ISSN: 0973-7510
E-ISSN: 2581-690X
Panax vietnamensis Ha et Grushv. is a high-value endemic ginseng species and a rich source of biologically active ginsenosides. However, low productivity and high susceptibility to phytopathogens limit supply relative to demand. Here, we evaluated the efficacy of bacterial strain VCCM 22715 against phytopathogenic fungi and assessed its effects on the growth of P. vietnamensis. Among 61 strains, VCCM 22715 showed the strongest activity against Rhizoctonia solani, inhibiting mycelial growth by 80.1 ± 5.7%. An ethyl acetate extract also inhibited R. solani (52.8 ± 5.3%), Magnaporthe oryzae (73.5 ± 0.3%), and Fusarium solani (52.8 ± 5.3%). Based on phenotypic characterization and molecular analyses, the strain was identified as Streptomyces albus VCCM 22715. Notably, the strain produced 24.3 ± 2.5 µg/mL indole-3-acetic acid (IAA) and significantly increased plant height, root length, fresh weight, and dry weight of P. vietnamensis (p < 0.05). Whole-genome analysis revealed an 8.0 Mb linear chromosome with 73% GC content, 6,796 protein-coding sequences, and 16 secondary metabolite biosynthetic gene clusters. Key genes implicated in IAA biosynthesis were also detected. Collectively, these results indicate that S. albus VCCM 22715 is a promising biocontrol and plant growth-promoting agent for field-cultivated ginseng.
Antifungal activity, Indole-3-acetic acid, Panax vietnamensis, Streptomyces albus, Whole-genome analysis
Medicinal crop production has traditionally relied heavily on agricultural chemicals, including inorganic fertilizers and pesticides.1 The emergence of pest resistance to synthetic agrochemicals has contributed to antibiotic resistance in pathogens and has increased the need for new pesticides.2,3 Consequently, alternatives to conventional agrochemicals are increasingly important for achieving sustainable agriculture.
One promising approach is the use of biostimulants derived from plant growth-promoting bacteria (PGPB). PGPB, including Pseudomonas, Bacillus, Streptomyces, and Azospirillum, can increase crop productivity, enhance resilience to abiotic stresses and phytopathogenic microbes, and reduce reliance on agrochemicals.4 Among these, Streptomyces spp. are prolific producers of bioactive metabolites and account for approximately 75% of antibiotics and antifungal compounds on the global market.5 Streptomyces lydicus and Streptomyces griseoviridis have been commercialized as the biofungicides Actinovate® and Mycostop®, respectively.6 Strawberry fruit rot caused by Botrytis cinerea, Mucor hiemalis, Rhizopus stolonifer, and Sclerotinia sclerotiorum is suppressed by reveromycins A and B produced by Streptomyces sp. 3-10.6 In addition, Streptomyces spp. can promote plant growth by traits such as nitrogen fixation, IAA production, and phosphorus and potassium solubilization, thereby increasing plant biomass and length.7,8 For example, Streptomyces albidoflavus St-220 promotes the growth of Salvia miltiorrhiza seedlings under greenhouse conditions by solubilizing inorganic phosphate, fixing nitrogen, and producing IAA.9 Accordingly, the discovery of new Streptomyces strains with plant growth-promoting and antifungal activities tailored to specific medicinal plants remains an active area of research and application.
Panax vietnamensis Ha et Grushv. is a valuable endemic species first discovered in Quang Nam and Kon Tum provinces, Vietnam.10 Overharvesting, slow growth, and poor regeneration have pushed wild populations toward extinction.11,12 In parallel, excessive use of chemical fertilizers and pesticides has raised concerns regarding phytochemical integrity and the accumulation of toxic compounds. Here, Streptomyces albus VCCM 22715 was screened for antifungal activity and identified to the species level. Broad-spectrum antifungal activity of its ethyl acetate extract was also assessed, and the effects of the strain on P. vietnamensis growth were evaluated under in vivo conditions. To elucidate the genetic basis of antifungal and plant growth-promoting properties, we performed genomic analysis.
Screening of bacterial strains for antifungal activity
Sixty-one bacterial strains isolated from mangrove plants, together with the phytopathogenic fungus Rhizoctonia solani, were obtained from the VAST-Culture Collection of Microorganisms (Institute of Biology, Vietnam Academy of Science and Technology). Antifungal activity was screened using a plate confrontation assay as described previously.13 Briefly, a 5 mm mycelial plug of R. solani was placed at the center of potato dextrose agar (PDA) plates (HiMedia, India). Test strains were inoculated at four equidistant points, each 25 mm from the plate center, and incubated for 5-7 days. Plates containing only R. solani served as the control. The percentage of fungal growth inhibition (GI) was calculated by comparing mycelial diameters in control and treated plates.13
Morphological characterization and 16S rRNA gene sequence analysis
The most active strain was cultured on ISP2 agar at 30 °C for 9 days to assess colony morphology, growth characteristics, pigmentation, and color as described previously.14 Genomic DNA from strain VCCM 22715 was extracted using the G-spin™ Total DNA Extraction Mini Kit (Intron Bio). Molecular identification was performed by PCR amplification of the 16S rRNA gene, followed by agarose gel electrophoresis, purification, and Sanger sequencing. The sequence was trimmed using BioEdit v7.2.5 and compared with the National Center for Biotechnology Information (NCBI) database using BLAST. A phylogenetic tree was constructed in MEGA v11.0 using the neighbor-joining approach with maximum-likelihood-based analysis.
Preparation of ethyl acetate extract and evaluation of antifungal activity
VCCM 22715 was cultivated in potato dextrose broth (PDB; HiMedia, India) for 3 days at 30 °C as a seed culture and then inoculated into five 1,000 mL flasks, each containing 250 mL fresh PDB. After incubation for 8 days at 30 °C, cultures were pooled, centrifuged at 8,000 rpm for 20 min, and extracted three times with ethyl acetate. The organic phase was evaporated at 50 °C to obtain a brown crude extract. Immediately before each assay, fresh stock solutions were prepared and incorporated into PDA to final concentrations of 0, 100, or 150 µg/mL. A 5 mm mycelial plug of Magnaporthe oryzae, Fusarium solani, or R. solani was placed at the center of each plate. Plates containing 5% (v/v) dimethyl sulfoxide (DMSO) served as the solvent control. After 7 days at 30 °C, colony diameters were measured and growth inhibition (GI %) was calculated as described previously.13 All assays were performed in triplicate, and data are presented as mean ± standard deviation (SD).
Determination of indole-3-acetic acid production
IAA production by VCCM 22715 was quantified as described previously.3 The strain was grown in ISP2 medium supplemented with 100 µg/mL tryptophan and incubated at 30 °C for 3 days (triplicate cultures). Cultures were centrifuged at 10,000 rpm for 10 min, and 1.0 mL supernatant was mixed with 1.0 mL Salkowski reagent. The mixture was incubated in the dark at room temperature for 30 min, and absorbance at 530 nm was measured. IAA concentration was calculated using the standard curve equation (y = 0.0065x + 0.0033), generated from serial IAA standards (0, 5, 10, 20, 40, and 80 µg/mL).
Pot experiment
Pot experiments were performed in a greenhouse in Yen Bai Province, Vietnam, where the mean annual temperature in 2024 was approximately 19.6 °C. Each plastic pot contained approximately 1 kg UV-sterilized soil (neutral pH). Six-month-old ginseng plants were pre-cultivated in pots for 1 month before treatment. Treatments included non-inoculated controls and pots inoculated with S. albus VCCM 22715. For inoculation, S. albus VCCM 22715 was grown on PDA for 7 days at 30 °C to obtain spores, which were applied to each pot to a final density of 104 CFU/g soil. Six plants were treated weekly for 3 weeks with the VCCM 22715 spore suspension, whereas six control pots received sterile water. After 6 months, plants were harvested to assess growth parameters, including plant height, root length, fresh weight, and dry weight of each P. vietnamensis plant. Experiments were performed in triplicate, and data are presented as mean ± SD. Statistical analyses were conducted using GraphPad Prism 9.
Whole-genome sequencing and genome mining
Genomic DNA from VCCM 22715 was extracted using the G-spin™ Total DNA Extraction Mini Kit (Intron Bio), followed by quality control and Illumina sequencing. Raw reads were trimmed and quality-assessed using Trimmomatic v3.0 and FastQC v0.12.1. Reads were assembled de novo with SPAdes v3.15.5 using default parameters. Genome annotation was performed using the NCBI Prokaryotic Genome Annotation Pipeline (PGAP) and RASTtk (Rapid Annotation using Subsystem Technology).15 The whole-genome sequence was deposited in DDBJ/ENA/GenBank under accession number JBPQDV010000000. Secondary metabolite biosynthetic gene clusters (BGCs) were predicted using antiSMASH v6.0.16 Species-level assignment was performed using the Type (Strain) Genome Server (TYGS: https://tygs.dsmz.de/) to construct a phylogenomic tree with default parameters and to calculate digital DNA-DNA hybridization (dDDH) values. Genes implicated in IAA biosynthesis were predicted using RASTtk with thresholds of >30% identity and >50% coverage.
Screening and identification of a bacterial strain with potent antifungal activity
A total of 61 actinomycete strains from a previous study were screened for antifungal activity against R. solani using a dual-culture assay. Among the strains tested, VCCM 22715 showed the strongest inhibition of R. solani mycelial growth (80.1 ± 5.7%). Therefore, VCCM 22715 was selected for further analyses.
VCCM 22715 grew well on ISP2 agar and formed colonies with dry, rough surfaces (Figure 1A). After 10 days, the vegetative mycelium developed white aerial hyphae and a spore mass (Figure 1B), and no pigment was observed on any of the tested media. These phenotypic characteristics were consistent with assignment to the genus Streptomyces.
For species-level identification, a 1,536-bp partial 16S rRNA gene sequence from VCCM 22715 was amplified and sequenced. VCCM 22715 showed 99.7% sequence similarity to S. albus NRRL B-1811T and S. albus NBRC 15415T. Phylogenetic analysis based on 16S rDNA sequences further showed that VCCM 22715 clustered with S. albus, with S. albus NRRL B-1811T and S. albus NBRC 15415T as the closest relatives (Figure 1C). Accordingly, the strain was designated S. albus VCCM 22715.
Figure 1. Identification of the strain VCCM 22715. Colonies grown on ISP2 agar plate (A) and aerial hyphae under phase contrast microscopy (B) of the strain VCCM 22715. (C) Phylogenetic tree built from 16S rRNA sequence of the VCCM 22715 and its related type strains
Antifungal potential of the crude extract from S. albus VCCM 22715
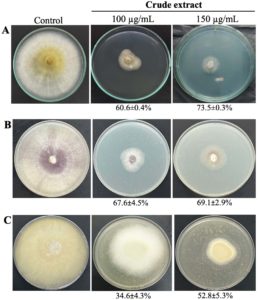
To determine whether S. albus VCCM 22715 exhibits broad antifungal activity, we tested its crude extract against three phytopathogenic fungi: Magnaporthe oryzae, Fusarium solani, and Rhizoctonia solani. The crude extract strongly inhibited mycelial growth of all three fungi, with inhibition ranging from 52.8% to 73.5% (Figure 2). At 150 µg/mL, the highest inhibition was observed for M. oryzae (73.5 ± 0.3%), followed by F. solani (69.1 ± 2.9%) and R. solani (52.8 ± 5.3%).
Figure 2. Antifungal activity of the VCCM 22715 extract against Magnaporthe oryzae (A), Fusarium solani (B), and Rhizoctonia solani (C)
Streptomyces spp. can suppress diverse phytopathogenic fungi through the production of secondary metabolites.17 For example, Fusarium oxysporum f. sp. cubense Tropical Race 4, Curvularia fallax, Colletotrichum gloeosporioides, Colletotrichum acutatum, and Fusarium graminearum are strongly inhibited by Streptomyces sp. YYS-7, with mycelial growth inhibition ranging from 48.6%-80.9%.18 In addition, S. parvulus VRR3 isolated from green gram exhibits biocontrol activity against F. solani under laboratory and greenhouse conditions.19 These reports are consistent with our findings. Biocontrol mechanisms against phytopathogenic fungi can include antimicrobial metabolites, nutrient competition, and antibiosis. However, the superior performance of VCCM 22715 may also reflect strain-specific genomic features.
In our previous study, S. albus RC2 showed strong activity against Lasiodiplodia theobromae, Fusarium fujikuroi, and Scopulariopsis gossypii, which cause serious crop damage in Vietnam.13 However, S. albus RC2 showed no inhibitory activity against R. solani in our preliminary screening. This observation supports the view that the antifungal potential of Streptomyces spp. depends on both the strain and the target phytopathogen.17 Collectively, these results indicate that VCCM 22715 is a promising candidate for improving plant growth and controlling disease as a biocontrol agent. However, comprehensive structural elucidation of secondary metabolites produced by VCCM 22715 is required for further studies.
IAA production and effects of VCCM 22715 inoculation on P. vietnamensis growth parameters
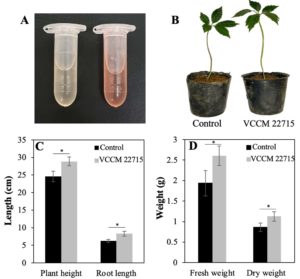
As IAA regulates diverse plant physiological processes,20 we evaluated IAA production by VCCM 22715. In the presence of 100 µg/mL tryptophan, VCCM 22715 produced 24.3 ± 2.5 µg/mL IAA after 3 days at 30 °C (Figure 3A). Multiple Streptomyces spp., including S. violaceus, S. albidoflavus, S. scabies, S. coelicolor, S. albus, and S. lividans, secrete IAA, with yields varying by medium composition and tryptophan supplementation.20 Notably, S. albidoflavus St-220 produces 30.4 µg/mL IAA in Gause medium containing 500 µg/mL tryptophan, which is higher than that reported for S. albus RC2.9 Together, these findings support the role of tryptophan as an IAA precursor and suggest that VCCM 22715 likely synthesizes IAA via a tryptophan-dependent pathway.
In pot experiments, VCCM 22715 treatment significantly enhanced growth relative to the control (Figure 3B). Plant height and root length increased by 17.3% and 33.5%, respectively (p < 0.05) (Figure 3C). Fresh weight and dry weight increased by 34.6% and 30.3%, respectively, in the treated group compared with the control (Figure 3D). These results indicate that VCCM 22715 strongly stimulates root elongation and development, likely through direct or indirect effects of IAA. Consistent with our findings, S. albidoflavus St-220 increases root fresh weight, total fresh weight, total dry weight, and root dry weight in Salvia miltiorrhiza.9 Similarly, S. fradiae NKZ-259 increases multiple growth parameters in tomato plants under greenhouse conditions.8
Figure 3. Effects of the VCCM 22715 on growth parameters of Panax vietnamensis plants. (A) IAA production by the VCCM 22715 in the ISP2 medium supplemented with 100 µg/mL tryptophan. (B) Representative images of ginseng plants inoculated with and without the VCCM 22715. Plant height, root weight (C) and fresh and dry weights (D) of ginseng in control group and VCCM 22715-treated group. The level *P < 0.05 was statistically considered significant using a Student’s unpaired two-tailed t-test by the GraphPad Prism 9 software
Wild-simulated cultivation of P. vietnamensis typically requires at least 6 years to reach harvest maturity.21 Extended growth cycles, disease-related replanting, and labor-intensive practices further complicate cultivation. Growth promotion of Panax ginseng by bacterial strains such as Bacillus velezensis and Arthrobacter nicotinovorans has been reported recently.12,22 B. velezensis NT35 promotes ginseng seed germination and controls ginseng rusty root rot under in vivo conditions.12 Treatment with A. nicotinovorans JI39 increases plant height, length, and weight of P. ginseng after 150 days in the field.22 To our knowledge, this is the first study demonstrating growth-promoting potential of Streptomyces spp. in P. vietnamensis, supporting its potential utility for sustainable ginseng cultivation. Nevertheless, the basis for developing VCCM 22715 as an effective biocontrol agent remains unresolved and warrants further investigation.
Genome features of S. albus VCCM 22715
To better understand the mechanisms underlying biocontrol and plant growth promotion, the genome of VCCM 22715 was sequenced using the Illumina platform. Sequencing generated 3,392,043 reads and 511,502,970 bases, with an N50 length of 279,269 bp. The draft genome comprised an 8.0 Mb linear chromosome with 85X coverage and 73.0% GC content and was predicted to contain 6,796 coding sequences (CDSs), 17S rRNA genes, 65 tRNA genes, and 3 ncRNA genes (Table 1). The estimated genome completeness was 98.22%. The chromosome sequence of VCCM 22715 was deposited in GenBank under accession number JBPQDV010000000. The genomic characteristics of VCCM 22715 were similar to those of S. albus DSM 40763T and S. albus N11-50.
Table (1):
Genome characteristics of the VCCM 22715 and its related strains
Attribute |
S. albus VCCM 22715 |
S. albus N11-50 |
S. albus DSM 40763T |
|---|---|---|---|
Genome size (Mb) |
8.0 |
8.29 |
8.09 |
GC content (%) |
73.0 |
72.7 |
72.6 |
rRNA genes |
17 |
18 |
19 |
tRNA genes |
65 |
62 |
61 |
ncRNA genes |
3 |
3 |
3 |
CDSs |
6796 |
7010 |
6888 |
Predicted genes |
6862 |
7093 |
6971 |
GenBank accession |
JBPQDV010000000 |
NZ_BNEJ00000000 |
NZ_RCIY00000000 |
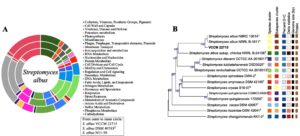
Genome annotation using RAST indicated subsystem and non-subsystem coverage of 18% and 82%, respectively. The top three subsystem categories were Amino Acids and Derivatives (330 genes), Carbohydrate Metabolism (266 genes), and Protein Metabolism (266 genes). Additional categories included Cofactors, Vitamins, Prosthetic Groups, and Pigments (189 genes) and Fatty Acids, Lipids, and Isoprenoids (133 genes). Compared with S. albus DSM 40763T, VCCM 22715 contained higher numbers of genes associated with Cell Wall and Capsule, Phages/Prophages/Transposable Elements/Plasmids, Membrane Transport, and Metabolism of Aromatic Compounds (Figure 4A). These features may contribute to energy conversion efficiency and improved adaptation to the host plant.
Consistent with 16S rRNA based identification, phylogenomic analysis using TYGS showed that S. albus VCCM 22715 is closely related to S. albus NRRL B-1811T and S. albus NBRC 13014T, with gene-content formula d4 values of 89.2% and 89.3%, respectively (Figure 4B). These results support assignment of the studied strain to S. albus.
Figure 4. (A) RAST annotations of S. albus genomes and (B) Phylogenomic tree constructed for VCCM 22715 and the type strains of Streptomyces data set on TYGS. Tree inferred with FastME 2.1.6.1 from GBDP distances calculated from genome sequences
Secondary metabolite biosynthetic gene clusters in S. albus VCCM 22715
AntiSMASH analysis identified 34 putative BGCs on the VCCM 22715 chromosome, of which 16 showed >21% similarity to known clusters. These included three nonribosomal peptide synthetase (NRPS) clusters, two type I polyketide synthase (T1PKS) clusters, two siderophore clusters, two lanthipeptide clusters, two terpene clusters, and five clusters associated with other secondary metabolites (Table 2). Five BGCs showed >96% similarity to clusters for tambjamine BE-18591, legonoxamine A, ectoine, A-factor, and geosmin. In contrast, the remaining 11 BGCs showed 21%-81% similarity, suggesting potential for production of novel metabolites by S. albus VCCM 22715.
Table (2):
Predicted secondary metabolite biosynthetic gene clusters in S. albus VCCM 22715
Region |
Size (bp) |
Synthetase type |
Similar known BGC (similarity) |
Biological activity |
|---|---|---|---|---|
1.1 |
35.020 |
Prodigiosin |
Tambjamine BE-18591 (96%) |
Antibacteria, antifungi, antitumor |
1.3 |
8.834 |
NI-siderophore |
Legonoxamine A (100%) |
Anticancer |
2.4 |
41.239 |
Nucleoside |
Pseudouridimycin (68%) |
Antibacteria |
2.5 |
22.715 |
Lanthipeptide |
SapB (75%) |
Morphogenetic peptide |
3.1 |
118.266 |
Lanthipeptide |
Xantholipin (81%) |
Antitumor, antibacteria |
3.2 |
10.405 |
Ectoine |
Ectoine (100%) |
Antioxidant |
4.1 |
26.705 |
Terpene |
Hopene (61%) |
Antimicrobial |
7.2 |
48.590 |
NRPS |
Griseobactin (53%) |
Iron chelation, anticancer |
9.1 |
69.737 |
T1PKS |
Ibomycin (58%) |
Antifungi |
15.3 |
58.331 |
NRPS |
Coelibactin (54%) |
– |
16.1 |
24.462 |
Terpene |
Isorenieratene (45%) |
– |
17.1 |
10.345 |
Butyrolactone |
A-factor (100%) |
– |
26.1 |
33.306 |
NI-siderophore |
Kinamycin (21%) |
Antibacteria |
39.1 |
32.093 |
NRPS |
Dudomycin A (65%) |
Antibacteria |
50.1 |
14.850 |
Terpene |
Geosmin (100%) |
– |
57.1 |
8.407 |
T1PKS |
Griseochelin (53%) |
Antibacteria |
With respect to bioactivity, four BGCs were associated with the biosynthesis of compounds reported to have anticancer activity, including tambjamine BE-18591, legonoxamine A, xantholipin, and griseobactin. BGCs linked to pseudouridimycin, hopene, kinamycin, dudomycin A, and griseochelin were associated with antibacterial activity. Notably, only two BGCs, tambjamine BE-18591 and ibomycin, were linked to antifungal activity in S. albus VCCM 22715. However, the ibomycin BGC can be silent under standard conditions; for example, co-cultivation of Streptomyces sp. WAC2288 with Cryptococcus neoformans induces expression of genes involved in ibomycin biosynthesis.23 Tambjamine produced by Pseudoalteromonas tunicata exhibits antifungal activity against Candida albicans and Malassezia furfur.24 Microbial co-culture can activate silent BGCs in Streptomyces strains by inducing defensive and nutritional metabolite production, thereby promoting synthesis of antifungal compounds.25 Accordingly, exposure to phytopathogenic fungi may induce VCCM 22715 to express known or previously uncharacterized antifungal BGCs, which warrants further study.
Identification of genes associated with IAA biosynthesis
To support the observed IAA production, we mined the VCCM 22715 genome for genes involved in IAA biosynthesis. Genes encoding tryptophan monooxygenase IaaM (orf_1879) and indole-3-acetamide hydrolase IaaH (orf_1880) were identified, consistent with an indole-3-acetamide pathway. In contrast, genes required for the indole-3-acetaldoxime/indole-3-acetonitrile pathway and the indole-3-pyruvate pathway were not detected. In Streptomyces sp. AgN23, the complete biosynthetic routes for multiple pathways leading to IAA production have been reported.7 In pathogenic Streptomyces spp. that cause scab disease, the gene encoding IaaM, which is required for IAA biosynthesis, is absent.26 Collectively, these results suggest that S. albus VCCM 22715 produces IAA primarily via a single pathway.
In the present study, the bacterial strain exhibiting the strongest antifungal activity against R. solani was identified as S. albus VCCM 22715. The ethyl acetate extract of VCCM 22715 showed broad-spectrum antifungal activity against M. oryzae, F. solani, and R. solani. Pot experiments further demonstrated that S. albus VCCM 22715 significantly enhanced the growth of P. vietnamensis, consistent with the IAA concentration measured in the cell-free supernatant. Genome analysis indicated that VCCM 22715 has substantial potential to produce antifungal metabolites and IAA. Collectively, these findings support the use of this strain as a bioinoculant for sustainable agriculture and plant nutrient management. Future studies should evaluate its biocontrol efficacy and elucidate the underlying mechanisms under pot and field conditions.
ACKNOWLEDGMENTS
The authors would like to acknowledge the support of the Key Laboratory of Bioprocess and Biochemical Synthesis Engineering, Institute of Biology, Vietnam Academy of Science and Technology.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This research was funded by the Vietnam Academy of Science and Technology, grant number NVCC08.02/24-25.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
This article does not contain any studies on human participants or animals performed by any of the authors.
- Nuruzzaman M, Bahar MM, Naidu R. Diffuse soil pollution from agriculture: Impacts and remediation. Sci Total Environ. 2025;962:178398.
Crossref - Tyagi A, Lama Tamang T, Kashtoh H, et al. A review on biocontrol agents as sustainable approach for crop disease management: applications, production, and future perspectives. Horticulturae. 2024;10(8):805.
Crossref - Ahmed W, Zhou G, Yang J, et al. Bacillus amyloliquefaciens WS-10 as a potential plant growth-promoter and biocontrol agent for bacterial wilt disease of flue-cured tobacco. Egypt J Biol Pest Control. 2022;32(1):25.
Crossref - Jakubowska Z, Gradowski M, Dobrzynski J. Role of plant growth-promoting bacteria (PGPB) in enhancing phenolic compounds biosynthesis and its relevance to abiotic stress tolerance in plants: a review. Antonie van Leeuwenhoek. 2025;118(9):123.
Crossref - Ghareeb A, Fouda A, Kishk RM, El Kazzaz WM. Unlocking the therapeutic potential of bioactive exopolysaccharide produced by marine actinobacterium Streptomyces vinaceusdrappus AMG31: A novel approach to drug development. Int J Biol Macromol. 2024;276(Pt 2):133861.
Crossref - Lyu A, Liu H, Che H, et al. Reveromycins A and B from Streptomyces sp. 3-10: antifungal activity against plant pathogenic fungi in vitro and in a strawberry food model system. Front Microbiol 2017;8:550.
Crossref - Gayrard D, Nicolle C, Veyssiere M, et al. Genome sequence of the Streptomyces strain AgN23 revealed expansion and acquisition of gene repertoires potentially involved in biocontrol activity and rhizosphere colonization. PhytoFrontiers™. 2023;3(3):535-547.
Crossref - Myo EM, Ge B, Ma J, et al. Indole-3-acetic acid production by Streptomyces fradiae NKZ-259 and its formulation to enhance plant growth. BMC Microbiol. 2019;19(1):155.
Crossref - Du Y, Wang T, Jiang J, et al. Biological control and plant growth promotion properties of Streptomyces albidoflavus St-220 isolated from Salvia miltiorrhiza rhizosphere. Front Plant Sci. 2022;13:976813.
Crossref - Le HTT, Nguyen LN, Pham HLB, et al. Target capture reveals the complex origin of Vietnamese ginseng. Front Plant Sci. 2022;13:2022.
Crossref - Titova MV, Lunkova MK, Tyurina TM, et al. Suspension cell cultures of Panax vietnamensis as a biotechnological source of ginsenosides: growth, cytology, and ginsenoside profile assessment. Front Plant Sci. 2024;15:1349494.
Crossref - Li X, Wang J, Shen H, et al. Biocontrol and growth promotion potential of Bacillus velezensis NT35 on Panax ginseng based on the multifunctional effect. Front Microbiol. 2024;15:2024.
Crossref - Quach NT, Vu THN, Nguyen TTA, et al. Metabolic and genomic analysis deciphering biocontrol potential of endophytic Streptomyces albus RC2 against crop pathogenic fungi. Braz J Microbiol. 2023;54(4):2617-2626.
Crossref - Quach NT, Vu THN, Bui TL, et al. Genomic and physiological traits provide insights into ecological niche adaptations of mangrove endophytic Streptomyces parvulus VCCM 22513. Ann Microbiol. 2022;72(1):27.
Crossref - Overbeek R, Olson R, Pusch GD, et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 2013;42(D1): 206-214.
Crossref - Blin K, Shaw S, Vader L, et al. antiSMASH 8.0: extended gene cluster detection capabilities and analyses of chemistry, enzymology, and regulation. Nucleic Acids Res. 2025;53(1):32-38.
Crossref - Khan S, Srivastava S, Karnwal A, Malik T. Streptomyces as a promising biological control agents for plant pathogens. Review. Front Microbiol. 2023;14:2023.
Crossref - Wei Y, Zhao Y, Zhou D, et al. A newly isolated Streptomyces sp. YYS-7 with a broad-spectrum antifungal activity improves the banana plant resistance to Fusarium oxysporum f. sp. cubense Tropical Race 4. Front Microbiol. 2020;11:1712.
Crossref - Kadaikunnan S, Alharbi NS, Khaled JM, Alobaidi AS. Biocontrol property of Streptomyces parvulus VRR3 in green gram plant (Vigna radiata L.) against Fusarium solani in greenhouse. Physiol Mol Plant Pathol. 2023;128:102128.
Crossref - Devi S, Sharma M, Manhas RK. Investigating the plant growth promoting and biocontrol potentiality of endophytic Streptomyces sp. SP5 against early blight in Solanum lycopersicum seedlings. BMC Microbiol. 2022;22(1):285.
Crossref - Trong TT, Truong D-H, Nguyen HC, et al. Biomass accumulation of Panax vietnamensis in cell suspension cultures varies with addition of plant growth regulators and organic additives. Asian Pac J Trop Dis. 2017;10(9):907-915.
Crossref - Jiang Y, Song Y, Jiang C, et al. Identification and characterization of Arthrobacter nicotinovorans JI39, a novel plant growth-promoting rhizobacteria strain from Panax ginseng. Front Plant Sci. 2022;13:2022.
Crossref - Robbins N, Spitzer M, Wang W, et al. Discovery of ibomycin, a complex macrolactone that exerts antifungal activity by impeding endocytic trafficking and membrane function. Cell Chem Biol. 2016;23(11):1383-1394.
Crossref - Sakai-Kawada FE, Ip CG, Hagiwara KA, Awaya JD. Biosynthesis and bioactivity of prodiginine analogs in marine bacteria, Pseudoalteromonas: A mini review. Mini Review. Front Microbiol. 2019;10:2019.
Crossref - Li J, Zhang L, Yao G, et al. Synergistic effect of co-culture rhizosphere Streptomyces: A promising strategy to enhance antimicrobial activity and plant growth-promoting function. Front Microbiol. 2022;13:2022.
Crossref - Henao L, Zade RSH, Restrepo S, Husserl J, Abeel T. Genomes of four Streptomyces strains reveal insights into putative new species and pathogenicity of scab-causing organisms. BMC Genomics. 2023;24(1):143.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.