevent infection. Therefore , the clones were determined their genotype according to virulence factors and resistance to antibiotic .
- Al-Abbasi,Abdul Ridha and Aema,sabah M.2015.The cholera epidemic in Iraq during 2015.TOFIQ Journal of Medical Sciences.
- Nair,G.B. National Institute of Cholera and Enteric Diseases, Calcutta, India, with contributions from J. Bartram, Water, Sanitation and Health, World Health Organization, Geneva, Switzerland; A.H. Havelaar, Microbiological Laboratory of Health Protection, Rijksinstituut voor Milieuhygiene en Voldsgezondheid, Bilthoven, Netherlands: J. Hueb, Water, Sanitation and Health, World Health Organization, Geneva, Switzerland; and J. Jacob, Institute für Wasser-, Boden- and Lufthygiene, Bad Elster, Germany.
- Joseph,P.V.; Sundararaj,R.T.; Rajkumar,S.; Anup,A.and Banu,A.S. Genotyping of Vibrio cholerae strains based on the cholera toxin and virulence associated genes.medical scines, 2015; 5(7).
- Neelam Taneja, Garima Sangar, Goutam Chowdhury*, T. Ramamurthy*, Arti Mishra,Meenakshi Singh & Meera Sharma Molecular epidemiology of Vibrio cholerae causing outbreaks & sporadic cholera in northern India. Indian J Med Res, 2012; 136: pp 656-663.
- Carlos S. and Eduardo G., 1996. Cholera: Overview of Epidemic, Therapeutic, and Preventive Issues Learned from Recent Epidemics. International Journal of Infection 1:37-46.
- Shears P. Recent developments in cholera. Curr. Opin. Infect. Dis. 2001; 14: 553 – 558.
- DSMZ. Leibniz-Institu Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH. Bacterial diversity. [cited 2012 Jan. 24]. Available from: http://www.dsmz. de/bacterial-diversity.html
- Yamai S, Okitsu T, Shimada T, Katsube Y. Distribution of serogroups of Vibrio cholera non-O1 non-O139 with specific reference to their ability to produce cholera toxin, and addition of novel serogroups. Kansenshogaku Zasshi, 1997; 71:1037–1045
- Al-Abbassi AM, Ahmed S, Al-Hadithi T (2005) Cholera epidemic in Baghdad during: clinical and bacteriological profile of hospitalized cases. East Mediterr Health J, 1999; 11:6–13
- Valenciano M, Coulombier D, Lopes Cardozo B. Challenges for communicable disease surveillance and control in southern Iraq, April–June 2003. JAMA, 2003; 290: 654–658
- Khwaif JM, Hayyawi AH, Yousif TI. Cholera outbreak in Baghdad in 2007: an epidemiological study. East Mediterr Health J, 2010; 16: 584–589
- Saleh TH, Sabbah MA, Jasem KA, Hammad ZN. Identification of virulence factors in Vibrio cholerae isolated from Iraq during the 2007–2009 outbreak. Can J Microbiol, 2011; 57:1024–1031
- Juliana, M.V.B., Ronaldo M.A., Irma N.G.R., Dalia P. R., David K.R.K. and Campos C, Prevalence of virulence-associated genes in clinical and environmental Vibrio cholerae strains isolated in Brazil between 1991 and 1999. FEMS Microbiology Letters, 2002; 215: 15-21.
- Chen, C.H., Shimada, T., Elhadi, N., Radu, S., and Nishibushi, M. Phenotypic and Genotypic characteristic and epidemiological significance of ctx + strains of Vibrio cholerae isolated from seafood in Malaysia. Applied and Environmental Microbiology, 2004; 70: 1964-1972
- Waldor, M. K., Tschape, H. and Mekalanos J. J. A new type of conjugative transposon encoded resistance to sulfamethoxazole, trimethoprim, and streptomycin in Vibrio cholerae 0139. Journal of Bacteriology, 1996; 178: 4157-65
- Patrick, G. B., 3Nishibuchi, M., 2Tunung, R. and 2Son, R. Molecular characterization of clinical isolate of Vibrio cholerae isolated from outbreaks cases in Malaysia. International Food Research Journal, 2012; 19(3): 1267-1274
- Fasano, A., B. Baudry, D. W. Pumplin, S. S. Wasserman, B. D. Tall, J. M. Ketley, and J. B. Kaper. Vibrio cholerae produces a second enterotoxin which affects intestinal tight junctions. Proc. Nat. Acad. Sci. USA, 1991; 88:5242– 5246.
- Ghosh, C., R. K. Nandy, S. K. Dasgupta, G. B. Nair, R. H. Hall, and A. C. Ghose. A search for cholera toxin (CT), toxin coregulated pilus (TCP), the regulatory element ToxR and other virulence factors in non-O1/non- O139 Vibrio cholerae. Microb. Pathog. 1997; 22: 199–208.
- Karasawa, T., T. Mihara, H. Kurazono, B. G. Nair, S. Garg, T. Ramamurthy, and Y. Takeda. Distribution of the zot (zonula occludens toxin) gene among strains of Vibrio cholerae O1 and non-O1. FEMS Microbiol. Lett. 1993; 106:143–146.
- Chowdhury, M. A. R., R. T. Hill, and R. R. Colwell. A gene for the enterotoxin zonula occludens toxin is present in V. mimicus and V. cholera O139. FEMS Microbiol. Lett. 1994; 119:377–380.
- Rivera,I.N.G.; Chun,J.; Huq,A.;Sack,R.B.; And Colwellr.R. Genotypes Associated With Virulence In Environmental Isolates Of Vibrio Cholera. Applied And Environmental Microbiology; 2001; 67(6) : 2421–2429.
- Theophilo GND, Rodrigues DP, Leal NC, Hofer E. Distribution of virulence markers in clinical and environmental Vibrio cholerae non-O1/non-O139 strains isolated in Brazil from 1991 to 2000. Rev Inst Med Trop Sao Paulo. 2006; 48:65-70.
- Menezes, F.G.; Neves,S.S.; Sousa,O.V.; Vila-Nova,M.V.; Maggioni,R.; Diogo,G.N.; Theophilo; Hofer,E.; & Vieira,R.H.2014. Detection Of Virulence Genes In Environmental Strains Of Vibrio Cholerae From Estuaries In Northeastern Brazil. Rev. Inst. Med. Trop. Sao Paulo 56(5):427-432,
- Goel AK, Jain M, Kumar P, Jiang SC. Molecular characterization of Vibrio cholerae outbreak strains with altered El Tor biotype from southern India. World J Microbiol Biotechnol. 2010; 26: 281-7.
- Goel AK, Ponmariappan S, Kambol DV, Singh L. Single multiplex polymerase chain reaction for environmental surveillance of toxigenic-pathogenic O1 and non-O1 Vibrio cholerae. Folia Microbiol. 2007 ; 52:81-5.
- Izumiya H, Matsumoto K, Yahiro S, Lee J, Morita M, Yamamoto S, et al. Multiplex PCR assay for identification of three major pathogenic Vibrio spp., Vibrio cholerae, Vibrio parahaemolyticus, and Vibrio vulnificus. Mol Cell Probes. 2011 ; 25:174-6.
- Jain M, Goel AK, Bhattacharya P, Ghatole M, Kamboj DV. Multidrug resistant Vibrio cholerae O1 El Tor carrying classical ctxB allele involved in a cholera outbreak in South Western India. Acta Trop. 2011 ; 117:152-6.
- Gonçalves EGR, Leal NC, Hofer E. Estudo molecular de Vibrio cholerae não-O1 isolado de zooplâncton da Baía de São Marcos/São Luis – MA, Brasil. Rev Soc Bras Med Trop. 2004;37:324-8.
- Ratnam,P.; Sundararaj,T.; Rajkumar,S.; Anand,A.; Banu,S. Genotyping of Vibrio cholerae strains based on the cholera toxin and virulence associated genes. Medical Sciences, 2005; 5 : 196-199.
- Pourshafie,M.; Grimont,F. Kohestani,S. And Grimont,P.A.D. A molecular and phenotypic study of Vibrio cholerae in Iran. J. Med. Microbiol, 2002 ; 51: 392–398.
© The Author(s) 2018. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


