Worldwide concern about the energy security and search for alternate energy source as fuel has created attention because of increased demand and fast depleting nonrenewable resources of fuel. In order to meet the increasing energy demand, we need to approach the sustainable, cost-effective alternates which should be able to cater unlimited resource utilization potential. Among the next-generation biofuels, butanol is the one which shows the ability to meet both the current and upcoming future energy crisis over the existing biofuels, for its distinguished merits; high energy density, high air-fuel ratio and lower hygroscopicity which make butanol superior over the other. In this review, we summarized all the methods attempted to screen out a high-stress tolerant strain through the various approach like; variation in sampling methods, media optimization, manipulation in physiochemical parameters etc. and then adapting them to high butanol environment. Secondly, the possible techniques involved in the identification of butanol tolerant gene and its overexpression in heterologous host or butanol tolerant microbes through genetic engineering, manipulating the metabolic pathway through engineering synergistically integrated with other disciplines such as synthetic biology as well as systems biology, has been discussed in detail. Apart from this, the present review also highlights the various bottlenecks, achievements and prospects for the commercialization of the butanol as the next generation of biofuel.
Butanol, Clostridium sp., Biofuels, Solvent toxicity, Tolerance, Energy.
Worldwide concern about the energy security and search for alternate energy source as fuel has created attention because of increased demand and fast depleting nonrenewable resources of petroleum. Alternatively, sustainable and renewable liquid transport fuels of biological origin have gained interest including ethanol which so far constitutes 90% of the total biofuels in the market1.Biofuels produced from sugar, starch, vegetable oil are termed as first-generation biofuel which includes ethanol, gasoline, vegetable oil, bio ether, biogas, and solid biofuel2.Microorganisms like; Clostridium acetobutylicum, Zymomonas mobilis, and Saccharomyces cerevisiae have been the native microbes which are employed for the current production of biofuels but these often suffer from limitations of slower growth, complex nutrient requirement, complex life-cycle and the major limitation being the production of a variety of by-products which suffer various challenges in downstream processing including high-cost recovery of targeted product3,4,1.In European countries, biodiesel is the major biofuel manufactured by the trans-esterification of the animal fats as well as oils of plant origin supplemented with solvent methanol5.Biofuel still suffers from various limitations like; wax formation in the fuel in the low temperature and transportation in the current fuel distribution infrastructure1. There have been attempts for the production of the first-generation biofuels from the rich and unused biological resources like lignocelluloses, biomass from algal cultures and gases like CO and CO2, but these resources still suffer from various limitations. Lignocelluloses, the most abundant biomass present on the earth contains 70% sugars but the sugar it contains requires various thermal, chemical and biochemical energy mediated process before being released for microbial fermentation. Similarly, algal biomass, which can also efficiently contribute for the production of first-generation biofuel, also suffers from the limitation of collection and dewatering1.
All these limitations of the first-generation biofuels have pushed the interest of the microbiologists and scientists of related field towards the development of the second-generation biofuel. the second-generation biofuels are obtained from biomass of non-food origin. Various engineering techniques like; metabolic engineering, genetic engineering, evolutionary engineering, system biology, and synthetic biology employed in synergy, aiming towards the production of the advanced biofuels have become comparatively easier. During the development of the advanced biofuels, factors like engine type, energy content, combustion quality i.e. odor, water miscibility and cost etc. has been taken into account6,1. During the process of the development of the advanced biofuels the major problem faced by the scientists is to trouble shoot the indigenous regulation for high yield of biofuel using the native hosts1. The reconstruction of the advanced biofuel pathways using the heterologous hosts like; E. coli and Saccharomyces cerevisiae will make us to overcome the above major challenges but the reconstruction of the biofuel pathways itself suffers from the challenges of limitations in production of enzyme for maximum metabolic outcome at the same time1.
Butanol is one of the most advanced biofuels as it possesses various distinguished features. Butanol has drawn attention for its major suitable chemical properties which turns to be superior in terms of energy content, corrosiveness, and volatility7,3. Butanol for its longer chain is more efficient than the currently used fuel ethanol8. Improved technical characteristics: viz higher ratio of air to fuel (11.1 vs. 9.0), higher amount of energy density (29.2 vs. 19.6 MJ/L), and lower hygroscopicity makes butanol more sought after than the existing biofuels8.Besides the all above advantages butanol has one of the major characteristics of being easily transported by the existing gas line distribution infrastructure and can be easily replaced with the existing car engines2.Butanol has many isomeric forms like iso-butanol, sec-butanol, ter-butanol, n-butanol, but out of all n-butanol and iso-butanol can only act as a potential solvent for industry and other isomers of butanol are highly soluble in water9. Butanol has advantage over ethanol because of its scarce solubility as well as resemblance in other characteristic properties with gasoline,.
The butanol production from microbes merits concern and symbolize as fascinating alternatives to present fuels through biotechnological intervention10. Microbial fermentation of Clostridium fulfills our current need for biofuel although Clostridium also suffers from various process limitations11. So, today the main concern of the microbiologists is to find an alternative microorganism which may emerge as the second-generation host for the production of advanced biofuel butanol. Higher concentration of butanol in the reaction environment causes toxicity which leads to the lower production of butanol, and the cell toxicity caused by the higher concentration of butanol is the basic challenge faced today. A cellular system which can give the optimum yield efficiency and is cost effective too, is being tried to develop by the metabolic engineers1.In this review we discuss the different biofuel producing bacteria, their bottlenecks, achievements and prospective for prospect directions in the bacterial biofuel production.
Physico-chemical characteristics of Butanol
Butanol, popularly called as butyl alcohol, is basically an alcohol with four carbon chain having chemical formula; C4H9OH. It is principally used as intermediate in the synthesis of different chemicals and solvent at industrial scale. Butanol is also used in life sciences and medicine as cleaning and sterilizing agent. In pharmaceuticals it has a great demand for production of drugs. In addition, most recently butanol has been used as potential fuel (Fig.1). Butanol occurs in different isomeric forms: iso-butanol, 2-butanol, n-butanol and tert-butanol, Presence of a straight chain carbon length with an alcohol group at the terminal carbon are commonly called butanol.
The isomer 2-butanol or sec-butanol is actually a butanol having an alcoholic functional group at an internal carbon. The chemical structure and arrangement of alcoholic functional group in different isomers is presented in Fig. 2.
The physiochemical properties of different isomers of butanol differ due the basic differences in their chemical structure. These isomers differ in their solubility, melting and boiling points as well as other so many properties. The details of variation in physical and chemical characteristics of all the four isomers of butanol are present for a comparative view in Table 1.
Table (1):
Physiochemical properties of different isomers of butanol
Characteristics |
n-butanol |
iso-butanol |
sec-butanol |
tert-butanol |
|---|---|---|---|---|
Formula |
C4H10O |
C4H10O |
C4H10O |
C4H10O |
IUPAC name |
Butan-1-ol |
2-methylpropan-1-ol |
Butan-2-ol |
2-Methylpropan-2-ol |
Molecular weight [gm/mol] |
74.1 |
74.1 |
74.1 |
74.1 |
Boiling point (°C) |
117 |
108 |
100 |
83 |
Melting point (°C) |
-90 |
-108 |
-115 |
25.7 |
Blending RON* |
94°,96° |
113° |
101° |
104-110° |
Blending MON† |
78°,81° |
94° |
91° |
89-98° |
Solubility in water wt% at 20°C |
7.7 |
8.5 |
12.5 |
Miscible |
Surface tension at 20°CmN/m |
24.7 |
23 |
– |
20.7 |
Toxicity |
Most toxic |
Least toxic among butanol |
Less toxic |
Less toxic |
Suitability |
Most suitable for bio fuel |
Suitable for bio fuel. |
Suitable as cleaning agent and paint remover. |
Suitable as gasoline octane booster, ethanol denaturant. |
Self-ignition temperature [°C] |
343 |
415.6 |
406.1 |
477.8 |
Viscosity at 25°C [mPa·s] |
2.544 |
4.312 |
3.096 |
– |
*RON: Blending Research Octane Number; †MON: Blending Motor Octane Number
Source: 1. Adanced Motors fuels (http://www.iea-amf.org/content/fuel_information/butanol/properties); 2. PubChem (https://pubchem.ncbi.nlm.nih.gov/); 3. Encyclopedia of Chemical Processing and Design: Volume 20- Ethanol as Fuel: Options: Advantages, and Disadvantages to Exhaust Stacks: Cost [Hardcover 1984]; By John J Mc Ketta, Mc Ketta J Mc
Butanol as potential alternative biofuel to ethanol
Butanol as compared to ethanol, is primary alcohol with short carbon chain which exists in isomeric forms of as mentioned earlier. Due to the presence of two more methylenes, butanol tends to be more hydrophobic than ethanol resulting in less volatile, high energy dense and more miscible with gasoline12.The physical and chemical properties has been compared and presented in the Table 2 which shows highly remarkable variations and are responsible for the properties of advantage to mankind. The above-mentioned advantages of butanol over ethanol have drawn the attention of biologists in recent years to consider it to be potential alternative biofuel. Till date most of the butanol production is being reported from the Clostridium bacteria through the anaerobic process which leads to the production of acetone, butanol and ethanol6. Apart from this, several other common bacteria like; Lactobacillus, Bacillus, E.coli as well as Pseudomonas are known for their potential to yield butanol but are unable to tolerate butanol beyond 2% which prevents their growth and activity. This makes industrial production uneconomic and irrelevant13. To overcome the problem of toxicity and downstream processing, steps have been taken to introduce the non-native strains of Bacillus subtilis, Pseudomonas putida, Lactobacillus and E. coli through genetic manipulations9,14. Thus the presently available technology seems to be unfavorable economically for the commercialization of butanol12. Higher concentration of the substrate requirements, sensitivity towards higher pH, temperature, and concentration of product, lack of cost-effective product recovery methods etc., draws the attention of the biologists to develop new tolerant strains14.Therefore, butanol tolerating mechanisms in the microbes need to be observed and studied at cellular and molecular level and thus synthetic biology approach have drawn the attention of microbiologist for the development of toxicity tolerant non- native strains1.
Table (2):
Difference in physiochemical properties of ethanol and butanol
Properties |
Ethanol [C2H5OH] |
n-Butanol [C4H10OH] |
|---|---|---|
Cetane number |
8 |
25 |
Octane number |
108 |
96 |
Viscosity(mm2/s)at 40°C |
1.08 |
2.63 |
Saturation pressure (kPa) at 38°C |
13.8 |
2.27 |
Flammability limits (% vol) |
4.3-19 |
1.4-11.2 |
Boiling point (°C) |
78.4 |
117.7 |
Oxygen content(% wt) |
34.8 |
21.6 |
Density (g/ml) at 20°C |
0.79 |
0.808 |
Auto ignition temperature (°C) |
434 |
385 |
Lower heating value (MJ/kg) |
26.8 |
33.1 |
Methods of screening of butanol tolerant strains
Various methods like chemical mutagenesis, continuous culture and serial enrichment process have helped to develop tolerant strains mostly of the Clostridium sp.15.Due to the intolerance of the butanol produced by microbial cultures, various methods used to remove butanol from the broth are either gas stripping or membrane stripping. This resulted in enhancement of butanol production in the culture broth16.
From recent studies, it has been understood that some stains like; p260 and BA101 of Clostridium beijerinckii are able the degrade biomass of lignocellulose origin and convert its hydrolysates into valued chemical solvents having biofuel properties17-22.Staphylococcus haemolyticus is reported for the tolerance for higher solvent concentrations. S. cerevisiae and E. coli were reported to yeild butanol although with low yield after engineering with clostridium genes key to butanol synthesis pathways23.Grsw2-B1 strain of the Bacillus subtilis has emerged as a potential host for the production of the biobutanol up to 2%9. Gram positive lactobacillus strains have also been reported to tolerate butanol up to 3%. Adopted strains of the pseudomonas have also reported to tolerate butanol up to 6% but their normalized growth kinetics observed was low.
Various microbes isolated from the environmental samples have been reported to show the butanol tolerance (Table3). Nine genera i.e. Rummeliibacillus, , Coprothermobactoer, Bacillus, Caloribacterium, Lysinibacillus, Enterococcus, Hydrogenoanaero bacterium, Cellulosimicrobium and Brevibacillus within the phyla Firmicutes and Actinobacteria have been experimentally tested by Kanno et al.14 identified two botanol tolerant strains which can trive at a concentration higher than 3.0% and further experimentally concluded that the membrane fluidity was maintained due to the enhanced amount of fatty acids of longer carbon chain, and thus were able to tolerate the higher concentration of butanol24,18,21,22,25,26,14. Similarly different other species have also shown tolerance to the butanol like Pseudomonas sp., Bacillus subtilis, Escherichia coli, Clostridium sp., but to different concentrations of butanol. Hybrid strains have also been generated between L. brevis and E. coli by protoplast fusion which resulted in the high tolerance of butanol and comparatively growth inhibitory effect of butanol was also less27. Compared to the cyanobacteria the % tolerance observed in C. acetobutylicum, Pseudomonas putida, Z. mobilis, E. coli, was 10 times high28.
Table (3):
Percentage tolerance of microbial isolates for next-generation biofuel butanol
| S. No. | Microorganism | Tolerance (% V/V) | References |
|---|---|---|---|
| 1 | Escherichia coli | 1.50 | Fischer et al. 2008 |
| 2 | Pseudomonas putida | Rühl et al. 2009 | |
| 3 | Zymomonas mobilis | Green 2011 | |
| 4 | Clostridium acetobutylicum | 1.60 | Ezeji et al. 2010 |
| 5 | Clostridium beijerinckii P260 | 1.95 | Qureshi et al. 2012 |
| 6 | Clostridium beijerinckii BA101 | 1.96 | Qureshi et al. 2007 |
| 7 | Bacillus amyloliquefaciens | 2.00 | Kanno et al. 2013 |
| 9 | Brevibacillus reuszeri | ||
| 10 | Caloribacterium cisternae | ||
| 12 | Cellulosimicrobium cellulans | ||
| 13 | Coprothermobacter proteolyticus | ||
| 14 | Garciella nitratireducens | ||
| 15 | Hydrogenoanaerobacterium sacchrovorans | ||
| 16 | Rummeliibacillus pycnus | ||
| 18 | Saccharomyces cerevisiae | Fischer et al. 2008 | |
| 19 | Bacillus mycoides | 2.50 | Kanno et al. 2013 |
| 21 | Clostridium pasteurianum | ||
| 22 | Lysinibacillus xylanilyticus | ||
| 24 | Eubacterium cylindroides | 3.00 | |
| 26 | Lactobacillus brevis | Knoshaug and Zhang 2009; Liu and Qureshi 2009 | |
| 28 | Lactobacillus delbrueckii | ||
| 30 | Enterococcus faecalis (AB012212) | 3.50 | Kanno et al. 2013 |
| 31 | Pseudomonas sps. | 6.00 | Park et al. 2007; Rühl et al. 2009 |
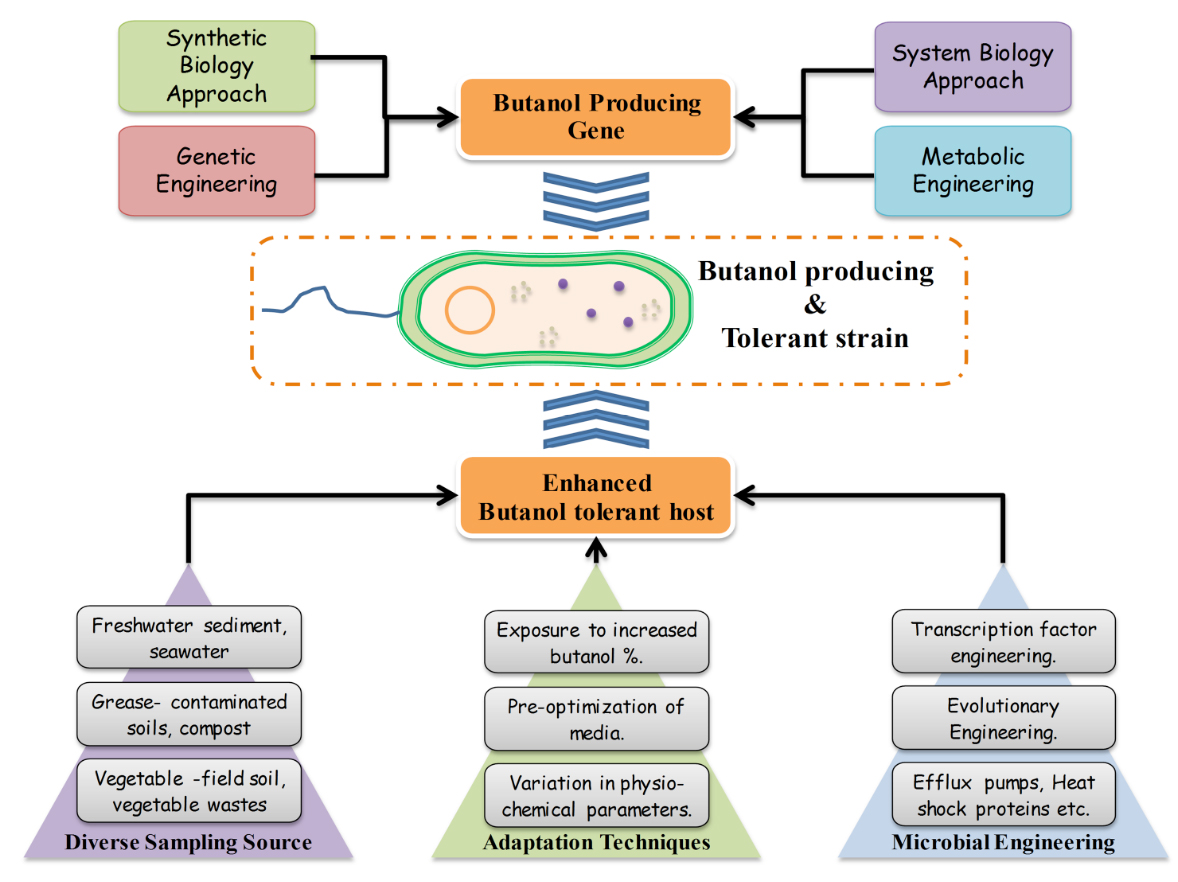
Attempts are taken to trouble shoot the bottlenecks commonly encountered while producing butanol using microorganism as a biological agent. The most important hurdle, scientists’ face in production of butanol has been the toxicity of butanol in the fermentation broth beyond a certain level which generally varies from 2-6%. Therefore, obtaining a high butanol tolerant strain through various approaches like variation in sampling source, media optimization, change in other physiochemical parameters etc. and then adapting them to the butanol stress. Identification of butanol tolerant gene and it’s over expression in heterologous host or butanol tolerant microbes through genetic engineering, manipulating the metabolic pathway through metabolic engineering synergistically integrated with system biology and synthetic biology provides a great scope of coming up with a solution for large scale production of advanced biofuels (Fig.3).
Manipulation of media and cultivation techniques for butanol tolerance
Substrate versatility and metabolic efficiency of butanol tolerating microbes led the researchers to draw their attention towards manipulation of different physiological and biochemical factors involved in butanol tolerating strains. Different optimized conditions were used for the isolation, screening and development of the butanol tolerating microbes14.
Media optimization is the key technique being reported by the biologists in various research experiments for the development of butanol tolerant microbe29,26. In the research conducted using different strains of Pseudomonas putida DOT-TIE, KT2440, S12, VLB120, different media like M9 minimal media and LB medium supplemented with glucose, maximum tolerance up to the 6% was achieved by the adopted strain30,26.In the other experiment conducted by Kanno et al.14 , butanol tolerance at increased concentrations was checked for the samples collected from the varied environmental sources by manipulating the media using different carbon sources (glucose, tryptone, yeast extract), salts (NH4)2SO4, KH2PO4 and K2HPO4), vitamins and the incubation time (48 hr for short and 3 to 9 months for long term)14.Some researchers favored RG medium to initiate the culture and MRS medium to test alcohol tolerance to avoid the complication of too many osmo-protectants in the latter 31,25,26,32.
Mechanisms behind butanol tolerance
Tolerance in butanol strains, so far achieved have been due to modifications of various in situ processing and recovery techniques like; manipulation of temperature, pH, substrate enrichment, time, and change in metabolic pathways etc.11,9,14.These techniques led the microbes to tolerate the butanol toxicity and enhanced their growth in the reaction environment. Other than the above approaches, alternate multiple tolerance strategies have also been employed which includes engineering of the biofuel export system, membrane modification, and heat shock proteins, metabolic engineering and synthetic biology approach33,11.
Efflux pump mediated butanol tolerance
The significant role of efflux pumps in biofuel tolerance has been understood as these behave as membrane transporters in exporting the toxic molecules out of the cells by utilizing proton motive force34-37.Various efflux pumps have been reported from several solvent tolerant species like; the solvent resistance pumps (srpABC) from Pseudomonas putida S1238,39, 3 solvent resistant efflux pumps from pseudomonas putida DOT-TIE40,39, acrAB pump from Escherichia coli41,39, three mex pumps from Pseudomonas aeruginosa39,42and the RND pumps, most commonly found in the gram negative bacteria are reported for the best tolerance of biofuels39,33.An efflux pump encoded by focA over expression showed the tolerance towards the n-butanol as reported in E. coli37,1.Fiocco et al. 2007 reported the AcrB efflux pump which by acting on the n-butanol, a non-native substrate can support the growth of E. coli by 25% in the presence of n-butanol.
Heat shock proteins conferring butanol resistance
Heat shock proteins were known to help in improving the biofuel tolerance which could be due to the over expression of proteins; rpoH, dnaj, htpG, and ibpABas reported in the E.coli43,44.Whereas, Gros ESL, dnaKJ, hsp18, hsp90 genes have been over expressed in Clostridium acetobutylicum to bring in tolerance to butanol45. Evidences have been there for the over expression of the heat shock proteins from Lactobacillus plantarum46.In E. coli, modification to the chaperons and various heat shock proteins including rpoH, dnaJ have been reported which are involved in the butanol stress mechanism39,44.Thus conclusion was drawn that over expression of the heat shock proteins can help in overcoming the solvent stress and makes the strain tolerant to butanol43,45,39.In addition, the role of heat shock proteins in response to the n-butanol and iso-butanol stress have been identified from the transcriptomics and proteomic studies44,13.
Over expression of the GroESL chaperone system resulted in 12-fold butanol production 33,47,48.11 genes in E. coli were reported to be over expressed under n-butanol49 of which three were the heat shock genes47.The two genes encoding the tail of butanol synthesis pathway has been cloned by 51in the genus lactobacillus were named as Bcs-operon and thl. Up regulation of the heat shock proteins have also been reported in the lacto bacillus in response to the butanol tolerance17,27. Unlike the involvement of the GroESL in E. coli, over expression of the GroESL have led to the survival of Lactobacillus paracasei strains in the presence of butanol toxicity50,46,51,27.Up regulation of the fatty acid synthetic pathway towards the response for butanol stress in L. brevis have been also reported by R hl et al.26.Cyclopropane fatty acid were reported for the stabilizing of the membrane lipids and in the cell membrane fluidity in response to various stress unlike acid, thermal, osmotic in various microbes52,53 in E. coli54, P. putida55, lactobacillus56,27.Reyes et al.47further confirmed the role of the cyclo propane fatty acid in the solvent tolerance especially for n-butanol in E. coli. Genes like ygfo, setA, mdtA and pgsA have been reported for their role in the enhancement of the butanol tolerance in the wild type E. coli47,49.Over expression of the sodB an iron dismutase is also reported to increase the n-butanol tolerance in E. coli58,27. PepB Proteinase was reported to enhance the tolerance of butanol to an extend of 40% in over expressed condition33. The GroESL chaperone system was activated in response to n-butanol toxicity in E. coli and C. acetobutylicum33,48.
Membrane structure modification and butanol tolerance
Modification in the membrane especially in phospholipids chain has been reported to increase the solvent tolerance59.Cis to trans conversion of the unsaturated fatty acid as one of the membrane modifications adopted by various strains which decreases the membrane fluidity under the solvent toxicity46.Various mechanism for tolerance to bio-butanol by bacterial strains have been reported which includes cell membrane modification by change in the fatty-acid composition, generally from cis to trans unsaturated fatty acid as reported in the gram positive strain of Staphylococcus haemolyticus60,12, and by extracellular capsule thickness which helps in the maintenance of their structural integrity as reported in the clostridium sp.39,61,62,3,12,9,63,14.Such modifications in the membrane structure results in incorporation of butanol into the cytoplasmic membrane and interferes with the functions of the cells. Mutation in regulators and transcriptional factors regulation in cell of E. coli towards iso-butanol have been also cited for the tolerance towards stress39,64,65.Nicolaou et al.37 reported that during the solvent toxicity reactive oxygen species are found to be highly increased. Elevation in the level of the reactive oxygen species (ROS) have also been reported during the solvent stress so the engineering in such genes have led the interest of the researchers for the development of new solvent tolerant strains39,47,49.
Metabolic engineering and synthetic biology for butanol tolerance
Metabolic engineering and synthetic biology approach have been designed for the pathway optimization of host and further for advanced biofuelsproduction1.Metabolic engineering techniques have been also adopted for the enhancement of the butanol production in Clostridium acetobutylicum6.The reconstructed butanol pathway of clostridium was introduced in E. coli66,23,9, Bacillus subtilis and Pseudomonas putida61,63, and lactic acid bacteria i.e. Lactobacillus brevis38.
Potential applications and scopes of butanol production
Among all the biofuels reported till date bio-butanol meets all the criteria of the advanced biofuel including performance of auto-engine and emissions as well as their combustion features. This has the potential to combat the prevailing energy crisis, more importantly for its use in internal combustion engines. In addition, the other significant applications of butanol are: (i) use in spark ignition engine as an alternative fuel of gasoline, (ii) use as potential fuel supplement for compression ignition in diesel engine (iii) use in burning reactors in order to carry out fundamental combustion tryouts. Looking at its potential applications and scope of uses from the literature, it is clear that more exhaustive investigations are still lacking before considering butnol as a fuel substitute as compared to the its counterparts.
Perspective remarks
Lack of tolerance to the solvent toxicity by the native and non-native bacterial strains is today the major obstacle in the development of the next generation biofuel. Various synergistic approaches have been developed to obtain the non-native bacterial strains exhibiting tolerant to the high solvent toxicity. To meet the increasing energy demand and to ensure the energy security, the development of the next generation biofuel is a panacea in today’s world. Among all the biofuels reported till date bio-butanol meets all the criteria of the advanced biofuel which has the potential to combat the prevailing energy crisis. Classical butanol producing bacteria Clostridium sp. suffers many challenges like low percentage tolerance, formation of various by-products etc. The major bottleneck today is that we lack in solvent tolerant butanol producing strain. Aerobic strains can meet all the challenges faced by the anaerobic strains but the problem occurs at the genetic modification where aerobic strains do not have the stress tolerant genes which can cope up with the high level of solvent toxicity like butanol. So, in order to develop a tolerant butanol producing aerobic strain we need to adapt various screening methods integrated with the genetic, metabolic engineering, synthetic biology and system biology approach to come up with a solution and meet the energy security in a cost effective way.
Acknowledgements
We are thankful to Chairman, Siksha ‘O’ Anusandhan (Deemed to be) University, Bhubaneswar, Odisha for their infrastructure support for carrying out of this work.
Conflict of Interest
The authors declares that there is no conflict of interest.
Authors’ Contribution
RKS drafted the manuscript. AD and SS gathered information from the literature. SD and MG designed the figures and tables. ES supervised and reviewed the manuscript.
Funding
None.
Data Availability
All datasets generated or analyzed during this study are included in this review manuscript.
Ethics Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
- Liu G., Zhang J., Bao J. Cost evaluation of cellulase enzyme for industrial-scale cellulosic ethanol production based on rigorous Aspen Plus modeling. Bioprocess Biosyst. Eng., 2016; 39: 133–140.
Crossref - Gonzבlez-Ramos D., van den Broek M., van Maris A.J., Pronk J.T., Daran J.M. Genome-scale analyses of butanol tolerance in Saccharomyces cerevisiae reveal an essential role of protein degradation. Biotechnol. Biofuels., 2013; 6: 48.
Crossref - Lee S.Y., Park J.H., Jang S.H., Nielsen L.K., Kim J., Jung K.S. Fermentative butanol production by clostridia. Biotechnol. Bioeng., 2008; 101: 209–228.
Crossref - Liao J.C., Mi. L., Pontrelli S., Luo S. Fuelling the future: microbial engineering for the production of sustainable biofuels. Nat. Rev. Microbiol., 2016; 14: 288-304.
Crossref - Banerjee R., Proshlyakov Y., Lipscomb J.D., Proshlyakov, D.A. Structure of the key species in the enzymatic oxidation of methane to methanol. Nature., 2015; 518: 431–434.
Crossref - L tke-Eversloh T., Bahl H. Metabolic engineering of Clostridium acetobutylicum: recent advances to improve butanol production. Curr. Opin. Biotechnol., 2011; 22: 634–647.
Crossref - Ravinder K., Pradeep K. Future microbial applications for bioenergy production: A perspective. Front. Microbiol., 2017; 8: 450.
Crossref - Pבsztor A., Kallio P., Malatinszky D., Akhtar M.K., Jones P.R. A synthetic O2-tolerant butanol pathway exploiting native fatty acid biosynthesis in Escherichia coli. Biotechnol. Bioeng., 2015; 112: 120–128.
Crossref - Kataoka N., Tajima T., Kato J., Rachadech W., Vangnai A.S. Development of butanol-tolerant Bacillus subtilis strain GRSW2-B1 as a potential bioproduction host. AMB Express., 2011; 1: 10.
Crossref - Branduardi P., de Ferra F., Longo V., Porro D. Microbial n-butanol production from Clostridia to non-Clostridial hosts. Eng. Life Sci., 2014; 14: 16–26.
Crossref - Yang S., Mohagheghi A., Franden M.A., Chou Y. C., Chen X., Dowe N., Himmel M.E., Zhang M. Metabolic engineering of Zymomonas mobilisfor 2, 3-butanediol production from lignocellulosic biomass sugars. Biotechnol. Biofuels., 2016; 9:189.
Crossref - Liu S., Qureshi N. How microbes tolerate ethanol and butanol. N. Biotechnol., 2009; 26: 117–121.
Crossref - Jin H., Chen L., Wang J., Zhang W. Engineering biofuel tolerance in non-native producing microorganisms. Biotechnol. Adv., 2014; 32: 541–548.
Crossref - Kanno M., Katayama T., Tamaki H., Mitani Y., Meng X.-Y., Hori T., Narihiro T., Morita N., Hoshino T., Yumoto I., Kimura N., Hanada S., Kamagata Y. Isolation of butanol- and isobutanol-tolerant bacteria and physiological characterization of their butanol tolerance. Appl. Environ. Microbiol., 2013; 79: 6998–7005.
Crossref - Hermann M., Fayolle F., Marchal R., Podvin L., Sebald M., Vandecasteele J.P. Isolation and characterization of butanol-resistant mutants of Clostridium acetobutylicum. Appl. Environ. Microbiol., 1985; 50: 1238–1243.
- Ezeji T., Milne C., Price N.D., Blaschek H.P. Achievements and perspectives to overcome the poor solvent resistance in acetone and butanol-producing microorganisms. Appl. Microbiol. Biotechnol., 2010; 85: 1697–1712.
Crossref - VollherbstSchneck K., Sands J.A., Montenecourt B.S. Effect of butanol on lipid composition and fluidity of Clostridium acetobutylicum ATCC 824. Appl. Environ. Microbiol., 1984; 47: 193–194.
- Zhang Ch., Tinggang Li., Jianzhong He. Characterization and genome analysis of a butanol–isopropanol-producing Clostridium beijerinckii strain BGS1. Biotechnol. Biofuels.,2018; 11: 280.
Crossref - Kushwaha D., Srivastava N., Mishra I., Upadhyay S.N., Mishra P.K. Recent trends in biobutanol production. Rev. Chem. Eng., 2018; 37: 1-30.
- Ezeji T., Qureshi N., Blaschek H.P. Butanol production from agricultural residues: Impact of degradation products on Clostridium beijerinckii growth and butanol fermentation. Biotechnol. Bioeng., 2007; 97: 1460–1469.
Crossref - Qureshi N., Saha B.C., Cotta M.A. Butanol production from wheat straw hydrolysate using Clostridium beijerinckii. Bioprocess Biosyst. Eng., 2007; 30: 419–427.
Crossref - Qureshi N., Liu S., Ezeji T.C. Cellulosic butanol production from agricultural biomass and residues: Recent Advances in Technology, in: Advanced Biofuels and Bioproducts. Springer New York., 2012; 247–265.
Crossref - Inui M., Suda M., Kimura S., Yasuda K., Suzuki H., Toda H., Yamamoto S., Okino S., Suzuki N., Yukawa H. Expression of Clostridium acetobutylicum butanol synthetic genes in Escherichia coli. Appl. Microbiol. Biotechnol., 2008; 77: 1305–1316.
Crossref - Birgen C., D rre P., Preisig H., Wentzel A. Butanol production from lignocellulosic biomass: revisiting fermentation performance indicators with exploratory data analysis. Biotechnol. Biofuels., 2019; 12: 167.
Crossref - Knoshaug E.P., Zhang M. Butanol tolerance in a selection of microorganisms. Appl. Biochem. Biotechnol., 2009; 153: 13–20.
Crossref - R hl J., Schmid A., Blank LM. Selected Pseudomonas putida strains able to grow in the presence of high butanol concentrations. Appl Environ Microbiol., 2009; 75: 4653–6.
Crossref - Winkler J., Kao K.C. Transcriptional Analysis of Lactobacillus brevis to N-Butanol and Ferulic Acid Stress Responses. PLoS One., 2011; 6: e21438.
Crossref - Tian X., Chen L., Wang J., Qiao J., Zhang W. Quantitative proteomics reveals dynamic responses of Synechocystis sp. PCC 6803 to next-generation biofuel butanol. J. Proteomics., 2013; 78: 326–345.
Crossref - Fischer C.R., Klein-Marcuschamer D., Stephanopoulo G. Selection and optimization of microbial hosts for biofuels production. Metab. Eng., 2008; 10: 295–304.
Crossref - Park J.-B., B hler B., Panke S., Witholt B., Schmid A. Carbon metabolism and product inhibition determine the epoxidation efficiency of solvent-tolerant Pseudomonas sp. strain VLB120ִC. Biotechnol. Bioeng., 2007; 98: 1219–1229.
Crossref - Wen Z., Ledesma-Amaro R., Lin J., Jiang Y., Yang S. Improved n-butanol production from Clostridium cellulovorans by integrated metabolic and evolutionary engineering. Appl. Environ. Microbiol., 2019; 85: e02560-18.
Crossref - Zhiqiang Wen, Rodrigo Ledesma-Amaro, Jianping Lin, Yu Jiang, Sheng Yang Li J., Zhao J.B., Zhao M., Yan Y.L., Jiang W.H., Yang S. Screening and characterization of butanol-tolerant micro-organisms. Lett. Appl. Microbiol., 2010; 50: 373–379.
Crossref - Dunlop M.J. Engineering microbes for tolerance to next-generation biofuels. Biotechnol. Biofuels., 2011; 4: 32.
Crossref - He X., Xue T., Ma Y., Zhang J., Wang Z., Hong J., Hui L., Qiao J., Song H., Zhang M. Identification of functional butanol-tolerant genes from Escherichia coli mutants derived from error-prone PCR-based whole-genome shuffling. Biotechnol. Biofuels., 2019; 12: 73.
Crossref - Okochi M., Kurimoto M., Shimizu K., Honda H. Increase of organic solvent tolerance by overexpression of manXYZ in Escherichia coli. Appl. Microbiol. Biotechnol., 2007; 73: 1394–1399.
Crossref - Nikaido H., Takatsuka Y. Mechanisms of RND multidrug efflux pumps. Biochim. Biophys.Acta., 2009; 1794: 769–781.
Crossref - Nicolaou S.A., Gaida S.M., Papoutsakis E.T. A comparative view of metabolite and substrate stress and tolerance in microbial bioprocessing: From biofuels and chemicals, to biocatalysis and bioremediation. Metab. Eng., 2010; 12: 307–331.
Crossref - Kieboom J., Dennis J.J., de Bont J.A.M., Zylstra G.J. Identification and molecular characterization of an efflux pump involved in Pseudomonas putida S12 Solvent Tolerance. J. Biol. Chem., 1998; 273: 85–91.
Crossref - Nielsen L.E., Kadavy D.R., Rajagopal S., Drijber R., Nickerson K.W. Survey of Extreme solvent tolerance in gram-positive Cocci: Membrane fatty acid changes in Staphylococcus haemolyticus grown in toluene. Appl. Environ. Microbiol., 2005; 71: 5171–5176.
Crossref - Rojas A., Duque E., Mosqueda G., Golden G., Hurtado A., Ramos J.L., Segura A. Three Efflux pumps are required to provide efficient tolerance to toluene in Pseudomonas putida DOT-T1E. J. Bacteriol., 2001; 183: 3967–3973.
Crossref - Basler G., Thompson M., Tullman-Ercek D., Keasling J. A Pseudomonas putida efflux pump acts on short-chain alcohols. Biotechnol. Biofuels., 2018; 11: 136.
Crossref - Zheng Z., Li C., Liu H., Zhang Y., Zhong X., Yao M. Experimental study on diesel conventional and low temperature combustion by fueling four isomers of butanol. Fuel., 2015; 141: 109–119.
Crossref - Tomas C.A., Welker N.E., Papoutsakis E.T. Overexpression of groESL in Clostridiumacetobutylicum results in increased solvent production and tolerance, prolonged metabolism, and changes in the cell’s transcriptional program. Appl. Environ. Microbiol., 2003; 69: 4951–4965.
Crossref - Rutherford B.J., Dahl R.H., Price R.E., Szmidt H.L., Benke P.I., Mukhopadhyay A., Keasling J.D. Functional genomic study of exogenous n-butanol stress in Escherichia coli. Appl.Environ. Microbiol., 2010; 76: 1935–1945.
Crossref - Tomas C.A., Beamish J., Papoutsakis E.T. Transcriptional analysis of butanol stress and tolerance in Clostridium acetobutylicum. J. Bacteriol., 2004;186: 2006–2018.
Crossref - Fiocco D., Capozzi V., Goffin P., Hols P., Spano G. Improved adaptation to heat, cold, and solvent tolerance in Lactobacillus plantarum. Appl. Microbiol. Biotechnol., 2007; 77: 909–915.
Crossref - Reyes L.H., Almario M.P., Kao K.C. Genomic library screens for genes involved in n-butanol tolerance in Escherichia coli. PLoS One., 2011; 6: e17678.
Crossref - Zingaro K.A., Terry Papoutsakis E. GroESL overexpression imparts Escherichia coli tolerance to i-, n-, and 2-butanol, 1,2,4-butanetriol and ethanol with complex and unpredictable patterns. Metab. Eng., 2013; 15: 196–205.
Crossref - Reyes L.H., Almario M.P., Winkler J., Orozco M.M., Kao K.C. Visualizing evolution in real time to determine the molecular mechanisms of n-butanol tolerance in Escherichia coli. Metab. Eng., 2012; 14: 579–590.
Crossref - Desmond C., Fitzgerald G.F., Stanton C., Ross R.P. Improved stress tolerance of GroESL-overproducing Lactococcus lactis and probiotic Lactobacillus paracasei NFBC 338. Appl. Environ. Microbiol., 2004; 70: 5929–5936.
Crossref - Abdullah-Al-Mahin Sugimoto, S., Higashi C., Matsumoto S., Sonomoto K. Improvement of multiple-stress tolerance and lactic acid production in Lactococcus lactis NZ9000 under conditions of thermal stress by heterologous expression of Escherichia colidnaK. Appl. Environ. Microbiol., 2010; 76: 4277–4285.
Crossref - Guerzoni M.E., Lanciotti R., Cocconcelli P.S. Alteration in cellular fatty acid composition as a response to salt, acid, oxidative and thermal stresses in Lactobacillus helveticus. Microbiology., 2001; 147: 2255–2264.
Crossref - Muסoz-Rojas J., Bernal P., Duque E., Godoy P., Segura A.,Ramos J-L.Involvement of cyclopropane fatty acids in the response of Pseudomonas putida KT2440 to freeze-drying. Appl. Environ. Microbiol., 2006; 72: 472–477.
Crossref - Jin C., Yao M., Liu H., Lee C.F., Ji J. Progress in the production and application of n-butanol as a biofuel. Renew. Sustain. Energy Rev., 2011; 15: 4080–4106.
Crossref - Montanari C., SadoKamdem S.L., Serrazanetti D.I., Etoa F.-X., Guerzoni M.E. Synthesis of cyclopropane fatty acids in Lactobacillus helveticus and Lactobacillus sanfranciscensis and their cellular fatty acids changes following short term acid and cold stresses. Food Microbiol., 2010; 27: 493–502.
Crossref - Borden J.R., Papoutsakis E.T. Dynamics of Genomic-Library Enrichment and Identification of Solvent Tolerance Genes for Clostridium acetobutylicum. Appl. Environ. Microbiol., 2007; 73: 3061–3068.
Crossref - Kim Y.H., Cho K., Yun S.-H., Kim J.Y., Kwon K.-H., Yoo J.S., Kim S. Il. Analysis of aromatic catabolic pathways in Pseudomonas putida KT 2440 using a combined proteomic approach: 2-DE/MS and cleavable isotope-coded affinity tag analysis. Proteomics., 2006; 6: 1301–1318.
Crossref - Lynd L.R., Van Zyl WH., McBride JE., Laser M. Consolidated bioprocessing of cellulosic biomass: An update. Curr. Opin. Biotechnol., 2005; 16: 577–583.
Crossref - Ramos J.L., Duque E., Gallegos M.-T., Godoy P., Ramos-Gonzבlez M.I., Rojas A., Terבn W., Segura A. Mechanisms of solvent tolerance in gram-negative bacteria. Annu. Rev. Microbiol., 2002; 56: 743–768.
Crossref - Atsumi S., Hanai T., Liao J.C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature., 2008; 451: 86–89.
Crossref - Nielsen D.R., Leonard E., Yoon S.-H., Tseng H.-C., Yuan C., Prather K.L.J. Engineering alternative butanol production platforms in heterologous bacteria. Metab. Eng., 2009; 11: 262–273.
Crossref - Alper H., Moxley J., Nevoigt E., Fink G.R., Stephanopoulos G. Engineering yeast transcription machinery for improved ethanol tolerance and production. Science., 2006 ; 314: 1565–1568.
Crossref - Dragosits M., Mozhayskiy V., Quinones-Soto S., Park J., Tagkopoulos I. Evolutionary potential, cross-stress behavior and the genetic basis of acquired stress resistance in Escherichia coli. Mol. Syst. Biol., 2013; 9: 643–643.
Crossref - Santos C.N.S., Stephanopoulos G. 2008. Combinatorial engineering of microbes for optimizing cellular phenotype. Curr. Opin. Chem. Biol., 2008;12: 168–176.
Crossref - Berezina O. V., Zakharova N. V., Brandt A., Yarotsky S. V., Schwarz W.H., Zverlov V. V. Reconstructing the clostridial n-butanol metabolic pathway in Lactobacillus brevis. Appl. Microbiol. Biotechnol., 2010; 87: 635–646.
Crossref - Atsumi S., Cann A.F., Connor M.R., Shen C.R., Smith K.M., Brynildsen M.P., Chou K.J.Y., Hanai T., Liao,J.C. Metabolic engineering of Escherichia coli for 1-butanol production. Metab. Eng., 2008; 10: 305–311.
Crossref
© The Author(s) 2019. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.