ISSN: 0973-7510
E-ISSN: 2581-690X
The karst ecosystem, distinguished by its unique geophysical characteristics, hosts a wide variety of microbes. In these environments, cave-dwelling bacteria are renowned for producing compounds with diverse biological activities. Therefore, this study was conducted to explore the antibacterial potential of cave-dwelling bacteria from the Maros-Pangkep Karst Area in South Sulawesi, Indonesia. Soil samples were collected from Leang Kassi, Leang Jarie, and Leang Ulu Wae caves. The three caves are located at different points within the Maros-Pangkep karst system, each representing an independent and spatially separated sampling site. The bacterial isolation was performed, and antibacterial agar diffusion assays were conducted against pathogenic bacteria as a preliminary qualitative screening approach. The secondary metabolites from promising isolates were extracted using ethyl acetate and tentatively characterized using GC-MS. Additionally, the putative compounds were then screened and evaluated for their potential interaction with the protein target through molecular docking analysis as an in silico analysis approach. Results showed that isolates exhibited distinct morphological and antibacterial properties, with one from Leang Ulu Wae showing notable inhibitory effects in preliminary screening assays. 16S rRNA identified the Leang Ulu Wae isolate as Exiguobacterium. GC-MS analysis detected three major compounds with a library match (Similarity Index, SI) ≥90%. In silico analysis suggested that Pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methylpropyl), exhibited the most favorable predicted binding interaction with the protein target, followed by Cyclo(L-prolyl-L-valine) and 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a]. The findings suggest that the bacteria located in the caves of the Maros-Pangkep Karst Area possess considerable potential for future antibacterial research.
Antibacterial Activity, Cave Bacteria, GC-MS Analysis, Karst Ecosystem, Secondary Metabolites
Antibacterial resistance poses a significant global threat, negatively affecting clinical and therapeutic outcomes.1 Antibacterial resistance refers to pathogenic bacteria developing the ability to resist the effects of antibacterial.2 If some factors stop their ability to grow, such as antibacterial, genetic modifications can arise that enable them to survive, thereby developing resistance to treatment. This is the natural process of bacteria to develop drug resistance.3 The ongoing battle against pathogenic infections involves the continuous exploration of new sources of antibacterial.1
Prolonged and excessive use of antibacterial agents has caused pathogenic bacteria to adapt, ultimately becoming resistant to treatment, despite the development of various antibacterial agents. When bacteria become resistant, infections caused by these microbes become more difficult to treat. This often necessitates the use of stronger or combination antibacterial, which may further accelerate the development of resistance.4 The decline in effectiveness of plant-derived metabolites and synthetic bioactive compounds has led to an increasing recognition of microorganisms as a potential source of novel antibacterial agents. These microbes produce diverse bioactive compounds with antibacterial, antifungal, and cytotoxic properties, enabling them to adapt to various environmental conditions. Recent research emphasizes their unique biological characteristics as an underexplored reservoir for developing new antibacterial treatments.5
Microorganisms, including bacteria, can serve as sources of antibacterial agents. Unique environmental conditions, such as those found in caves, enable certain bacteria to produce secondary metabolites.6 Many cave systems are located within karst regions, which are characterized by total darkness, high humidity, elevated mineral content (especially calcium carbonate), and low levels of organic matter. These nutrient-limited and pH-extreme environments (either alkaline or acidic) force microbes to develop specific adaptations, including the production of distinct secondary metabolites with potential antibacterial activity. The ability to synthesize bioactive compounds is often part of their survival strategy under such extreme conditions. In addition, carbonate-producing bacteria found in karst areas may function as natural conservation agents by forming protective layers of calcium carbonate (CaCO3). The CaCO3 deposits produced by these bacteria are primarily in the form of vaterite, which has the potential to act as a natural protective coating on cave paintings, thereby slowing degradation due to environmental exposure. However, the metabolic activity and biofilm formation of certain bacterial species can also accelerate the deterioration of these artworks.7,8 The dissolution of calcium carbonate in karst environments contributes to the formation of unique microhabitats that support bacterial development with specialized metabolic capabilities. Consequently, microorganisms adapted to cave environments represent a promising source of novel bioactive compounds, including antibacterial agents.8
One of the major karst regions in South Sulawesi is the Maros-Pangkep Karst Area. UNESCO has designated this region as part of the UNESCO Global Geopark network in Indonesia on May 24, 2023.7 Despite its ecological significance, karst ecosystems remain largely unexplored. These ecosystems harbor a potentially vast diversity of microorganisms. Cave-dwelling microbes are considered promising sources of bioactive compounds, as many of them produce antibacterial substances either as a defense mechanism or to outcompete other organisms in nutrient-limited environments. This study aims to identify a new source of antibacterial compounds, specifically from cave-dwelling bacteria in the Maros-Pangkep Karst Area.
Sampling
Soil samples were collected from caves in the Maros-Pangkep karst area, including Leang Jarie (LJ) (05°01’57.1″S 119°41’12.9″E), Leang Ulu Wae (LU) (04°59’04.0″S 119°40’23.1″E), and Leang Kassi (LK) (04°50’07.4″S 119°35’23.5″E). These three caves were selected because they represent spatially separated karst microhabitats with distinct environmental characteristics, including differences in humidity, light penetration, organic matter availability, and airflow, all of which can influence microbial diversity and biosynthetic potential within cave ecosystems. Sampling was performed using a sterile spatula to collect soil from each cave site. The soil samples were stored in sterile containers, kept in a cooler box at 4-8 °C, and transported to the laboratory on the same day for microbial analysis.
Bacterial Isolation
Bacteria were isolated by performing serial dilutions of the soil samples up to 10-3, followed by inoculation onto tryptic soy agar (TSA) medium (Merck, Cat. No. 105458). Individual colonies with distinct morphologies were picked and subcultured 2-3 times on TSA to obtain pure isolates.
Preparation of Test Bacteria Suspension. The 24-hour-old test bacteria from the slanted agar were suspended using sterile distilled water. Using a spectrophotometer, the suspension density was adjusted to achieve the desired dilution equivalent to the 0.5 McFarland standard (OD600 = 1×108 CFU/mL), compared to a sterile distilled water blank.
Selection of Antibacterial Compound-Producing Bacteria
The selection of potential antibacterial compound-producing bacteria was performed using an antagonistic assay as a preliminary qualitative screening method, in which bacterial isolates were cultured together with test bacteria (S. aureus ATCC 25923 and E. coli ATCC 25922) to evaluate their ability to inhibit bacterial growth. The bacterial isolates were streaked in a straight line on the surface of Mueller-Hinton agar (MHA) medium. Isolates capable of producing antibacterial compounds will form a clear zone (zone of inhibition) in the streaked bacterial colonies.
Bacterial characterization
Observation of colony morphology
Bacterial isolates were grown in Nutrient Agar (NA) medium (Oxoid, Thermo Fisher Scientific, Cat. No. CM0003). After incubation, the resulting colonies were examined using a stereo microscope (Olympus SZ61, Japan). The colonies were characterized based on standard morphological parameters, including shape (circular, irregular, filamentous), margin (entire, undulate, lobate), elevation (flat, convex, umbonate), and color, following established microbiological guidelines.
Cell morphology observation
Gram staining
A bacterial colony was smeared thinly on a glass slide. Then it is fixed on a Bunsen flame. The slide was then sequentially treated with crystal violet (primary stain) (Merck, Cat. No. 101245), Lugol’s iodine solution (mordant) (Merck, Cat. No. 100213), 95% alcohol (decolorizer) (Merck, Cat. No. 100983), and safranin (counterstain) (Merck, Cat. No. 109217). After each staining step, the slide was rinsed gently with distilled water. Finally, the stained slide was observed under a light microscope (Olympus CX23, Japan) at 100× magnification.
Endospore observation
A bacterial colony was prepared on a glass slide and stained with malachite green (Merck, Cat. No. 101228). The slide was then placed over a water bath to facilitate stain penetration. Following this, the slide was counterstained with safranin (Merck, Cat. No. 109217) and rinsed with distilled water. The stained preparations were subsequently examined under a light microscope (Olympus CX23, Japan) at 100 × magnification.
Biochemical tests
Motility Test, Indole Test, and H2S (Hydrogen Sulfide) Test. A bacterial suspension was inoculated into SIM (Sulfide Indole Motility) (Oxoid, Cat. No. CM0435) medium using the stab method and then incubated at 37 °C for 24 hours. A positive result was shown by growth diffusion from the stab line and medium turbidity, indicating bacterial motility due to flagella. After an additional 48 hours at the same temperature, Kovac’s reagent was added to test for indole production from tryptophan, indicated by a violet-red ring on the medium’s surface. The presence of black precipitates suggested a reaction with sulfur groups via the enzyme desulfurase.9,10
TSIA (Triple Sugar Iron Agar) test
A bacterial suspension was taken using a straight inoculating needle and inoculated into TSIA medium (Oxoid, Cat. No. CM0277) by continuously streaking the slant. The culture was incubated for 24 hours at 37 °C. The color change in the medium from red to yellow demonstrates that the isolated bacteria fermented carbohydrates such as glucose, sucrose, and lactose, resulting in acid production. In addition, gas production by the bacteria was observed as the lifting of the medium.11
Citrate test: A bacterial suspension was taken using a loop and inoculated onto Simmons Citrate Agar (SCA) medium (Oxoid, Cat. No. CM0155) by streaking the surface continuously. The color change from green to blue demonstrates that the bacteria utilized citrate as the sole carbon source.11
MR (Methyl Red) test
A bacterial suspension (0.2 mL) was taken using a pipette and transferred into a test tube containing MR-VP (Methyl Red-Voges Proskauer) medium (Oxoid, Cat. No. CM0435). The culture was incubated for 5 days at 37 °C. After incubation, methyl red indicator (Merck, Cat. No. 106066) was added. A positive result was indicated by color change to red, suggesting that the bacteria produced mixed acids from glucose fermentation.
VP (Voges Proskauer) test
A bacterial suspension (0.2 mL) was taken using a pipette and transferred into a test tube containing MR-VP medium. After incubation for 3 days at 37 °C, 0.2 mL of potassium hydroxide (KOH) (Merck, Cat. No. 105033) and 0.6 mL of a-naphthol (Merck, Cat. No. 800980) were added. The color changes from yellow to red, suggesting that the bacteria carried out 2,3-butanediol fermentation, resulting in the detection of acetoin in the bacterial culture.
Catalase test
A bacterial isolate was inoculated onto a glass slide. Subsequently, 2 to 3 drops of hydrogen peroxide (H2O2) (Merck, Cat. No. 107209) were added to the smear. The formation of gas bubbles on the slide suggests that the bacteria produced catalase.12
Molecular identification
Molecular identification based on partial 16S rRNA gene sequencing was performed using the Geneaid Presto™ Mini gDNA Bacteria Kit with a special procedure for Gram-positive bacteria. Genomic DNA was extracted, and the 16S rRNA gene was amplified using universal bacterial primers 63F (5′-CAGGCCTAACACATGCAAGTC-3′) and 1387R (5′-GGGGGGTGTGTACAAGGC-3′). The resulting PCR product (~1500 bp) was sequenced, and the obtained sequence was compared with reference sequences in the NCBI database using BLAST analysis. Identification was assigned at the genus level based on the highest sequence similarity.
Gas chromatography-mass spectrometry (GC-MS) analysis
The bacterial isolates were cultured in MHB medium at 150 rpm for 3 days. The cell-free supernatant was collected via centrifugation, then extracted with ethyl acetate in a 1:1 ratio. The extract underwent rotary evaporation at 70 °C for 40 minutes. For GC-MS analysis using Shimadzu Ultra QP 2010, 100 µL of the extract was injected with an injector temperature of 250 °C, at a pressure of 76.9 kPa, and a flow rate of 14 mL/min. The ion source and interface temperatures were set at 200 °C and 280 °C, respectively. The scanning range was from 40-700 m/z, with the column temperature initially at 70 °C, increasing to 200 °C and then to 280 °C. Chromatogram data were analyzed using NIST 17 and Wiley 9 libraries, identifying compounds with a Similarity Index (SI) ≥90% as acceptable.
In silico test with molecular docking
The results of the GC-MS analysis were further examined using molecular docking techniques. The compound structures were downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). A literature review was conducted focusing on commonly studied antibacterial target proteins, which were validated using UniProt (https://www.uniprot.org). Protein targets were selected based on three key parameters: essentiality, drug-targetability, and availability of experimentally resolved 3D structures, allowing reliable molecular docking analysis.2,13,14 The selected bacterial proteins from S. aureus and E. coli served as receptor targets, with their respective native ligands used as control ligands. The structure of target proteins was retrieved from the Protein Data Bank (https://www.rcsb.org/) and then visualized using PyMOL.2 The grid box was determined based on the native ligand coordinates extracted from the receptor file. The docking process involving the test ligands, the native ligand (control ligand), and the receptor was conducted using the PyRx software. Subsequently, the visualization was conducted using Biovia Discovery Studio Visualizer software to examine the interactions and types of bonding between the receptor protein and the ligands.15
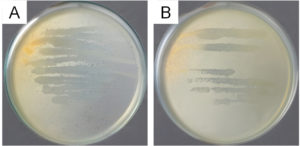
The results of the antagonistic test, 6 bacterial isolates exhibited preliminary antibacterial effects, as proven by clear zone formation. Furthermore, the 6 isolates were each distinguished based on macroscopic colony morphological characters, such as shape, edge, elevation, color, and surface texture (Table 1). Although precise measurements were not recorded at the time of testing, isolate LU3 exhibited a visibly wider inhibition zone compared to the other isolates, based on qualitative visual observation (Figure 1).
Table (1):
Morphology of colonies and cells from bacterial isolates obtained from sampling locations LJ, LU, and LK
| Isolate | Colony | Cell | |||||
|---|---|---|---|---|---|---|---|
| Shape | Elevation | Edge | Color | Shape | Gram | Endospore | |
| LJ8 | Circular | Raised | Entire | Beige | Long bacilli | Positive | + |
| LK4 | Circular | Convex | Entire | Beige | Short bacilli | Negative | – |
| LK5 | Circular | Raised | Entire | Beige | Long bacilli | Positive | + |
| LU1 | Circular | Convex | Entire | Beige | Short bacilli | Negative | – |
| LU2 | Circular | Convex | Entire | Beige | Short bacilli | Negative | – |
| LU3 | Irregular | Flat | Curled | Beige | Short bacilli | Positive | – |
*Numeric codes refer to the colony isolation sequence*+: Endospore detected; -: Endospore undetected
Figure 1. Antagonistic test result of LU3 isolates against test bacteria (A) S. aureus and (B) E. coli
Although the cross-streak antagonistic assay reflects primarily contact-dependent antibacterial effects, the consistent formation of clear zones produced by LU3 suggests that this isolate has the potential to be associated with antibacterial activity in preliminary screening assays. These potential isolates may produce various bioactive compounds, including antibacterial, pigments, toxins, or enzymes.16-18 Because the contact-based method does not allow confirmation of inhibition mediated by diffusible or secreted secondary metabolites, further chemical analysis was performed. Therefore, LU3 was subsequently subjected to ethyl acetate extraction followed by GC-MS analysis to tentatively characterize the secondary metabolite profiles. This approach allows us to explore whether LU3 produces putative bioactive compounds that may be associated with the antibacterial effects observed during preliminary screening.
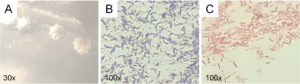
Additionally, macroscopic morphological characteristics of the bacterial colonies were differentiated based on colony shape, edge, elevation, and color. Microscopic morphological characteristics included Gram staining, cell shape, and the presence of bacterial endospores.19 The morphological observations of the LU3 bacterial isolate are shown in Figure 2.
Figure 2. Microscopic and macroscopic characterization of LU3 isolate: (A) colony morphology, (B) gram characteristics, and (C) Schaeffer–Fulton staining showing absence of endospores (no green structures observed)
Colony morphology analysis of the six active isolates showed clear differences in shape, elevation, and edge (Table 1). Observation using a stereo microscope revealed that colonies of the LU3 isolates exhibited an irregular shape, flat elevation, curled edges, and a beige color. Moreover, LU3 isolate was identified as a Gram-positive bacterium with a short bacillus morphology. Gram-positive bacteria form a stable complex between crystal violet and iodine. During the decolorization step with alcohol, Gram-positive cells dehydrate, the pores in the cell wall shrink, and the complex remains trapped within the cell, causing it to retain the purple color.20 Endospore staining of the LU3 isolate produced negative results, as evidenced by the absence of green-stained endospores, indicating the lack of endospore formation. While this trait does not impact its antibacterial metabolite production, it may necessitate careful storage and handling for long-term viability. Endospores are intracellular structures with a thick, retractile wall that enables bacterial survival under adverse environmental conditions.21
Endospore staining using the Schaeffer–Fulton method showed that isolate LU3 did not produce endospores. Under microscopic observation, vegetative cells stained red with safranin, while no green-stained endospores were detected, confirming the absence of endospore formation under the growth conditions tested. The lack of endospores suggests that LU3 is a non-sporulating bacterium, which is consistent with several genera commonly found in cave environments, such as Pseudomonas and Bacillus-like variants that lose sporulation ability under nutrient-rich laboratory conditions. This result supports the preliminary phenotypic characterization of LU3 and provides additional information for distinguishing it from other isolates obtained in this study.
Furthermore, the LU3 isolate which exhibited antimicrobial activity, was subjected to a series of biochemical tests to identify and classify the bacterium based on its metabolic activities. These tests assisted in determining the ability of the bacterium to degrade various compounds, produce specific enzymes, or utilize certain nutrients. The results of the biochemical tests are presented in Table 2 and Figure 3.
Table (2):
The Results of Biochemical Tests
| Isolate | SIM | TSIA | MR-VP | citrate | catalase | ||||
|---|---|---|---|---|---|---|---|---|---|
| Motility | Indole | H2S | sugar | Gas | MR | VP | |||
| LU3 | + | – | – | + | – | + | + | – | + |
Figure 3. Biochemical Test Results of LU3 Bacterial Isolate
(A) Motility, indole, and H₂S tests; (B) TSIA test; (C) MR test; (D) VP test; (E) Citrate test; and (F) Catalase test
Molecular identification
Based on 16S rRNA gene sequence analysis, isolate LU3 showed high similarity to members of the genus Exiguobacterium. BLAST analysis against the NCBI GenBank database revealed that the 16S rRNA sequence of isolate LU3 (approximately 1422-1511 bp in length, with 98% query coverage) shared a maximum sequence identity of 98.11% with several Exiguobacterium species, including Exiguobacterium sp., Exiguobacterium acetylicum, Exiguobacterium indicum, and related Exiguobacterium strains (Table 3). A slightly lower sequence similarity (98.03%) was observed with Ornithinibacillus scapharcae, indicating a more distant relationship compared to Exiguobacterium species. Based on these results, isolate LU3 was assigned to the genus Exiguobacterium. Species-level identification was not pursued due to the close similarity values among multiple Exiguobacterium species and the limitations of partial 16S rRNA gene resolution.
Table (3):
Closest GenBank matches of isolate LU3 based on 16S rRNA gene BLAST analysis
Total length (bp) |
Coverage % |
Close relative |
Identity % |
Gene bank accession no. |
|---|---|---|---|---|
1435 |
98% |
Exiguobacterium sp. |
98.11% |
MK414878.1 |
1462 |
98% |
Exiguobacterium acetylicum |
98.11% |
MW041269.1 |
1442 |
98% |
Exiguobacterium sp. F30 |
98.11% |
GU120647.1 |
1470 |
98% |
Exiguobacterium sp. IARI-R-140 |
98.11% |
JX429020.1 |
1453 |
98% |
Exiguobacterium sp. CP-2 |
98.11% |
JN642681.1 |
1511 |
98% |
Exiguobacterium indicum |
98.11% |
LC873072.1 |
1422 |
98% |
Ornithinibacillus scapharcae |
98.03% |
KU601305.1 |
GC-MS Analysis
Metabolite profiles of the ethyl acetate fraction from bacterial isolates that showed preliminary antibacterial activity were analyzed using GC-MS. The profiles of the tentatively identified compounds are presented in Table 4. Peaks with a library match (Similarity Index, SI) ≥90% were considered acceptable for tentative compound assignment based on library matching. Raw chromatogram data, mass spectra, and library match details have been mentioned in the supplementary Figures 1 and 2, and Table 1 for transparency and reproducibility.
Table (4):
Tentative GC-MS Profile of ethyl acetate extract compounds of isolate LU3
Peak |
Retention Time |
Tentatively Identified Compound |
Library Match (%) |
Area (%) |
|---|---|---|---|---|
2 |
5.772 |
Styrene |
95 |
0.08 |
32 |
18.014 |
2,5-Piperazinedione, 3-methyl-6-(1-methylethyl) |
96 |
1.29 |
33 |
18.752 |
1,4-diazabicyclo[4.3.0]nonan-2,5-dione, 3-methyl |
94 |
0.64 |
35 |
19.853 |
Cyclo(L-prolyl-L-valine) |
98 |
10.95 |
39 |
21.454 |
Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropy) |
92 |
6.12 |
40 |
21.828 |
Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropyl) |
94 |
4.50 |
41 |
21.944 |
5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a] |
91 |
6.14 |
42 |
22.268 |
Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropyl) |
91 |
5.99 |
50 |
32.251 |
Cyclohexane, 1,3,5-triphenyl- |
93 |
0.34 |
E. coli and S. aureus are significant human pathogens involved in a wide range of infections. Both species exhibit various patterns of antibacterial resistance, making them representative models for the development of new antibacterial agents. E. coli is a Gram-negative bacillus associated with diarrheal diseases, including dysentery, and is a primary cause of uncomplicated cystitis. It also contributes to other extraintestinal infections, such as pneumonia.22 In contrast, S. aureus is a Gram-positive bacterium that causes numerous skin infections and a significant number of severe invasive infections globally each year, including pneumonia.23 In this context, the present study demonstrated the preliminary antibacterial potential of metabolites associated with cave-derived bacterial isolates based on initial screening assays. However, these findings are derived from qualitative antagonistic tests and do not provide quantitative measures of antibacterial potency, such as minimum inhibitory concentration (MIC) or minimum bactericidal concentration (MBC) values. Accordingly, the absence of MIC/MBC determinations represents a limitation of this study.
The GC-MS data (Table 3) suggest that the compound with the highest concentration, accompanied by a peak with a library match (Similarity Index, SI) ≥90% in the ethyl acetate extract from isolate LU3 was detected at peak 35, with a retention area of 10.95% and a library match value of 98%. This compound was tentatively identified as Cyclo(L-prolyl-L-valine). This diketopiperazine-type compound has been reported to modulate pathogenicity without contributing to antibacterial resistance.24 The second most abundant compound was tentatively detected at peak 41, contributing 6.14% and a library match value of 91%. This compound was tentatively identified as 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a], which has been reported to exhibit antifungal properties, the potential to treat fungal infections, and other antimicrobial activities.25 Additionally, the third most abundant compound was tentatively detected at peak 39, 40, and 42, with consecutive concentrations of 6.12%, 4.50%, and 5.99% and the library’s consecutive match rate is 92%, 94% and 91%. This compound was tentatively identified as pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methylpropyl), which has been reported to possess antibacterial and antioxidant properties.26 As procedural blanks were not included, the GC-MS-based compound assignments should be considered tentative and may include potential background or media-derived components. BLAST analysis of the 16S rRNA gene sequence (1,313 bp, 100% query coverage) assigned isolate LU3 to the genus Exiguobacterium. The sequence showed high similarity (up to 98.11% identity) to multiple Exiguobacterium species, supporting genus-level identification rather than species-level resolution.
In silico
The conformation of the bioactive compound and the protein target can be visualized using molecular docking methods.2,27 Three bioactive compounds included compound 1 (Cyclo(L-prolyl-L-valine)), compound 2 (5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a]) and compound 3 (Pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methylpropyl)) was selected for in silico evaluation to explore their potential interactions with bacterial target proteins. The macromolecular targets used in this study were bacterial proteins involved in vital cellular functions, selected to provide mechanistic insight into possible antibacterial interactions rather than direct evidence of biological activity.
Enzymes from test bacteria act as proteins or receptors, one of which is the PBPs (Penicillin Binding Proteins) enzyme from S. aureus and E. coli. PBPs is essential for the bacterial cell wall and is a major target for antibacterial agents. The structural modification of the PBPs leads to the emergence of resistance. PBPs is a type of enzyme found in the cell membrane that helps build the cell wall by connecting certain parts of the sugar chains.28,29 The PBP enzyme from S. aureus (PDB ID 8VBV) is paired with its native ligand (63U: native inhibitor of the protein), while the PBP enzyme from E. coli (PDB ID 2EX9) is paired with its native ligand (35P: native inhibitor of the protein). This native ligand serves as a comparison or control ligand. Penicillin G (benzylpenicillin) is also used as a control since structural evidence shows that PBPs form enzyme-acyl complexes with benzylpenicillin (Penicillin G), indicating an interaction at the active site of the transpeptidase enzyme. Therefore, benzylpenicillin represents an appropriate positive control for validating docking protocols targeting PBPs.30
Molecular docking predicts interactions between compounds and their targets based on the compound’s structure, generating possible binding conformations.31 This approach provides a computational estimation of ligand receptor interactions rather than direct experimental evidence of biological activity. The method’s validity is confirmed with an RMSD value ≤ 2 Å, indicating lower deviation errors during docking. Smaller RMSD values correspond to reduced errors in the docking process. The RMSD value for structural conformational alignment is considered acceptable if it is less than 3, and optimal if it is less than 2. The closer the value is to 0, the better the alignment. Regarding binding affinity, it is important to describe how the ligand interacts with the receptor. In molecular docking, more negative binding affinity values indicate a predicted favorable interaction between the ligand and the receptor within the docking model, while a higher value suggests a weaker affinity.32,33 Based on the molecular docking results presented in Table 5, the PBP from S. aureus (PDB ID code 8VBV) showed a binding affinity of -7.8 kcal/mol for the native ligand. In comparison, all three test ligands exhibited comparable predicted binding affinities to the native ligand. Among the test ligands, Pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methylpropyl) had the binding energy closest to the native ligand, at -7.3 kcal/mol. Benzylpenicillin, used as a control, displayed a more negative binding affinity of -7.6 kcal/mol compared to the compound ligands. These findings indicate that ligands with more negative binding affinity values demonstrate stronger interactions with the receptor.33
Table (5):
Binding affinity score of target proteins and ligands
| Target Protein | Ligand | Binding Affinity (kcal/mol) | RMSD |
|---|---|---|---|
| PBP of S. aureus (PDB code ID 8VBV) | Ligand native: 63U | -7.8 | 0.0 |
| 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a] | -6.4 | 0.0 | |
| Cyclo(L-prolyl-L-valine) | -6.1 | 0.0 | |
| Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropyl) | -7.3 | 0.0 | |
| Benzylpenicillin | -7.6 | 0.0 | |
| PBP of E. coli (PDB code ID 2EX9) | Ligand native: 35P | -6.4 | 0.0 |
| 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a] | -5.1 | 0.0 | |
| Cyclo(L-prolyl-L-valine) | -5.3 | 0.0 | |
| Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropyl) | -6.2 | 0.0 | |
| Benzylpenicillin | -6.3 | 0.0 |
The molecular docking study of E. coli’s penicillin-binding protein (PDB ID 2EX9) revealed a binding score of -6.4 kcal/mol for the native ligand. Three test ligands showed predicted binding affinities within a comparable range. Pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methylpropyl) at -6.2 kcal/mol, Cyclo(L-prolyl-L-valine) at -5.3 kcal/mol, and 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a] at -5.1 kcal/mol.
A lower binding score signifies a more stable ligand receptor interaction, influenced by hydrogen bonds, van der Waals interactions, and hydrophobic interactions. Hydrogen bonds are typically stronger due to their directional nature, while hydrophobic interactions enhance stability by minimizing interactions between nonpolar residues and water.32 Visualizations of these interactions are shown in Figures 4 and 5.
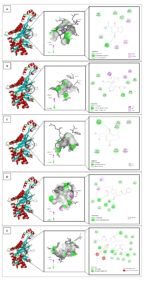
Figure 4. In silico interaction analysis of Cyclo(L-prolyl-L-valine) ligand (A); 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a] ligand (B); Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropyl) ligand (C); Benzylpenicillin ligand (D); and native ligand 63U (E) with PDB ID: 8VBV. A specific visualization type for each ligand as follows: 3D visualization (highlighted by black circle); Binding distance and receptor surface representation, hydrogen bonds indicated by dashed lines with green/pink surfaces as donors/acceptors; and 2D visualization showing interaction types, number of bonds, and involved amino acids (e.g., van der Waals, hydrogen bonds, unfavorable contacts)
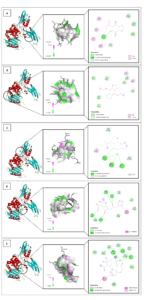
Figure 5. In silico interaction analysis of Cyclo(L-prolyl-L-valine) ligand (A); 5,10-Diethoxy-2,3,7,8-tetrahydro-1H,6H-dipyrrolo[1,2-a] ligand (B); Pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro-3-(2-methylpropyl) ligand (C); Benzylpenicillin ligand (D); and native ligand 35P (E) with PDB ID: 2EX9. A specific visualization type for each ligand as follows: 3D visualization (highlighted by black circle); Binding distance and receptor surface representation, hydrogen bonds indicated by dashed lines with green/pink surfaces as donors/acceptors; 2D visualization showing interaction types, number of bonds, and involved amino acids (e.g., van der Waals, hydrogen bonds, unfavorable contacts)
The binding energy and stability of molecular complexes are significantly influenced by amino acid residues and the interactions between ligands and receptors. An increase in hydrogen bonds leads to enhanced stability, while van der Waals and hydrophobic interactions also play a crucial role. A strong binding interaction is characterized by the formation of stable hydrogen bonds and engagement with key amino acid residues at the receptor’s active site.34
Based on the binding affinity values and the structural interactions, the three compounds from the LU3 bacterial isolates demonstrated predicted binding poses within the PBP active site that are comparable to those of reference ligands in the docking model. The molecular docking results revealed that the positive control, Penicillin G (benzylpenicillin), exhibited the most favorable conformation, closely matching the native ligand of each protein receptor. Among the test compounds, Pyrrolo[1,2-a]pyrazine-1,4-dione, hexahydro-3-(2-methylpropyl) displayed the closest interaction profile to the control, followed by Cyclo(L-prolyl-L-valine) and, finally, 5,10-Diethoxy- 2,3,7,8-tetrahydro-1H,6H-dipyrrolo [1,2-a]. These findings suggest that the identified compounds may interact with key amino acid residues within the PBP binding pocket according to the docking simulations. However, such in silico predictions do not constitute direct evidence of antibacterial activity, and further experimental validation is required to confirm their inhibitory potential against PBP in S. aureus and E. coli.
The formation of inhibition zones against pathogenic bacteria suggests preliminary antibacterial activity from the metabolic extracts of bacteria isolated from LU3. These findings are supported by molecular docking results, which indicated potential binding interactions of three major compounds identified in the extracts with the selected target proteins of the tested bacteria. The binding affinity results demonstrated that these compounds formed favorable binding conformations with the native ligands of the respective bacterial protein receptors. Based on 16S rRNA gene sequence analysis, the LU3 isolate was identified as belonging to the genus Exiguobacterium. Overall, these results indicate that bacterial metabolites from the Maros-Pangkep karst area, particularly from Leang Ulu Wae, have promising potential as sources of antibacterial candidates. These findings provide a preliminary foundation for developing new antibacterial treatments, particularly given the rising issue of antimicrobial resistance. Future research should focus on purifying the active compounds, elucidating their structures, and conducting quantitative antibacterial assays and in vivo validation to assess their clinical efficacy and safety profiles.
Additional file: Table S1 and Figure S1-S2.
ACKNOWLEDGMENTS
The authors would like to express their gratitude for the support provided by the Department of Biology, Faculty of Mathematics and Natural Sciences, Universitas Hasanuddin.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
This study was funded through research grant under the Fundamental Collaborative Research scheme (Contract No. 00323/UN4.22/PT.01.03/2023).
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript and/or in the supplementary files.
ETHICS STATEMENT
Not applicable.
- Nwobodo DC, Ugwu MC, Anie CO, et al. Antibiotic resistance: The challenges and some emerging strategies for tackling a global menace. Wiley. 2022;36(9):1-10.
Crossref - Husain DR, Wardhani R. Antibacterial activity of endosymbiotic bacterial compound from Pheretima sp. earthworms inhibit the growth of Salmonella typhi and Staphylococcus aureus: in vitro and in silico approach. Iran J Microbiol. 2021;13(4):537-543.
Crossref - Uddin TM, Chakraborty AJ, Khusro A, et al. Antibiotic resistance in microbes: History, mechanisms, therapeutic strategies and future prospects. J Infect Public Health. 2021;14(12):1750-1766.
Crossref - Siahaan S, Herman MJ, Fitri N. Antimicrobial Resistance Situation in Indonesia: A Challenge of Multisector and Global Coordination. J Trop Med. 2022;7:1-10.
Crossref - Danquah CA, Minkah PAB, Junior IOD, Amankwah KB, Somuah SO. Antimicrobial Compounds from Microorganisms. Antibiotics. 2022;11(3):285.
Crossref - Farda B, Djebaili R, Vaccarelli I, Gallo MD, Pellegrini M. Actinomycetes from Caves: An Overview of Their Diversity, Biotechnological Properties, and Insights for Their Use in Soil Environments. Microorganisms. 2022;10(2):1-19.
Crossref - Haedar N, Iqram M, Ambeng, Yusriana, Priosambodo D, Lebe R. Bacterial Communities on Degraded Prehistoric Rock Paintings in Maros-Pangkep Global Geopark. Philipp J Sci. 2024;153(1):391-402.
Crossref - Zhafirah NA, Haedar N, Sentosa S, Gani F. Micromorphology Characterization Of Crystal Calcium Carbonate And Exopolysaccharides Quantification Carbonatogenic Bacterial LTP4-d Isolated from Historical Painting of Maros-Pangkep Karst Area, Indonesia. J Biodivers. 2024;25(5):2139-2147.
Crossref - Sasaki-Imamura T, Yano A, Yoshida Y. Production of indole from L-tryptophan and effects of these compounds on biofilm formation by Fusobacterium nucleatum ATCC 25586. Appl Environ Microbiol. 2010;76(13):4260-4268.
Crossref - Yang Z, Helmann T, Baudin M, et al. Genome-wide identification of novel flagellar motility genes in Pseudomonas syringae pv. tomato DC3000. Front. Microbiol. 2025;16:1-16.
Crossref - Pardosi L, Manalu AI, Lisnahan CV. Isolation and Characterization of Superior Symbiotic Bacterial Isolates from the Oenggae Sea with Potential as Antibacterial. J Biol Sci. 2024;24(3):403-411.
Crossref - Chai B, Qiao Y, Wang H, et al. Identification of YfiH and the Catalase CatA As Polyphenol Oxidases of Aeromonas media and CatA as a Regulator of Pigmentation by Its Peroxyl Radical Scavenging Capacity. Front Microbiol. 2017;8:1-12.
Crossref - Wardhani R, Darsih C, Iwansyah AC, Indriati A, Hamid HA, Husain DR. In Silico Molecular Docking Approach and In Vitro Antioxidant and Antimicrobial Activity of Physalis angulata L. Extract. J Comput Biophys Chem. 2024;23(2):161-174.
Crossref - Wijaya T, Akmal AAM, Herman N, Hasan AA, Hafiz A, Widyastuti H. Apoptotic Effects Sulfated Polysaccharides of Caulerpa racemosa Extract on Colorectal Cancer Cells through Caspase-3. Mol Cell Biomed Sci. 2025;9(3):171-78.
Crossref - Ayodele PF, Bamigbade AT, Bamigbade OO, et al. Illustrated Procedure to Perform Molecular Docking Using PyRx and Biovia Discovery Studio Visualizer: A Case Study of 10kt With Atropine. Prog Drug Discov Biomed Sci. 2023;6(1):1-31.
Crossref - Nurbailis, Djamaan A, Rahma H, Liswarni Y. Secondary Metabolite Production by Trichoderma spp and its Potential as Antibacteria. Int J Curr Microbiol Appl Sci. 2019;8(4):196-201.
Crossref - Setiaji J, Feliatra F, Teruna HY, et al. Antibacterial activity in secondary metabolite extracts of heterotrophic bacteria against Vibrio alginolyticus, Aeromonas hydrophila, and Pseudomonas aeruginosa. F1000Res. 2020;9(1491):1-11.
Crossref - Gajic I, Kabic J, Kekic D, et al. Antimicrobial Susceptibility Testing: A Comprehensive Review of Currently Used Methods. Antibiotics. 2022;11(4):1-26.
Crossref - Fitri DA, Asih ENN, Kartika AGD, et al. Morphological characteristics of halophilic bacteria in traditional salt production. Indones J Mar Life Util. 2022;3(1):1-7.
Crossref - Vijayakumar T, Divya B, Vasanthi, Narayan M, Kumar AR, Krishnan R. Diagnostic Utility of Gram Stain for Oral Smears. J Microsc Ultrastruct. 2023;11(3):130-134.
Crossref - Yao PY, Annamaraju P. Clostridium perfringens infection. In: StatPearls [Internet]. StatPearls Publishing. 2023. https://www.ncbi.nlm.nih.gov/books/NBK559049/. Accessed July 28, 2025.
- Mueller M, Rausch-Phung EA, Tainter CR. Escherichia coli infection. In: StatPearls [Internet] StatPearls Publishing; 2025. https://www.ncbi.nlm.nih.gov/books/NBK564298/. Accessed December 20, 2025.
- Cheung GYC, Bae JS, Otto M. Pathogenicity and virulence of Staphylococcus aureus. Virulence. 2021;12(1):547-569.
Crossref - Kachhadia R, Kapadia C, Singh S, et al. Quorum Sensing Inhibitory and Quenching Activity of Bacillus cereus RC1 Extracts on Soft Rot-Causing Bacteria Lelliottia amnigena. ACS Omega. 2022;7(29):25291-25308.
Crossref - Hongjuan L, Lu L, Shuwen Z, Wenming C, Jiaping L. Identification of antifungal compounds produced by Lactobacillus casei AST18. Curr Microbiol. 2012;65(2):156-161.
Crossref - Kiran GS, Priyadharsini S, Sajayan A, Ravindran A, Selvin J. An antibiotic agent pyrrolo[1,2-a]pyrazine-1,4-dione,hexahydro isolated from a marine bacteria Bacillus tequilensis MSI45 effectively controls multi-drug resistant Staphylococcus aureus. RSC Adv. 2018; 8(32):17837-17846.
Crossref - Husain DR, Wardhani R, Erviani AE. Antibacterial activity of bacteria isolated from earthworm (Pheretima sp.) gut against Salmonella typhi and Staphylococcus aureus: in vitro experiments supported by computational docking. Biodiversity. 2022;23(2):1125-1131.
Crossref - Ambade SS, Gupta VK, Bhole RP, Khedekar PB, Chikhale RV. A Review on Five and Six-Membered Heterocyclic Compounds Targeting the Penicillin-Binding Protein 2 (PBP2A) of Methicillin-Resistant Staphylococcus aureus (MRSA). Molecules. 2023;28(20):1-53.
Crossref - Baran A, Kwiatkowska A, Potocki L. Antibiotics and Bacterial Resistance-A Short Story of an Endless Arms Race. Int J Mol Sci. 2023;24(6):1-34.
Crossref - Moon TM, D’Andrea ֹD, Lee CW, et al. The structures of penicillin-binding protein 4 (PBP4) and PBP5 from Enterococci provide structural insights into b-lactam resistance. J Biol Chem. 2018;293(48):18574-18584.
Crossref - Agu PC, Afiukwa CA, Ezeh EM, et al. Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Sci Rep. 2023;13(1):1-18.
Crossref - Millan-Casarrubias EJ, Garcia-Tejeda YV, Rosa CHGI, Ruiz-Mazon L, Hernandez-Rodriguez YM, Cigarroa-Mayorga OE. Molecular Docking and Pharmacological In Silico Evaluation of Camptothecin and Related Ligands as Promising HER2-Targeted Therapies for Breast Cancer. Curr Issues Mol Biol. 2025;47(3):1-20.
Crossref - Spassov DS. Binding Affinity Determination in Drug Design: Insights from Lock and Key, Induced Fit, Conformational Selection, and Inhibitor Trapping Models. Int J Mol Sci. 2024;25(13):1-24.
Crossref - Chen D, Oezguen N, Urvil P, Ferguson C, Dann SM, Savidge TC. Regulation of protein-ligand binding affinity by hydrogen bond pairing. J Comput Chem. 2016;2(3):1-16.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.