ISSN: 0973-7510
E-ISSN: 2581-690X
Vulvovaginal candidiasis (VVC) is a commonly found disease in antenatal women caused by Candida species. The usual presentations are persistent curdy white vaginal discharge with itching, bad odour, irritation, pain in the lower abdomen and local induration of vulva. Therefore this infection is an unpleasant and cause of intolerance for women. Several virulence factors are present in Candida species such as Secreted aspartyl proteinase (Sap), phospholipase, lipase, haemolysin, germ tube formation and production of biofilm. There are 10 SAP genes (SAP1 to SAP10) present in all Candida species which help for virulence. The objective of this research was to analyse the genotypic patterns of SAP genes in Candida species. The present research was done in SRM MCH & RC, Chennai, India, from March 2017 to December 2018. DNA was isolated and the SAP gene was detected by a polymerase chain reaction. Totally 35 Candida species were isolated from 92 suspected case of VVC. Out of 35 Candida species, 16/35 (45.7%) were from symptomatic women and 19/35 (54.3%) were from asymptomatic women. Out of 35 Candida species, C. albicans were 15(42.8%) and Non-albicans Candida (NAC) were 20(57.2%). Totally 28 genotypic patterns were found and 6 (21.4%) Candida isolates presented genotypic patterns of all the SAP genes. There is a predominance of the SAP1 gene in Candida isolates from asymptomatic women whereas SAP1 and SAP9 genes in Candida isolates from symptomatic women. The statistical analysis showed a significant difference for the SAP7 and SAP8 gene frequency among the Candida isolated from symptomatic and asymptomatic women.
Vulvovaginal candidiasis, symptomatic, asymptomatic, antenatal, Candida albicans, Secreted aspartyl proteinase.
In recent years there is an increased frequency of Candida infection and it has been the cause of a major rise in morbidity and mortality1. Vulvovaginal candidiasis (VVC) is a commonly found disease in antenatal women. The usual presentations are persistent curdy white vaginal discharge with itching, bad odour, irritation, pain in the lower abdomen and induration of vulva. Therefore, this infection is an unpleasant, intolerant condition for women2. The causative agents of this disease are Candida species which are yeast-like fungus present as a commensal of the skin, mucosal cavity, GI tract and vagina causing local and systemic infection in patients with the immunocompromised condition3. VVC is mostly seen in women with reproductive age group between 18 to 45 years2,3. Various studies show that about 75% of all women will have at least one episode of candidiasis during their lifetime3,4. The prevalence rate of vulvovaginal candidiasis in antenatal women is more than 40% worldwide and 5-10% suffers from Recurrent Vulvovaginal Candidiasis (RVVC). During pregnancy, VVC has been associated with complications such as low birth weight of babies, miscarriage, preterm birth and premature rupture of the membrane5,6. Candida species possess several virulence factors like extracellular production of hydrolytic enzymes, hyphae formation, phenotype switching and cell adhesion7,8. The emission of the extracellular hydrolytic enzyme by Candida species appeared to be the most important virulence factors9. The most vital hydrolytic enzymes produced by C. albicans are “Secreted aspartyl proteinases (Sap), phospholipase B, esterase and lipases”. C. albicans isn’t the only organism which produced Sap other Candida species have also been found to demonstrate SAP genes, including C. dubliniensis10, C. tropicalis and C. parapsilosis11. Sap enzyme production is related to various other virulence factors of Candida species including, hyphal development, adhesion to the surface, phenotypic exchanging, and dimorphism12. The sap is encoded by a multi-gene family including ten diverse highly regulated genes (SAP1 to SAP10). The presence of 10 SAP genes and their controlled articulation and guideline brings up various issues regarding the role and elements of this gene amid the infective progression. Being thereof 10 SAP genes, during their sequential activation in particular phases of infection, they organize as well as constitute a robust body and indicate that distinct members of this gene family having a significant role in the adaptive response of Candida species to its environment, including its host13. Thus the point of this investigation was to assess the genotypic patterns of SAP (SAP1 to SAP10) genes in Candida causing VVC in antenatal women.
It was a prospective study done in SRM tertiary healthcare Centre, from March 2017 to December 2018 after getting approval from the Institutional Ethical Committee (IEC, 1090/IEC/2017). Antenatal women were divided into two groups: (a) symptomatic patients characterised by the presence of white curdy discharge, itching, burning, oedema, erythema of vulva and vagina, (b) Asymptomatic patients. After getting informed consent from the antenatal women, two consecutive vaginal swabs were taken for direct microscopy and culture respectively. All samples were collected from the patient prior to the antifungal treatment. All samples were examined in the direct microscope by 10 % Potassium hydroxide (KOH) mount followed by Gram stain and cultured on “Sabouraud’s Dextrose Agar (SDA)” with supplement (gentamycin) to prevent bacterial contamination and incubated for 48 hours at 37°C and 25°C. Candida was identified by Gram stain to demonstrate budding yeast cell, hyphae and pseudohyphae. Germ tube test was performed to differentiate Candida albicans from Non-albicans Candida (NAC). Subsequently, Candida species were identified by cultured on chrome agar (Hi-media, India) a differential media for detection of coloured colonies and corn meal agar with tween 80 for chlamydospores and blastopores formation. Sugar fermentation and assimilation tests were done to identify Candida species, C. albicans ATCC90028 was used as a control strain. Besides the identification of Candida species by the conventional method, the species were also confirmed by “Matrix-Assisted Laser Desorption/Ionization Time-Of-Flight” (MALDI-TOF), for which samples were sent to Micro lab diagnostic centre, Coimbatore, Tamil Nadu, India.
Genomic DNA extraction and Purification
Candida genomic DNA of each Candida isolates was extracted and purified by using PureFast® fungal DNA mini spin purification kit (Helini biomolecules, Chennai) according to the manufacturer’s guidelines. Concisely, 1ml of the overnight culture was centrifuged at 6000rpm for 5min, the supernatant was discarded and the pellet was suspended in 0.2ml Phosphate Buffer Saline (PBS). 180µl of lysozyme digestion buffer and 20µl of lysozyme [10mg/ml] was added and incubated at 37oC for 15min. 400µl of Binding buffer, 5µl of internal control template and 20µl of proteinase K was added, mixed well by inverting several times and incubated at 56°C for 15min. 300µl of ethanol was added and mixed well. The entire sample was transferred into the spin column and centrifuged for 1 min. The supernatant was discarded and 500µl of wash buffer-1 was added then centrifuged for 30-60 seconds and discarded the flow-through. Again 500µl of wash buffer-2 was added and centrifuged for 30-60 seconds. The flow-through discarded and centrifuged for an additional 1 min, then transferred into a fresh 1.5 ml microcentrifuge tube. 100µl of elution buffer was added then incubated for 1 min at room temperature and centrifuged for 2 min. The purified DNA was stored at -20°C. Quality and quantity of extracted DNA were checked by loading in 1% agarose gel and 5µl of extracted DNA was used for PCR amplification.
PCR amplification and Gel Electrophoresis
SAP Genes were identified by PCR based on the method proposed by Bassyouni et al.20 with minor modification. Specific primers of SAP genes were used for the detection of Candida SAP genes by PCR as shown in table 1 (Helini biomolecules, Chennai).
Amplification program as it follows an initial denaturation at 95°C for 5 minutes, then 35 amplification cycles consisting of denaturation: at 95°C for 30 seconds, annealing: at 58°C for 30 seconds, extension: at 72°C for 30 seconds and final extension: at 72°C for 5 minutes. DNA amplification was done by thermal cycler; the final volume of the reaction mixture was 25µl. In that, sample (DNA) was 5µl, master mix- 5µl, sterile water- 11µl, primer- 2µl of each forward and reverse. After amplification 5µl of each PCR product were analysed by gel electrophoresis. For gel electrophoresis, 1% agarose was used. One gram of agarose was liquefied in 100 ml of 1x “Tris Borate EDTA (TBE)”. 1µl of “ethidium bromide” was added. 5µl of the amplified product along with the negative control and 1 kb DNA marker was laden into the well. The gel was run in electrophoresis at 50V till the dye reached three fourth of the gel and bands were observed in “UV Transilluminator” [Figs. 1-5].
Table (1):
Primers used to identify SAP gene
| Genes | Primers | Sequence (5´-3´) | Size of amplified product (bp) |
|---|---|---|---|
| SAP1 | SAP1F | CTGTTGGTTTTGGTGGTGCT | 200 |
| SAP1R | CAGCATTGGGAGAGTTGAGA | ||
| SAP2 | SAP2F | GCTGTCCCAGTGACTTTACACA | 409 |
| SAP2R | CATAACTACCACCGGCTTCA | ||
| SAP3 | SAP3F | AATGTTACTGGTCCCCAAGGT | 605 |
| SAP3R | AAAGCAATCAATGTCCCACT | ||
| SAP4 | SAP4F | ACTGGCTCTTCTGACTTGTGG | 733 |
| SAP4R | AAGCAGGAACGGAAATCTTG | ||
| SAP5 | SAP5F | CCCGTCGATGAGACTGGTAG | 868 |
| SAP5R | CAGGAACGGAGATCTTGAGG | ||
| SAP6 | SAP6F | ACAAAAATTCCCGTCGATGA | 794 |
| SAP6R | CCAGCAGAATCAAAATGCAC | ||
| SAP7 | SAP7F | AGCTTCCTCGGCATCTTCTT | 600 |
| SAP7R | TAACCTTGGACAGCATCACG | ||
| SAP8 | SAP8F | TTGACCGCATTACCAATCAC | 394 |
| SAP8R | TGGTGTTCCCATCAAGATCA | ||
| SAP9 | SAP9F | TGGGTGTTATGCAACAATCG | 213 |
| SAP9R | TGGTAGAGGTGCCAGATGAA | ||
| SAP10 | SAP10F | CAAAGTCCTGACACCATCCA | 102 |
| SAP10R | TTCGAACCGATCTCCAATTC |
SAP= Secreted Aspartyl Proteinase, F= forward, R= reverse primer.
Statistical Analysis
Data were entered into an excel sheet and analysed by using R version 3.5.2 software. Z test has been used to find the difference in proportions and a 5% level of significance has been used.
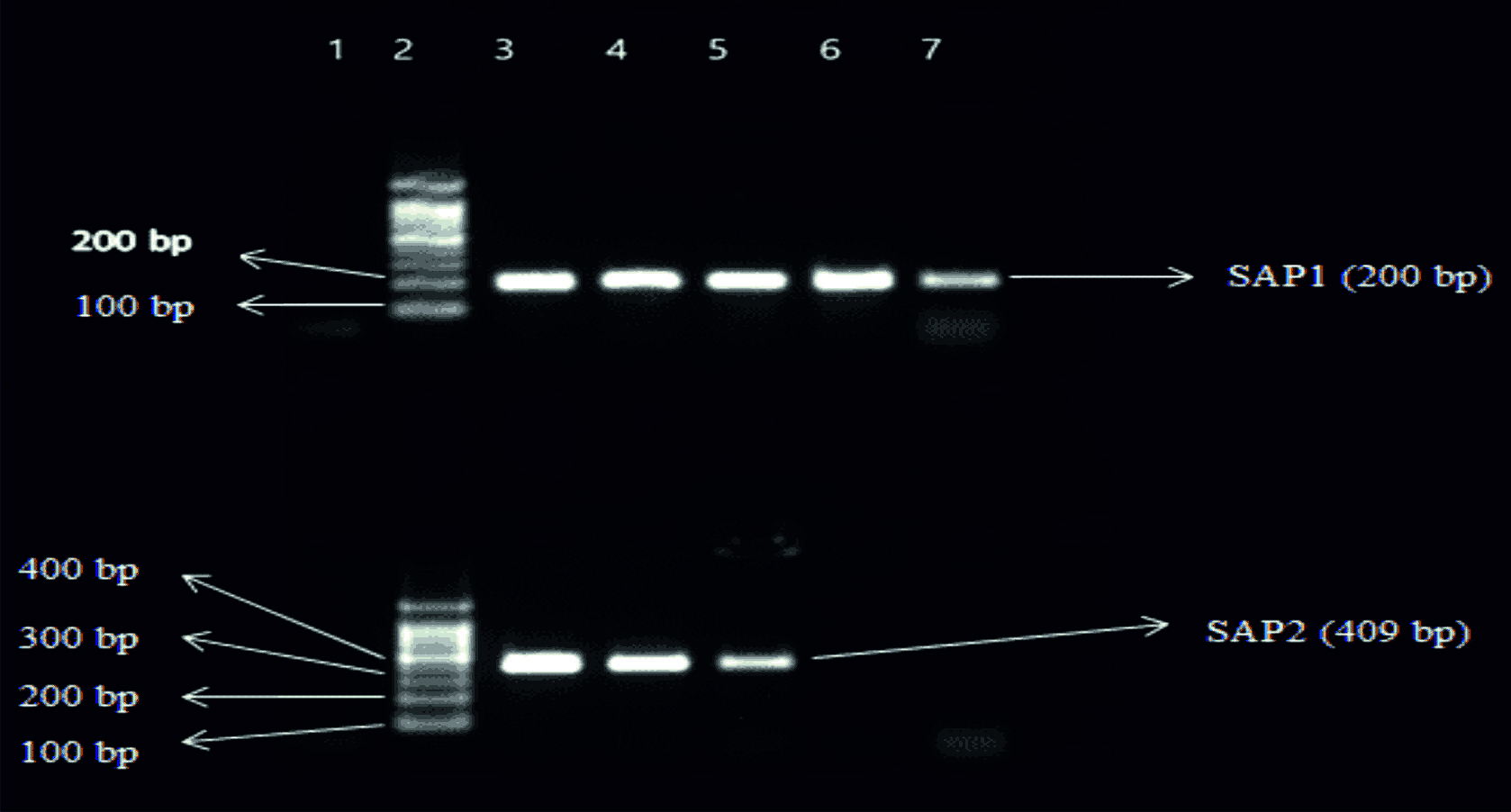 Fig. 1. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP1(200bp) of Candida albicans, 4- SAP 1 of Candida albicans, 5-SAP1 of Candida tropicalis, 6-SAP1 of Candida krusei, 7-SAP1 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP2(409bp) of Candida albicans, 4- SAP2 of Candida albicans, 5- SAP2 of Candida tropicalis, 6- SAP2 of Candida krusei, 7- SAP2 of Candida glabrata.
Fig. 1. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP1(200bp) of Candida albicans, 4- SAP 1 of Candida albicans, 5-SAP1 of Candida tropicalis, 6-SAP1 of Candida krusei, 7-SAP1 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP2(409bp) of Candida albicans, 4- SAP2 of Candida albicans, 5- SAP2 of Candida tropicalis, 6- SAP2 of Candida krusei, 7- SAP2 of Candida glabrata. Fig. 2. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP3(605bp) of Candida albicans, 4- SAP3 of Candida albicans, 5-SAP3 of Candida tropicalis, 6-SAP3 of Candida krusei, 7-SAP3 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP4(733bp) of Candida albicans, 4- SAP4 of Candida albicans, 5- SAP4 of Candida tropicalis, 6- SAP4 of Candida krusei, 7- SAP4 of Candida glabrata.
Fig. 2. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP3(605bp) of Candida albicans, 4- SAP3 of Candida albicans, 5-SAP3 of Candida tropicalis, 6-SAP3 of Candida krusei, 7-SAP3 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP4(733bp) of Candida albicans, 4- SAP4 of Candida albicans, 5- SAP4 of Candida tropicalis, 6- SAP4 of Candida krusei, 7- SAP4 of Candida glabrata. Fig. 3. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP5 (868bp) of Candida albicans, 4- SAP5 of Candida albicans, 5-SAP5 of Candida tropicalis, 6-SAP5 of Candida krusei, 7-SAP5 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP6 (794bp) of Candida albicans, 4- SAP6 of Candida albicans, 5- SAP6 of Candida tropicalis, 6- SAP6 of Candida krusei, 7- SAP6 of Candida glabrata
Fig. 3. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP5 (868bp) of Candida albicans, 4- SAP5 of Candida albicans, 5-SAP5 of Candida tropicalis, 6-SAP5 of Candida krusei, 7-SAP5 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP6 (794bp) of Candida albicans, 4- SAP6 of Candida albicans, 5- SAP6 of Candida tropicalis, 6- SAP6 of Candida krusei, 7- SAP6 of Candida glabrata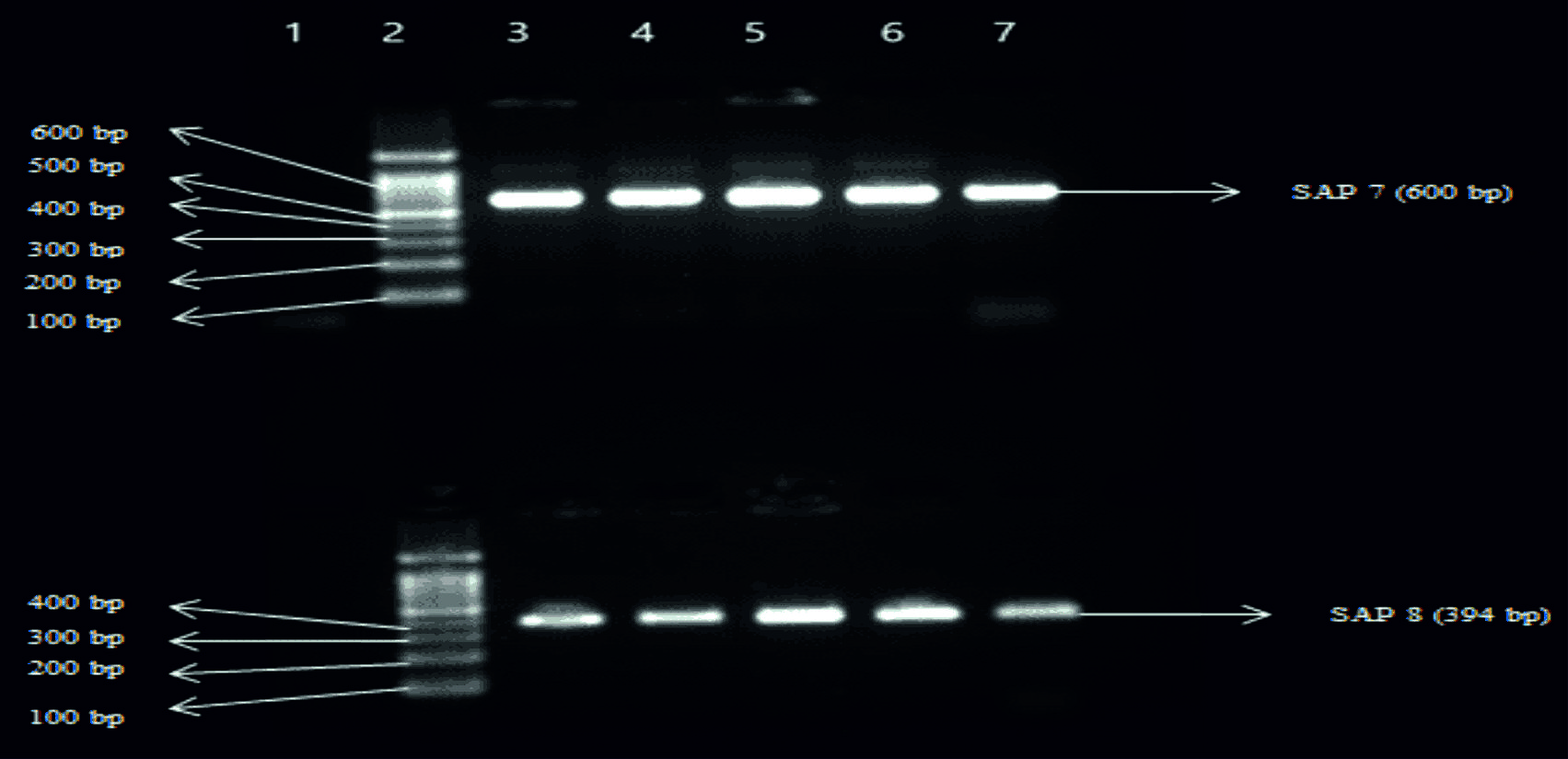 Fig. 4. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP7 (600bp) of Candida albicans, 4- SAP7 of Candida albicans, 5-SAP7 of Candida tropicalis, 6-SAP7 of Candida krusei, 7-SAP7 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP8 (394bp) of Candida albicans, 4- SAP8 of Candida albicans, 5- SAP8 of Candida tropicalis, 6- SAP8 of Candida krusei, 7- SAP8 of Candida glabrata.
Fig. 4. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP7 (600bp) of Candida albicans, 4- SAP7 of Candida albicans, 5-SAP7 of Candida tropicalis, 6-SAP7 of Candida krusei, 7-SAP7 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP8 (394bp) of Candida albicans, 4- SAP8 of Candida albicans, 5- SAP8 of Candida tropicalis, 6- SAP8 of Candida krusei, 7- SAP8 of Candida glabrata.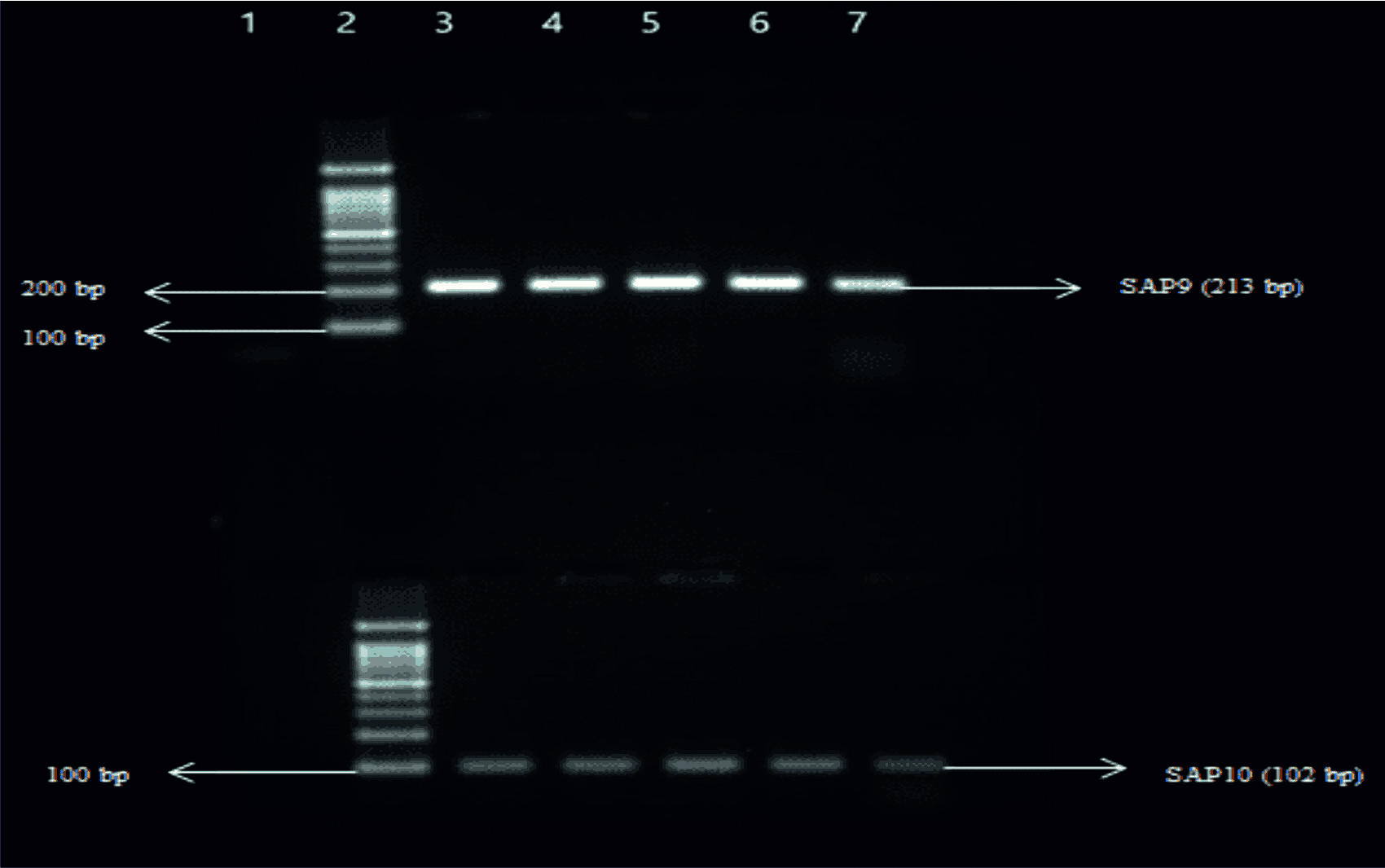 Fig. 5. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP9 (213bp) of Candida albicans, 4- SAP9 of Candida albicans, 5-SAP9 of Candida tropicalis, 6-SAP9 of Candida krusei, 7-SAP9 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP10 (102bp) of Candida albicans, 4- SAP10 of Candida albicans, 5- SAP10 of Candida tropicalis, 6- SAP10 of Candida krusei, 7- SAP10 of Candida glabrata.
Fig. 5. Top Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP9 (213bp) of Candida albicans, 4- SAP9 of Candida albicans, 5-SAP9 of Candida tropicalis, 6-SAP9 of Candida krusei, 7-SAP9 of Candida glabrata. Bottom Gel- 1-Negative control, 2-Ladder (1000bp), 3- SAP10 (102bp) of Candida albicans, 4- SAP10 of Candida albicans, 5- SAP10 of Candida tropicalis, 6- SAP10 of Candida krusei, 7- SAP10 of Candida glabrata.Thirty-five Candida species were isolated from 92 symptomatic and asymptomatic antenatal women. Out of 35 Candida species 16/35 (45.7%) were from symptomatic (Infection) women and 19/35 (54.3%) were from asympto-matic (Colonization) women. Out of 35 Candida species, 15/35 (42.8%) were C. albicans and 20/35 (57.7%) were NAC as shown in table 2.
Table (2):
Candida species distribution in symptomatic and asymptomatic women (n=35).
| Candida species
n (%) |
C. albicans
n (%) |
NAC*, n (%) | |||
|---|---|---|---|---|---|
| C. glabrata
n (%) |
C. tropicalis
n (%) |
C. parapsilosis
n (%) |
C. krusei
n (%) |
||
| Symptomatic women, (n=16) | 9(56.2) | 2(12.5) | 3(18.7) | 1(6.2) | 1(6.2) |
| Asymptomatic women, (n=19) | 6(31.5) | 6(31.5) | 4(21) | 2(10.5) | 1(5.2) |
| Total Candida species (n=35) | 15(42.8) | 8(22.8) | 7(20) | 3(8.5) | 2(5.7) |
*NAC = Non-albicans Candida
It was possible to detect the presence of SAP1 to SAP10 genes by PCR. Totally 16 isolates of Candida species from symptomatic women were subjected for SAP gene detection. Out of 16 isolates of Candida species, 9 were C. albicans followed by three C. tropicalis, two C. glabrata, one C. parapsilosis and C. krusei each. In C. albicans 100% (9/9) of SAP1, 100% (9/9) of SAP2, 66.6% (6/9) SAP3, 77.7% (7/9) of SAP4, 55.5% (5/9) of SAP5, 77.7% (7/9) of SAP6, 77.7% (7/9) of SAP7, 100% (9/9) of SAP8, 100% (9/9) of SAP9 and 66.6% (6/9) SAP10 genes were detected. For NAC data showed in table 3.
Table (3):
Detection of secreted aspartyl proteases (SAP) gene in Candida isolates from symptomatic women (n=16).
SAP genes |
SAP1
n (%) |
SAP2
n (%) |
SAP3
n (%) |
SAP4 n (%) |
SAP5n (%) |
SAP6
n (%) |
SAP7
n (%) |
SAP8 n (%) |
SAP9
n (%) |
SAP10
n (%) |
|---|---|---|---|---|---|---|---|---|---|---|
C. albicans (n=9) |
9 (100) |
9
(100) |
6
(66.6) |
7
(77.7) |
5
(55.5) |
7
(77.7) |
7
(77.7) |
9
(100) |
9
(100) |
6
(66.6) |
C. tropicalis
(n=3) |
3
(100) |
2
(66.6) |
2
(66.6) |
2
(66.6) |
2
(66.6) |
2
(66.6) |
3
(100) |
2
(66.6) |
3
(100) |
2
(66.6) |
C. glabrata
(n=2) |
2
(100) |
2
(100) |
2
(100) |
2
(100) |
1
(50) |
1
(50) |
1
(50) |
2
(100) |
2
(100) |
2
(100) |
C. parapsilosis (n=1) |
1
(100) |
Nil |
1
(100) |
1
(100) |
1
(100) |
Nil |
1
(100) |
Nil |
1
(100) |
1
(100) |
C. krusei (n=1) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
1
(100) |
Total Candida species (n=16) |
16
(100) |
14
(87.5) |
12
(75) |
13
(81.2) |
10
(62.5) |
11
(68.7) |
13
(81.2) |
14
(87.5) |
16
(100) |
12
(75) |
SAP= Secreted Aspartyl Proteinase
Totally 19 isolates of Candida species from asymptomatic women were subjected for SAP gene detection. Out of 19 isolates of Candida species, 6 were C. albicans and C. glabrata each followed by four C. tropicalis, two C. parapsilosis and one C. krusei. In C. albicans 83.3% (5/6) of SAP1, 100% (6/6) of SAP2, 33.3% (2/6) SAP3, 66.6% (4/6) of SAP4, 16.6% (1/6) of SAP5, 50% (3/6) of SAP6, 33.3% (2/6) of SAP7, 50%(3/6) of SAP8, 100% (6/6) of SAP9, 50%(3/6) of SAP10 genes were detected, for NAC data shown in table 4.
Table (4):
Detection of SAP gene in Candida isolates from Asymptomatic women (n=19).
SAP gene |
SAP1
n (%) |
SAP2 n(%) |
SAP3n (%) |
SAP4
n (%) |
SAP5 n (%) |
SAP6 n (%) |
SAP7
n (%) |
SAP8
n (%) |
SAP9
n (%) |
SAP10
n (%) |
|---|---|---|---|---|---|---|---|---|---|---|
C. albicans (n=6) |
5
(83.3) |
6
(100) |
2
(33.3) |
4
(66.6) |
1
(16.6) |
3
(50) |
2
(33.3) |
3
(50) |
5
(83.3) |
3
(50) |
C. glabrata (n=6) |
5
(83.3) |
3
(50) |
3
(50) |
3
(50) |
2
(33.3) |
3
(50) |
1
(16.6) |
2
(33.3) |
6
(100) |
3
(50) |
C. tropicalis (n=4) |
4
(100) |
2(50) |
2(50) |
2
(50) |
2
(50) |
2
(50) |
2
(50) |
4
(100) |
3
(75) |
1
(25) |
C. parapsilosis (n=2) |
1
(50) |
2
(100) |
Nil |
Nil |
Nil |
2
(100) |
Nil |
Nil |
2
(100) |
Nil |
C. krusei (n=1) |
1
(100) |
1
(100) |
Nil |
Nil |
Nil |
1
(100) |
1
(100) |
1
(100) |
Nil |
Nil |
Total Candida species (n=19) |
16
(84.2) |
12
(63.1) |
7
(36.8) |
9
(47.3) |
5
(26.3) |
11
(57.8) |
6
(31.5) |
10
(52.6) |
16
(84.2) |
7
(36.8) |
SAP= Secreted Aspartyl Proteinase
The highest frequency of SAP genes in Candida species isolated from antenatal women with symptomatic (infection) women of VVC was 100% (16/16) for SAP1 and SAP9 whereas SAP1 was the predominant in asymptomatic (colonization) women, for other SAP genes as data shown in table 5. Of all the genotypic patterns for the members of the SAP genes family in the Candida species, a total of 28 patterns were found in this present study. Out of 28 patterns 6 (21.4%) Candida isolates showed all the SAP genes
(Table 6).
Table (5):
Frequency of SAP genes in various Candida species isolated from vulvovaginal infections and colonization.
Genes |
Symptomatic/Infections (n=16)
n (%) |
Asymptomatic/Colonization (n=19)
n (%) |
Total (n=35)
n (%) |
P-value |
|---|---|---|---|---|
SAP1 |
16(100) |
15(78.9) |
31(88.5) |
0.1565 |
SAP2 |
14(87.5) |
14(73.6) |
28(80) |
0.5527 |
SAP3 |
12(75) |
7(36.8) |
19(54.2) |
0.0552 |
SAP4 |
13(81.2) |
9(47.3) |
22(62.8) |
0.0863 |
SAP5 |
10(62.5) |
5(26.3) |
15(42.8) |
0.0699 |
SAP6 |
11(68.7) |
10(52.6) |
21(60) |
0.5331 |
SAP7 |
13(81.2) |
7(36.8) |
20(57.1) |
0.0213* |
SAP8 |
14(87.5) |
9(47.3) |
23(65.7) |
0.0328* |
SAP9 |
16(100) |
14(73.6) |
30(85.7) |
0.0834 |
SAP10 |
12(75) |
9(47.3) |
21(60) |
0.1882 |
*Statistically significant (p<0.05), SAP= Secreted Aspartyl Proteinase
Table (6):
Genotypic patterns identified in vaginal isolates of various Candida species
Pattern |
Genotype |
n (%) |
|---|---|---|
1 |
SAP1 to SAP10 |
6(21.4) |
2 |
SAP1, SAP2 and SAP4 to SAP10 |
2(7.1) |
3 |
SAP1, SAP2, SAP3 and SAP5 to SAP10 |
2(7.1) |
4 |
SAP1, SAP2 and SAP5 to SAP10 |
2(7.1) |
5 |
SAP1, SAP2, SAP4 and SAP6 to SAP10 |
1(3.5) |
7 |
SAP1 to SAP3 and SAP7 to SAP10 |
1(3.5) |
8 |
SAP1, SAP3, SAP4 and SAP6 to SAP10 |
1(3.5) |
9 |
SAP1 to SAP4, SAP6, SAP8 and SAP9 |
1(3.5) |
10 |
SAP1, SAP3 to SAP5, SAP7, SAP9 and SAP10 |
1(3.5) |
11 |
SAP1 to SAP4 and SAP8 to SAP10 |
1(3.5) |
12 |
SAP1 to SAP3, SAP6, SAP7, SAP9 and SAP10 |
1(3.5) |
13 |
SAP1, SAP2, SAP4, SAP6, SAP7, SAP9 and SAP10 |
1(3.5) |
14 |
SAP1, SAP2, SAP3, SAP4, SAP8 and SAP9 |
1(3.5) |
15 |
SAP1, SAP2, SAP3, SAP6, SAP8 and SAP9 |
1(3.5) |
16 |
SAP1, SAP3, SAP5, SAP7, SAP9 and SAP10 |
1(3.5) |
17 |
SAP1, SAP2, SAP4, SAP6, SAP9 and SAP10 |
1(3.5) |
18 |
SAP1, SAP2, SAP3, SAP7, SAP8 and SAP9 |
1(3.5) |
19 |
SAP1, SAP2, SAP4, SAP5, SAP9 and SAP10 |
1(3.5) |
20 |
SAP2, SAP4, SAP6, SAP8 and SAP9 |
1(3.5) |
21 |
SAP1, SAP5, SAP7, SAP8 and SAP9 |
1(3.5) |
22 |
SAP1, SAP2, SAP6, SAP7 and SAP8 |
1(3.5) |
23 |
SAP4, SAP6, SAP9 and SAP10 |
1(3.5) |
24 |
SAP1, SAP2, SAP4 and SAP9 |
1(3.5) |
25 |
SAP1, SAP2, SAP4 and SAP8 |
1(3.5) |
26 |
SAP1, SAP2, SAP3 and SAP8 |
1(3.5) |
27 |
SAP1, SAP2 and SAP9 |
1(3.5) |
28 |
SAP2, SAP6 and SAP9 |
1(3.5) |
SAP= Secreted Aspartyl Proteinase
It has been four decades since the detection of Sap activity in Candida albicans22. Ten SAP genes have been identified till date and different expressions have been observed in vivo23. In the midst of pregnancy, increase hormone levels and glycogen aggregation in the vagina lead to two folds rises in the frequency of VVC in antenatal women compared with non-pregnant women14. VVC is a noteworthy reason for morbidity in pregnancy which leads to preterm delivery, miscarriages, Candida chorioamnionitis and emotional stress15-16. VVC caused by the Candida albicans and Non-albicans Candida (NAC), is able to inhabit the human body without generating any signs of infection such as the asymptomatic state. Despite the fact that under immunocompromised circumstances that disturb the sensitive harmony among the host and commensal, a parasitic relationship may happen to bring about the advancement of disease17–19. A study showed that the increased Sap enzymes production in diabetic patients have a higher prevalence of VVC20. Prolonged antibiotic treatment, diabetes mellitus and chronic stress may alter the cellular immune responses which lead to increase the susceptibility of infection by Candida21. Proteolysis activity of SAP genes of Candida shows vital success in pathogenicity of this opportunistic fungus. Sap production is one of the tightly regulated and controlled processes; therefore, SAP genes have multiple functions for virulence in Candida. SAP1 to SAP3 is essential for mucosal infection whereas SAP4 to SAP6 is responsible for systemic infections that take part in evasion of host immune response, host tissue damage and adherence to the host cell. The Sap is able to breakdown the human proteins such as mucin, collagen, keratin as well as antibodies, cytokines, complements of immune component27.
There are many newly existing methods for in vitro identification of Sap enzyme yet it is exceptionally dubious when utilizing plate methods with protein-containing media. The pH range of Saps is 3 to 5.5. If there is alteration in pH, that enzyme gets denatured easily. Furthermore, media contains an inhibitory substance which may alter the production of proteinase by Candida species and in vitro detection of Sap becomes challenging and unreliable24-25. Kilic et al.26 reported that detection of Sap by in vivo and in vitro may be useful. Nevertheless, molecular detection of SAP genes is more explicit than Sap enzymes detection by plate methods.
The present study revealed that the more prevalent SAP genes in Candida species isolated from symptomatic women were SAP1 and SAP9 (100%) followed by SAP2 and SAP8 (87.5%). Whereas SAP1 (78.9%) was more prevalent SAP gene among Candida species isolated from asymptomatic women. Bassyouni et al.20 C. albicans was isolated from diabetic and non-diabetic women with VVC and investigated the presence of SAP1 to SAP8 genes. The author found that SAP1 and SAP2 were the most predominant detected genes. Another study done by Kalkanci et al.28 reported that the most prevalent SAP gene in vaginal isolates were SAP1 to SAP3. A study done by Monroy et al.29 observed that SAP4 to SAP6 genes were more frequently expressed and suggesting that SAP gene plays a significant role in the pathogenesis of the infections. Bernardis et al.30 reported that SAP1 and SAP2 are responsible for the development of the disease. Schaller et al.31 demonstrated that SAP1, SAP2 and SAP5 were predominant genes identified in vaginal infection; the author also stated that SAP2 expression is found in both infection and colonization while SAP1 expression is higher than SAP2 in vaginal infection. The results of the earlier studies suggest the role of SAP1 for the infection process in VVC and are in agreement with the present study.
In the present study, SAP4 was detected 81.2% in Candida isolated from symptomatic VVC which is higher than the asymptomatic isolates. It is reported that SAP4 is responsible for biofilm formation and prevent phagocytosis33. The result of the present study does not correlate with the study done by Nas et al.34 where SAP4 was detected 40%. In this study, the SAP6 and SAP7 were predominantly detected in isolates of sympto-matic women. The SAP6 associated with the SAP4 to SAP6 subfamily, help for the development of hyphae and invasion of host cells12,33. Despite the fact that SAP6 and SAP7 genes were associated with tissue damage in the initial phase of infection in vitro model of vaginal candidiasis based on “Reconstituted Human Vaginal Epithelium (RHVE)”31. In the current study, SAP8 was detected 87.5% in isolates from symptomatic VVC and 47.3% in isolates from asymptomatic VVC. The result of the present study was higher than the study reported by Naglik et al.12 the researcher distinguished that 75% of C. albicans express SAP8 gene in the vaginal-infected group while 32% in the colonized group. Schild et al.32 proposed that SAP9 and SAP10 are responsible for biofilm formation, adherence and invasion into host cells.
The frequency of SAP1 and SAP3 are more in vaginal infection when compared with oral infection, thus indicating that the differential expression of genes varies according to the stage of infection and the anatomical site35. A study done by Lian et al.36 observed that SAP1 and SAP3 genes were present only in vaginal infection. Lin et al.37 reported that SAP2 and SAP5 are the most common genes expressed in the vaginal mucosa. However, in the present study, SAP1 and SAP9 were predominantly observed in vaginal infection compared to vaginal colonization. In the present study, statistical analysis showed a significant difference (p =0.0213) for the SAP7 gene frequency among the Candida isolated from infection (81.2%) and colonization (36.8%), similarly there is a significant difference (p=0.0328) for the SAP8 gene frequency among the Candida isolated from infection (87.5%) and colonization (47.3%). This suggests probable participation of SAP7 and SAP8 associated with tissue damage in the initial phase of VVC12,31.
The SAP1 to SAP10 genes can be expressed by Candida in the vaginal mucosa, both in colonization processes and in infections, but differential expression of these genes is seen when looking at the transcript levels in isolates of carriers and of active VVC, besides a predominance of certain members of the SAP family during vaginal infection. Medeiros et al.38 did not observe the differences in the expression of SAP gene among C. albicans isolated from patients with VVC and colonization, suggesting that the capacity to express virulence factors is significant in the pathogenesis of VVC, yet it appears not to be essential for the change from colonization to disease. In the present study, we did not identify a statistical difference for the presence of SAP1 to SAP6, SAP9 and SAP10 genes in isolates from vaginal infection and colonization.
Candida isolated from symptomatic women and asymptomatic women exhibited distinctive patterns of distribution for SAP1 to SAP10 genes. There is a predominance of the SAP1 gene in Candida isolates from asymptomatic women whereas SAP1 and SAP9 genes in Candida isolates from symptomatic women. The statistical analysis showed a significant difference for the SAP7 and SAP8 gene frequency among the Candida isolated from symptomatic and asymptomatic women. The existence of an SAP gene family in Candida species noticeably delivers the fungus with a proficient and adaptable proteolytic framework that may demonstrate fundamental to its prosperity as a sharp pathogen. Proteinases encoded the SAP genes as an exceptionally flexible and multi-functional harmful gene family of Candida. However, we did not identify a statistical difference between the presence of SAP1 to SAP6, SAP9 and SAP10 genes in isolates from vaginal infection and colonization. Moreover, this suggests that other factors that control the genetic fungal expression and characteristics inherent in the host are likely to influence the infection/colonization process. Though, it seems that we all know an excellent deal concerning the Candida SAP genes, further investigations involving a larger number of samples will yield a better conclusion and number of essential questions can be addressed in upcoming studies.
Acknowledgments
We are grateful to SRM Medical College Hospital and Research Centre to avail us with reagents, equipment’s and supporting our research work.
Conflict Of Interest
The authors declare that there is no conflict of interest.
Authors’ Contribution
All authors have made substantial, direct and intellectual contribution to the work and approved it for publication.
Funding
None.
Data Availability
Not Applicable.
Ethics Statement
The study was apporved from the Institutional Ethical Committee (IEC, 1090/ IEC/2017).
- Bhattacharjee P. Epidemiology and antifungal susceptibility of Candida species in a tertiary care hospital, Kolkata, India. Curr. Med. Mycol., 2016; 2(2): 20-27.
- Foxman B., Marsh J.V., Gillespie B. and Sobel J.D. Frequency and Response to Vaginal Symptoms among White and African American Women: Results of a Random Digit Dialing Survey. Journal of Women’s Health, 1998; 7(9): 1167-1174.
- Sobel J.D., Faro S., Force R.W., Foxman B., Ledger W.J., Nyirsejy P.R. et al. Vulvovaginal Candidiasis: Epidemiologic, Diagnostic, and Therapeutic Considerations. American Journal of Obstet- tricks and Gynecology, 1998; 178(2): 203- 211.
- Sobel J.D. Vaginitis. New England Journal of Medicine, 1997; 26: 1896-1903.
- Viudes A., Peman J., Canton E., Ubeda P., Lopez-Ribot J.L., Gobernado M. Candidemia at a tertiary-care hospital: epidemiology, treatment, clinical outcome and risk factors for death. Eur. J. Clin. Microbiol Infect. Dis, 2002; 21(4): 767–774.
- Pfaller M.A. Antifungal drug resistance: mechanisms, epidemiology, and consequences for treatment. Am. J. Med., 2012; 125(1): S3–S13.
- Odds F.C. Candida and candidiasis. 2nd ed. London: Balliere Tindall, 1988.; 30(5): 382-383
- Ruchel R., De Bernardis F., Ray T.L., Sullivan P.A., Cole G.T. Candida acid proteinases. Journal of Med. Vet. Mycol., 1992; 30(1): 123–132.
- Finlay B.B., Falkow S. Common themes in microbial pathogenicity. Microbiol. Rev., 1989; 53(2):210-30.
- Gilfillan G.D., Sullivan D.J., Haunes K., Parkinson T., Coleman D.C., Gow N.A. Candida dubliniensis: phylogeny and putative virulence factors. Microbiology, 1998; 144: 829-38.
- Monod M., Togni G., Hube B., Sangland D. Multiplicity of genes encoding secreted aspartic proteinases in Candida species. Mol. Microbiol., 1994; 13(2): 357-68.
- Naglik J.R., Challacombe S.J., Hube B. Candida albicans secreted aspartyl proteinases in virulence and pathogenesis. Microbiol. Mol. Biol. Rev., 2003; 67(3): 400-28.
- Miyasaki S.H., White T.C., and Agabian N. A fourth secreted aspartyl proteinase gene (SAP4) and a CARE2 repetitive element are located upstream of the SAP1 gene in Candida albicans. J. Bacteriol., 1994; 176(6): 1702–1710.
- Guideline vulvovaginal candidiasis of the German Society for Gynecology and Obstetrics, the Working Group for Infections and Infect immunology in Gynecology and Obstetrics, the German Society of Dermatology, the Board of German Dermatologists and the German Speaking Mycological Society. Mycoses, 2012; 55(3): 1-13.
- Kiss H., Petricevic L., Husslein P. Prospective randomised controlled trial of an infection screening programme to reduce the rate of preterm delivery. BMJ, 2004; 14: 329-371.
- Arul Anne Rose S., Sony P, Iyanar K. Prevalence of antenatal vulvovaginal candidiasis: our experience, 2017; 6(2): 443-446.
- Cassone A. Vulvovaginal Candida albicans infections: pathogenesis, immunity and vaccine prospects. BJOG, 2015; 122(6): 785–94.
- Sobel J.D. Recurrent vulvovaginal candidiasis. Am J. Obstet. Gynecol., 2016; 214(1): 15-21.
- Sobel J.D. Vulvovaginal candidosis. Lancet, 2007; 369: 1961–71.
- Bassyouni R.H., Wegdan A.A., Abdelmoneim A., Said W, AboElnaga F. Phospholipase and aspartyl proteinase activities of Candida species causing vulvovaginal candidiasis in patients with type 2 diabetes mellitus. J. Microbiol. Biotechnol., 2015; 25(10):1734–1741.
- Bonfim-Mendonca P.S., Ratti B.A., Godoy J.S., Negri M., Lima N.C., Fiorini A. et al. b-glucan induces reactive oxygen species production in human neutrophils to improve the killing of Candida albicans and Candida glabrata isolates from vulvovaginal candidiasis. Plos One, 2014; 9(9): e107805.
- Yamaguchi H., Iwata K. In vitro and in vivo protein synthesis in Candida albicans. Sabouraudia, 1970; 8(3): 177–188.
- Hube B., Naglik J. Candida albicans proteinases: resolving the mystery of a gene family. Microbiology, 2001; 147: 1997–2005.
- Dostal J. et al. Simple method for screening Candida species isolates for the presence of secreted proteinases: a tool for the prediction of successful inhibitory treatment. J. Clin. Microbiol. 2003; 41(2): 712–716.
- Cassone A., De Bernardis F., Mondello F., Ceddia T., Agatensi L. Evidence for a correlation between proteinase secretion and vulvovaginal candidosis, J. Infect. Dis., 1987; 156(5): 777–783.
- Kilic N., Kustimur S., Arslan S., Aldemir H. Fluorometric determination of acidic proteinase activity in vulvovaginal candidosis. Mycoses, 1996; 39(9): 347–351.
- Newport G. and Agabian N. KEX2 Influences Candida albicans Proteinase Secretion and Hyphal Formation. The Journal of Biological Chemistry, 1997; 272(46): 28954-28961.
- Kalkanci A., Bozdayi G., Birib A., Kustimur S. Distribution of secreted aspartyl proteinases using a polymerase chain reaction assay with SAP specific primers in Candida albicans isolates. Folia Microbiol., 2005; 50(5): 409-13.
- Monroy-Pיrez E., Paniagua-Contreras G., Vaca-Paniagua F., Negrete-Abascal E., Vaca S. SAP expression in Candida albicans strains isolated from Mexican patients with vaginal candidosis. Int. J. Clin. Med., 2013; 4(1): 25-3.
- Bernardis F., A. Cassone, J. Sturtevant and R. Calderone. Expression of Candida albicans SAP1 and SAP2 in Experimental Vaginitis. Infection and Immunity, 1995; 63(5): 1887–1892.
- Schaller M., Bein M., Korting C.H., Baur S., Hamm G., Monod M. et al. The Secreted Aspartyl Proteinases Sap1 and Sap2 Cause Tissue Damage in an in Vitro Model of Vaginal Candidiasis Based on Reconstituted Human Vaginal Epithelium. Infection and Immunity, 2003; 71(6): 3227-3234.
- Schild L., Heyken A., De Groot P.W. et al. Proteolytic cleavage of covalently linked cell wall proteins by Candida albicans Sap9 and Sap10. Eukaryot Cell, 2011; 10(1): 98-109.
- Winter M.B., Salcedo E.C., Lohse M.B., et al. Global identification of biofilm-specific proteolysis in Candida albicans. M. Bio., 2016; 7(5): e01514-16.
- Nas T., Kalkanci A., Ozkan S. Expression of ALS1 and SAP4 genes in Candida albicans strains isolated from women with vaginitis. Folia Microbiol., 2008; 53(2): 179-83.
- Naglik J.R., Rodgers C.A., Shirlaw P.J., et al. Differential expression of Candida albicans secreted aspartyl proteinase and phospholipase B genes in human’s correlates with active oral and vaginal infections. JID, 2003; 188(3): 469-79.
- Lian C.H., Liu W.D. Differential expression of Candida albicans secreted aspartyl proteinase in human vulvovaginal candidiasis. Mycoses, 2007; 50(5): 383-90.
- Lin N., Feng J., Tu Y., Feng A. Expression of Candida albicans secreted aspartyl proteinase in acute vaginal candidiasis. J. Huazhong Univ. Sci. Technol. Med. Sci., 2007; 27(3): 333-5.
- Medeiros M.A.P., Melo A.P.V., Sousa A. et al. Characterization of virulence factors of vaginal and anal isolates of Candida albicans sequentially obtained from patients with vulvovaginal candidiasis in north-east Brazil. J. Mycol. Med., 2017; 27(4): 567-72.
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


