ISSN: 0973-7510
E-ISSN: 2581-690X
A bacterial strain isolated from chilli pepper was identified as Bacillus amyloliquefaciens using VITEK 2, MALDI-TOF mass spectrometry, and 16S rRNA sequencing. Designated SBCHGBA1, this strain was assessed for its ability to inhibit the growth of Aspergillus species and suppress aflatoxin synthesis. On trypticase soy agar, SBCHGBA1 exhibited strong antifungal activity against Aspergillus flavus MTCC2798 and Aspergillus parasiticus MTCC411. Among 15 Aspergillus flavus strains tested, 13 produced detectable aflatoxin levels (7.0-257.4 µg/l) in trypticase soy broth after 14 days at 25 °C. Remarkably, co-cultivation with SBCHGBA1 completely inhibited aflatoxin production in all except in one strain. Additionally, a 77% reduction in aflatoxin content was observed in chilli pepper inoculated with Aspergillus flavus MTCC2798 when co-cultured with SBCHGBA1. These findings highlight Bacillus amyloliquefaciens SBCHGBA1 as a promising biocontrol agent for mitigating aflatoxin contamination in chilli peppers and related food systems.
Bacillus amyloliquefaciens, Aspergillus flavus, Aspergillus parasiticus, Aflatoxin
Aflatoxins are potent mycotoxins, primarily produced by the fungi Aspergillus flavus and Aspergillus parasiticus. They are a major cause of liver cancer in human and are commonly found on agricultural crops like spices, peanuts, maize, cottonseed, and tree nuts, especially in warm and humid climates. Crops can be contaminated by Aspergillus flavus and Aspergillus parasiticus during various stages, including in the field, at harvest, and throughout storage and transportation. In comparison to Aspergillus parasiticus, Aspergillus flavus is more aggressive and widely distributed aflatoxin producing fungal species.1 Among mycotoxins, aflatoxins are particularly toxic and carcinogenic.2 Aflatoxin B1 and Aflatoxin B2 are mainly produced by Aspergillus flavus, whereas Aspergillus parasiticus can produce all 4 types of aflatoxins namely aflatoxin B1, aflatoxin B2, aflatoxin G1, and aflatoxin G2. Aflatoxin B1 is produced in higher amounts than the other aflatoxins. Nayak and co-workers have reported that out of thirty-four Aspergillus flavus strains isolated from indoor air samples of farmers’ houses in the coastal region of Indian state, Odisha, 44% of these isolates were produced stable Aflatoxin B1.3 In 2012, International Agency for Research on Cancer (IARC) considered aflatoxin to be the most natural toxic compound and declared it a group 1 human carcinogen.4,5 Biological control methods have previously shown promise in managing aflatoxin contamination in food crops. Various microorganisms, including bacteria, yeasts, and non-toxigenic strains of Aspergillus flavus and Aspergillus parasiticus, have been evaluated for their effectiveness in controlling aflatoxin levels. In laboratory experiments, many bacteria such as Lactobacilli species, Rastonia species, Bacillus subtilis, Pseudomonas species, and Burkholderia species, have shown the ability to inhibit fungal growth and production of aflatoxins by Aspergillus species.6 Nayak and his team of researchers reported that both the active culture broth and the cell-free filtrate of Bacillus species, isolated from the Indian cow-shed environment, were highly effective in suppressing the mycelial growth of Aspergillus flavus, achieving up to a 90% reduction in fungal development.7
Chilli pepper (Capsicum annuum L.) is a significant commercial spice, commonly affected by aflatoxin contamination, particularly in its processed forms. It causes health hazards to consumers and affects the global market for chilli pepper and its products. The European Commission has established maximum allowable limits for aflatoxins in spices, including whole and ground chilli pepper are 5 µg/kg for aflatoxin B1 and 10 µg/kg for the combined total of aflatoxins B1, B2, G1, and G2.8 Aflatoxin produced by Aspergillus flavus, is the major problem observed in dried chilli pepper in tropical countries.9
In vitro and field conditions, there are integrated approaches for the management of aflatoxins contamination in chilli pepper that involve the use of plant extracts, bioagents, and fungicides against Aspergillus flavus.10
Bacillus amyloliquefaciens is a Gram-positive, rod-shaped, primarily aerobic bacterium characterized by its ability to produce endospores and exhibit filamentous, chain-forming cellular arrangements. Taxonomically, it is classified within the genus Bacillus, family Bacillaceae, and belongs to the Bacillus clade of the Firmicutes phylum. Bacillus amyloliquefaciens synthesizes a broad spectrum of structurally diverse antimicrobial secondary metabolites, notably including both cyclic and linear lipopeptides. It exhibits a close phylogenetic affinity to Bacillus subtilis, as evidenced by comparative genomic and molecular analyses.11 As per the recent studies, researchers have included four species of Bacillus namely Bacillus amyloliquefaciens, Bacillus nakamurai, Bacillus velezensis, and Bacillus siamensis under the operational group Bacillus amyloliquefaciens due to their similarities in cell wall compositions, Guanine-Cytosine (G+C) content, Fatty Acid Methyl Esters (FAME), DNA-DNA hybridization (DDH) studies, 16S rRNA sequence, morphological and physiological characteristics.12 Bacillus amyloliquefaciens is used as a biocontrol agent to prevent the infection of agricultural plants by plant pathogens.13,14
In this study, strain of Bacillus amyloliquefaciens was used to evaluate its potential to control the aflatoxin producing ability and growth of Aspergillus flavus MTCC2798, Aspergillus parasiticus MTCC411 and 14 other Aspergillus isolates from different spice products.
The Bacterial Strains used in the study
The bacterial strain, SBCHGBA1 was initially recovered as a contaminant on Sabouraud Dextrose Agar (SDA) plates during yeast and mold analysis of chilli pepper sample. The strain was subsequently isolated on nutrient agar (NA) plates and subjected to preliminary identification using the Bruker MALDI-TOF Biotyper and bioMérieux VITEK 2 systems. Further confirmation of SBCHGBA1 was achieved through 16S rRNA gene sequencing, as the strain demonstrated notable anti-aflatoxigenic activity. In addition, Bacillus megaterium SBCHGBM1, obtained from ground chilli pepper samples, was identified using MALDI-TOF and VITEK 2 methods and employed as a bacterial control in the anti-aflatoxigenic study. Pure cultures of both bacterial strains were maintained on trypticase soybean agar (TSA) medium and preserved at 2-8 °C for subsequent experimental use.
Fungal strains utilized in the study
Reference strains, Aspergillus flavus MTCC2798 and Aspergillus parasiticus MTCC411 were obtained from the Microbial Type Culture Collection and Gene Bank (MTCC), CSIR–Institute of Microbial Technology, Chandigarh, India, and maintained on potato dextrose agar (PDA) plates at 2-8 °C. The study also included fourteen Aspergillus flavus isolates recovered from various spice samples. Isolation of Aspergillus flavus from spices was performed following the USFDA BAM Chapter 18 protocol. During analysis, fungal colonies on dichloran 18% glycerol (DG18) agar were examined for morphological similarity to Aspergillus flavus. Colonies exhibiting comparable characteristics were subcultured on PDA and identified using MALDI-TOF as described below. The fourteen fungal isolates identified as Aspergillus flavus by MALDI-TOF with identification scores above 2 were selected for the anti-aflatoxigenic study. All fungal cultures were maintained on PDA and stored at 2-8 °C throughout the study.
Bacterial identification by VITEK 2
The bacteria to be identified were subcultured on trypticase soy agar (TSA) medium and prepared serial dilution of bacteria in 0.85% NaCl solution. The optical density (OD) values of bacterial dilutions were adjusted to a range of 1.80-2.20 by using McFarland 2.0 standard, as measured by the DensiChek (bioMérieux) turbidity meter. VITEK 2 system (bioMérieux, Inc., Hazelwood, MO) Version: 08.01 and VITEK 2 Bacillus identification card (BCL card) were used for the identification of bacteria.15
Bacterial and fungal identification by MALDI-TOF biotyper
Bacterial and fungal isolates were purified by culturing on nutrient agar (NA) and potato dextrose agar (PDA) prior to identification using the Bruker MALDI-TOF Biotyper (Bruker Daltonics, Billerica, MA). For MALDI-TOF MS analysis, 1 µL of 70% formic acid was dispensed into each well of a steel anchor 96-well plate, and bacterial and fungal colonies were transferred onto the separate wells containing formic acid and allowed to air dry. Next, 1 µL of matrix solution, comprised of α-cyano-4-hydroxycinnamic acid dissolved in 47.5% water, 50% acetonitrile, and 2.5% trifluoroacetic acid, was layered over the dried samples and permitted to dry prior to measurement. The instrument was calibrated before analysis using the bacterial test standard (BTS) provided by Bruker Daltonics.16
Bacterial identification by 16S rRNA gene sequencing
High-quality genomic DNA was isolated from a pure culture of strain SBCHGBA1, and its integrity was evaluated by running samples through a 1% agarose gel, where a clear high-molecular-weight band was noted and chosen for subsequent studies. The 16S rRNA gene was amplified using universal primers (27F and 1492R), producing a PCR product that appeared as a single band around 1500 bp after gel electrophoresis, which was then purified to remove impurities. Sequencing of this purified amplicon involved both forward and reverse primers on an ABI 3730xl Genetic Analyzer, using the BDT v3.1 Cycle Sequencing Kit. The paired reads were merged to form a consensus sequence with Aligner software, then analyzed using BLAST against the NCBI GenBank database to find sequences with the highest similarity scores. The ten closest matches were selected, aligned in Clustal W and MEGA7.17-19
Antifungal activity assays of Bacillus amyloliquefaciens SBCHGBA1
The antagonistic activity of Bacillus amyloliquefaciens SBCHGBA1 against Aspergillus flavus MTCC2798 and Aspergillus parasiticus MTCC411 was evaluated using dual culture assays on Sabouraud Dextrose Agar (SDA) and Tryptic Soy Agar (TSA). An inoculum of 0.1 ml, containing approximately 300 colony-forming units (CFU), was prepared by spread-plating Bacillus amyloliquefaciens SBCHGBA1 onto both SDA and TSA plates, with bacterial CFUs quantified via serial dilution and plating on TSA. After bacterial inoculation, 0.5 mm fragments of fungal mycelia were placed in the centre of each plate and parallel control plates containing only fungal isolates were also maintained. All plates were incubated at 25 °C for five days, after which fungal growth was monitored, and the diameter of colonies was measured in millimetres using a calibrated ruler.20
Assessment of anti-aflatoxigenic activity of Bacillus amyloliquefaciens SBCHGBA1 in tryptic soy broth
The experiment utilized 250 ml screw-cap borosilicate glass bottles, each containing 50 ml of tryptic soy broth (TSB), which were sterilized at 121 °C and 15 psi for 15 minutes in an autoclave to maintain aseptic conditions. Once cooled, individual bottles were inoculated with 1000 CFU per 50 ml of each of the fifteen Aspergillus flavus strains, including MTCC2798 and fourteen isolates from different spice sources and Bacillus amyloliquefaciens SBCHGBA1 at 1000 CFU per 50 ml of TSB. Controls included bottles with only bacterial or only fungal inoculation, as well as uninoculated media controls. The initial count of Bacillus amyloliquefaciens SBCHGBA1 in the inoculum was determined via standard plate counting on nutrient agar. To prepare fungal spores, Aspergillus flavus cultures were grown on PDA at 25 °C for seven days, followed by spore collection using a rinse with 10 ml sterile 0.05% Tween 80 and filtration through four layers of sterile cheesecloth to remove mycelial fragments. The spore concentration was measured through serial dilution in 0.1% peptone water and by plating on PDA. The pH of TSB in each bottle was set to 5.5 using sterile 1 N HCl. All cultures were incubated at 25 °C for fourteen days to promote microbial interactions, then subjected to final sterilization by autoclaving at 121 °C for 15 minutes. After cooling, a 25 ml sample was taken from each bottle for aflatoxin measurement.21
Assessment of anti-aflatoxigenic activity of Bacillus amyloliquefaciens SBCHGBA1 in chilli pepper
Fifty grams of sterilized ground chilli pepper were placed into 500 ml screw-cap glass bottles, labelled A through G. The treatments were arranged as follows, A: Blank control (no microbial inoculation); B: Inoculated with Aspergillus flavus MTCC2798; C: Inoculated with Bacillus amyloliquefaciens SBCHGBA1; D: Inoculated with Bacillus megaterium SBCHGBM1; E: Co-inoculated with Aspergillus flavus, Bacillus amyloliquefaciens, and Bacillus megaterium; F: Co-inoculated with Aspergillus flavus and Bacillus amyloliquefaciens; G: Co-inoculated with Aspergillus flavus and Bacillus megaterium. Each microorganism was added at an inoculum concentration of 50,000 CFU per gram of chilli pepper. All samples were incubated at 25 °C for 14 days. Following incubation, 25 g of chilli pepper were collected from each bottle for aflatoxin analysis.
Analysis of aflatoxins
For aflatoxin measurement, each 25 ml aliquot of TSB or 25 g chilli pepper was combined with 100 ml of an 80:20 methanol–water mixture and homogenized using a blender at 2000 RPM for 2 minutes. The mixture was filtered through Whatman No. 1 filter paper, and 20 ml of the filtrate was subsequently diluted to a total volume of 100 ml using 1 mM phosphate-buffered saline (PBS, pH 7.4) before passage through a 1.5 µm microfiber filter. Ten millilitres of this filtered solution were applied to an AflaPrep immunoaffinity column at a controlled flow rate of 1 ml/min. After washing the column twice with 10 ml of water, aflatoxins were eluted with 1 ml methanol. The eluate was analyzed via high-performance liquid chromatography (HPLC) equipped with a fluorescence detector, using a reversed-phase C18 column maintained at 40 °C. Separation was performed using an isocratic mobile phase of water and methanol (55:45 v/v) at a flow rate of 1 ml/min. Aflatoxin detection relied on fluorescence settings with excitation and emission wavelengths of 365 nm and 464 nm, respectively. To improve sensitivity for aflatoxin B1, aflatoxin B2, aflatoxin G1, and aflatoxin G2, a photochemical reactor with a 254 nm UV lamp and PTFE coil was placed between the column and detector. The procedure was validated over a concentration range of 0.5-300 µg/l or µg/kg using certified reference standards for all four aflatoxin compounds, and chromatographic outputs were processed using Shimadzu LC Solution software version 2.1 for quantification.22
Data analysis
All experiments were performed in three independent replicates, and data are expressed as mean values with their corresponding standard deviation (SD). Statistical calculations were carried out using Microsoft Excel.
Identification of bacterial strain, SBCHGBA1
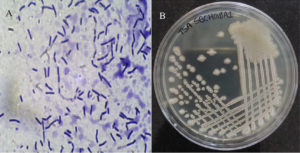
The bacterium isolated from the SDA plate during yeast and mold analysis of ground chilli pepper was further cultured on trypticase soyabean agar (TSA), producing creamy white, irregularly margined, and large circular colonies after 24 hours at 35 °C. Microscopically, it appeared as Gram-positive rods measuring 0.6-0.8 µm in width and 1.5-2.7 µm in length, occurring singly, in pairs, or chains (Figure 1). The isolate was conclusively identified as Bacillus amyloliquefaciens through a combination of advanced methods, including the Bruker MALDI-TOF Biotyper, VITEK 2 system, and 16S rRNA gene sequencing. The results demonstrated high confidence in identification, with a MALDI-TOF score of 2.08 and a VITEK 2 probability of 97%. The 16S rRNA gene sequence (1467 bp) was aligned with closely related sequences and the results are summarised in Table 1, revealing 99.86% similarity to Bacillus velezensis and Bacillus amyloliquefaciens, both part of the Bacillus amyloliquefaciens operational group known for close genetic relatedness.13 Due to limitations in species-level discrimination by 16S sequencing alone, the combined use of three techniques provided conclusive identification of the isolate as Bacillus amyloliquefaciens. The 16S ribosomal RNA gene, partial sequence is available under GenBank accession PV649863.
Table (1):
Top 10 Alignments for SBCHGBA1 16S rRNA Gene Sequence (Based on NCBI BLAST)
Description |
Max Score |
Total Score |
Query Cover |
E-value |
Percent |
Accession |
|---|---|---|---|---|---|---|
Bacillus amyloliquefaciens subsp. plantarum str. FZB42, complete genome |
2697 |
25075 |
100% |
0 |
99.86% |
CP000560.2 |
Bacillus velezensis strain Htq6 chromosome, complete genome |
2697 |
4261 |
100% |
0 |
99.86% |
CP050462.1 |
Bacillus velezensis strain LPL061 chromosome, complete genome |
2697 |
22411 |
100% |
0 |
99.86% |
CP042271.1 |
Bacillus velezensis strain CACC 316 chromosome, complete genome |
2697 |
24114 |
100% |
0 |
99.86% |
CP048002.1 |
Bacillus velezensis strain SRCM101368 chromosome, complete genome |
2697 |
24169 |
100% |
0 |
99.86% |
CP031694.1 |
Bacillus velezensis strain BA-26 chromosome, complete genome |
2697 |
23977 |
100% |
0 |
99.86% |
CP046918.1 |
Bacillus velezensis strain BvL03 chromosome, complete genome |
2697 |
24197 |
100% |
0 |
99.86% |
CP041192.1 |
Bacillus velezensis strain SYP-B637 chromosome, complete genome |
2697 |
24125 |
100% |
0 |
99.86% |
CP043546.1 |
Bacillus amyloliquefaciens strain X030 chromosome, complete genome |
2697 |
24203 |
100% |
0 |
99.86% |
CP040672.1 |
Bacillus velezensis strain B268, complete genome |
2697 |
24203 |
100% |
0 |
99.86% |
CP053764.1 |
Figure 1. (A) Gram-staining image of the bacterial strain SBCHGBA1; (B) Colonies of SBCHGBA1 on Trypticase Soybean Agar (TSA) medium
Antifungal studies
Bacillus amyloliquefaciens SBCHGBA1 exhibited potent antifungal activity when tested on SDA and TSA media against Aspergillus flavus MTCC2798 and Aspergillus parasiticus MTCC411. In control experiments, without the bacterium, Aspergillus flavus formed colonies measuring 64 mm on SDA and 62 mm on TSA, while co-cultivation with Bacillus amyloliquefaciens reduced these sizes to 27 mm and 3 mm, respectively. Aspergillus parasiticus exhibited a colony size of 52 mm on TSA in the absence of Bacillus amyloliquefaciens, but only 2 mm when both were present. These results, detailed in Table 2 and Figure 2, indicate that Bacillus amyloliquefaciens SBCHGBA1 inhibited Aspergillus flavus growth by approximately 58% on SDA and 95% on TSA, with a 37% higher inhibition rate on TSA. Suppression of Aspergillus parasiticus growth on TSA was even more pronounced, reaching 96%. The high dextrose (glucose) content in SDA serves as an abundant energy and carbon source, potentially promoting vigorous mold growth, as both media are maintained at an acidic pH to favour mold development.
Table (2):
Inhibition of Fungal Growth by Bacillus amyloliquefaciens SBCHGBA1
Plating Medium |
Fungal Strain(s) |
Colony Diameter After 5 Days (mm)* |
|---|---|---|
SDA |
Aspergillus flavus MTCC 2798 |
64 ± 1a |
SDA |
Aspergillus flavus MTCC2798 + SBCHGBA1 |
27 ± 2b |
TSA |
Aspergillus flavus MTCC 2798 |
62 ± 2a |
TSA |
Aspergillus flavus MTCC 2798 + SBCHGBA1 |
3 ± 1c |
TSA |
Aspergillus parasiticus MTCC411 |
52 ± 2d |
TSA |
Aspergillus parasiticus MTCC411 + SBCHGBA1 |
3 ± 1c |
Note: *Values are expressed as mean ± standard deviation (n = 3). Different superscript letters indicate statistically significant differences at P ≤ 0.05. This table presents the colony measurements for Aspergillus flavus and Aspergillus parasiticus, with and without Bacillus amyloliquefaciens SBCHGBA1, following a five days incubation at 25 °C using SDA and TSA media. The addition of SBCHGBA1 resulted in a marked reduction in fungal colony size, highlighting its antifungal activity.
Figure 2. Photographic documentation of antifungal assays conducted with Bacillus amyloliquefaciens SBCHGBA1.
Note: (A) Growth of Aspergillus flavus on TSA without Bacillus amyloliquefaciens; (B) Growth of Aspergillus flavus on TSA with Bacillus amyloliquefaciens; (C) Growth of Aspergillus parasiticus on TSA without Bacillus amyloliquefaciens; (D) Growth of Aspergillus parasiticus on TSA with Bacillus amyloliquefaciens; (E) Growth of Aspergillus flavus on SDA without Bacillus amyloliquefaciens; (F) Growth of Aspergillus flavus on SDA with Bacillus amyloliquefaciens
Anti-aflatoxigenic studies
Bacillus amyloliquefaciens SBCHGBA1 was evaluated for its ability to inhibit aflatoxin production by 15 Aspergillus flavus strains which including 14 spice-derived isolates and one standard strain in TSB medium. The results are summarized in Table 3. After 14 days at 25 °C, 13 strains produced aflatoxins ranging from 7.0-263.4 µg/l, while two strains showed no detectable levels. Notably, the presence of Bacillus amyloliquefaciens SBCHGBA1 suppressed aflatoxin synthesis across all tested strains, with 14 strains producing undetectable amounts (LOQ >0.5 µg/l), and only one strain producing a minimal 1 µg/l. Only aflatoxin B1 and aflatoxin B2 were detected, with no aflatoxin G1 or aflatoxin G2 observed. These data underscore the robust anti-aflatoxigenic effects of Bacillus amyloliquefaciens SBCHGBA1, which prevented total aflatoxin concentrations exceeding 1 µg/l.
Table (3):
Impact of Bacillus amyloliquefaciens on Aflatoxin Production in TSB (mean ± SD, n = 3)
| Fungal strain | Source | Aflatoxin Without Bacillus amyloliquefaciens (µg/l) | Aflatoxin With Bacillus amyloliquefaciens (µg/l) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| B1 | B2 | G1 | G2 | Total aflatoxin | B1 | B2 | G1 | G2 | Total aflatoxin | ||
| A. flavus MTCC 2798 | MTCC | 190.7 ± 2.4 | 5.2 ± 0.3 | ND* | ND | 195.9 ± 2.6 | ND | ND | ND | ND | ND |
| A. flavus SBCHGAF1 | Red chillies | 13.5 ± 0.6 | 0.9 ± 0.2 | ND | ND | 14.4 ± 0.8 | ND | ND | ND | ND | ND |
| A. flavus SBCHGAF2 | Chillies ground | 44.9 ± 1.5 | 1.0 ± 0.2 | ND | ND | 45.9 ± 1.7 | ND | ND | ND | ND | ND |
| A. flavus SBCHCAF3 | Chillies crushed | 257.4 ± 4.9 | 6.0 ± 0.4 | ND | ND | 263.4 ± 5.7 | ND | ND | ND | ND | ND |
| A. flavus SBCPAF4 | Curry powder | 35.7 ± 0.9 | 1.0 ± 0.3 | ND | ND | 36.7 ± 1.1 | 1.0 | ND | ND | ND | 1.0 |
| A. flavus SBCHCAF5 | Chilies ground | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND |
| A. flavus SBCPAF6 | Garam masala | 28.0 ± 1.8 | 2.6 ± 0.5 | ND | ND | 30.6 ± 2.3 | ND | ND | ND | ND | ND |
| A. flavus SBCHGAF7 | Chilies ground | 215.5 ± 2.4 | 13.3 ± 0.8 | ND | ND | 228.8 ± 3.2 | ND | ND | ND | ND | ND |
| A. flavus SBCHAF8 | Chillies | 160.6 ± 2.6 | 11.3 ± 0.8 | ND | ND | 171.9 ± 3.4 | ND | ND | ND | ND | ND |
| A. flavus SBCHGAF9 | Chillies ground | 16.1 ± 0.7 | ND | ND | ND | 16.1 ± 0.7 | ND | ND | ND | ND | ND |
| A. flavus SBCPAF10 | Curry powder | ND | ND | ND | ND | ND | ND | ND | ND | ND | ND |
| A. flavus SBCHAF11 | Chillies whole | 160.7 ± 5.3 | 4.4 ± 0.7 | ND | ND | 165.1 ± 6.0 | ND | ND | ND | ND | ND |
| A. flavus SBCHAF12 | Chillies whole | 7.0 ± 0.6 | ND | ND | ND | 7.0 ± 0.6 | ND | ND | ND | ND | ND |
| A. flavus SBCHAF13 | Chillies stem | 11.1 ± 0.9 | ND | ND | ND | 11.1 ± 0.9 | ND | ND | ND | ND | ND |
| A. flavus SBCHAF14 | Chillies ground | 147.3 ± 1.8 | 5.4 ± 0.6 | ND | ND | 152.7 ± 1.5 | ND | ND | ND | ND | ND |
Note: *ND: Not detected (LOQ < 0.5 µg/l). All values represent mean ± standard deviation for three replicates. This table displays the aflatoxin quantities (types B1, B2, G1, G2, and total) produced by various Aspergillus flavus strains from different sources, cultured in TSB both with and without Bacillus amyloliquefaciens SBCHGBA1. Across most strains, aflatoxin production was not detected in the presence of Bacillus amyloliquefaciens, demonstrating its strong inhibitory effect on aflatoxin synthesis
The results of the anti-aflatoxigenic study on Bacillus amyloliquefaciens SBCHGBA1 in chilli pepper are shown in Table 4. The tested Aspergillus flavus MTCC2798 strain produced approximately 17 µg/kg of total aflatoxins in chilli pepper after 14 days of incubation. According to EU standards, chilli pepper containing more than 10 µg/kg of aflatoxin is unacceptable. When Aspergillus flavus was co-cultivated with Bacillus megaterium, the total aflatoxins level remained around 16 µg/kg, showing no significant difference. However, co-cultivation with Bacillus amyloliquefaciens limited aflatoxin production to less than 3.9 µg/kg, which falls within the acceptable range under EU norms. This indicates an approximate 77% reduction in aflatoxin production by Aspergillus flavus in the presence of Bacillus amyloliquefaciens.
Table (4):
Aflatoxin content in chilli pepper after 14 days of microbial treatment
Chilli pepper samples |
B1 (µg/kg) |
B2 (µg/kg) |
G1 (µg/kg) |
G2 (µg/kg) |
Total aflatoxin (µg/kg) |
|---|---|---|---|---|---|
A |
ND* |
ND |
ND |
ND |
ND |
B |
16.2 ± 1.7 |
0.9 ± 0.1 |
ND |
ND |
17.1 ± 1.8 |
C |
ND |
ND |
ND |
ND |
ND |
D |
ND |
ND |
ND |
ND |
ND |
E |
4.3 ± 0.2 |
ND |
ND |
ND |
4.3 ± 0.2 |
F |
3.9 ± 0.1 |
ND |
ND |
ND |
3.9 ± 0.1 |
G |
15.5 ± 1.6 |
0.8 ± 0.1 |
ND |
ND |
16.3 ± 1.6 |
Note: A: Blank (no treatment), B: Chilli pepper treated with Aspergillus flavus, C: Chilli pepper treated with Bacillus amyloliquefaciens, D: Chilli pepper treated with Bacillus megaterium, E: Chilli pepper treated with Aspergillus flavus, Bacillus amyloliquefaciens, and Bacillus megaterium, F: Chilli pepper treated with Aspergillus flavus and Bacillus amyloliquefaciens, G: Chilli pepper treated with Aspergillus flavus and Bacillus megaterium. *ND: Not detected (LOQ < 0.5 µg/kg). All values represent mean ± standard deviation for six replicates
Previous studies involving various antifungal and antibacterial strains of Bacillus amyloliquefaciens have identified three families of Bacillus lipopeptides: surfactins, iturins, and fengycins. These lipopeptides were investigated for their antagonistic activity against a broad spectrum of potential phytopathogens, including both bacteria and fungi. Their antimicrobial properties primarily stem from their unique amphiphilic structure, which enables them to disrupt the integrity of the plasma membrane or cell wall of target microorganisms in a manner like most antimicrobial detergents.23-26 In one of the earlier reported research, fengycins isolated from Bacillus amyloliquefaciens JFL21 exhibited antimicrobial activity against Listeria monocytogenes, Aeromonas hydrophila, and Colletotrichum gloeosporioides at wide range of pH and temperature. Remarkably, these compounds retained their activity even after exposure to ultraviolet sterilization, chemical reagents, and proteolytic enzymes.27 Bacillus amyloliquefaciens SBCHGBA1, sourced from chilli pepper, effectively suppressed both fungal growth and aflatoxin production of Aspergillus species, aligning with previous reports of Bacillus amyloliquefaciens strains functioning as probiotic and/or biocontrol agent in food systems. These results suggest that Bacillus amyloliquefaciens SBCHGBA1 may operate via similar antifungal pathways.
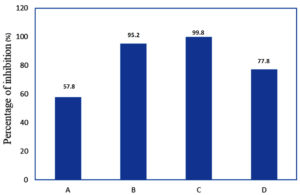
Depending on the variety and stage of ripeness, the pH of chilli peppers typically ranges between 4.9 and 6.2, placing them within a slightly acidic spectrum. The strain of Bacillus amyloliquefaciens isolated from chilli pepper demonstrated both anti-aflatoxigenic and antifungal properties when tested in acidic environments such as Sabouraud Dextrose Agar (SDA) and within chilli pepper itself. Prior research demonstrates the utility of these bacteria in animal feed for promoting gut health and productivity, as well as mediating aflatoxin detoxification through enzymatic degradation and gene expression modulation.28-33 The antifungal and anti-aflatoxigenic properties of Bacillus amyloliquefaciens SBCHGBA1, assessed in this study, are presented graphically in Figure 3. The findings support the application of Bacillus amyloliquefaciens SBCHGBA1 as a targeted biocontrol agent against aflatoxin-producing Aspergillus species in chilli pepper-based products and their derivatives.
Figure 3. Anti-aflatoxigenic properties of Bacillus amyloliquefaciens SBCHGBA1 against Aspergillus flavus.
Note: A: Growth inhibition on SDA, B: Growth inhibition on TSA, C: Aflatoxin inhibition on TSB (The percentage of inhibition was determined by calculating the average reduction in aflatoxin production across 13 strains of Aspergillus flavus.), D: Aflatoxin inhibition on chilli pepper
Chilli pepper powder is known to carry substantial microbial populations, often reaching levels of 106 to 108 CFU per gram of aerobic bacteria. A significant portion of this microbial load consists of heat-tolerant bacterial spores, particularly those belonging to Bacillus species.34-37 Methods are available for incorporating the beneficial strain Bacillus amyloliquefaciens SBCHGBA1 into chilli peppers as a biocontrol agent to prevent aflatoxin production. Prior to drying, chilli peppers typically undergo treatments designed to reduce their natural surface microflora, followed by steps to remove any treatment residues. After this preparation, the chilli peppers can be exposed to a harmless diluent containing the desired concentration of Bacillus amyloliquefaciens SBCHGBA1. Once treated, the peppers are dried by either naturally or through controlled methods and subsequently ground into powder that retains the introduced Bacillus amyloliquefaciens SBCHGBA1 culture. These methods must be standardized through scientifically validated protocols to enable the practical application of Bacillus amyloliquefaciens in chilli pepper.
Industrial strains belonging to subspecies of Bacillus amyloliquefaciens hold GRAS status from the USFDA for broad applications in food, feed, agriculture, enzyme production, probiotics, and biocontrol. In 1999, the USFDA issued a final rule affirming GRAS status for carbohydrase and protease enzyme preparations derived from Bacillus amyloliquefaciens, supported by scientific evidence and a history of safe use as direct food additives under relevant 21 CFR regulations.38,39
The bacterium isolated from chilli pepper was conclusively identified as Bacillus amyloliquefaciens using three complementary techniques. Antifungal assays revealed that Bacillus amyloliquefaciens SBCHGBA1 inhibited Aspergillus flavus MTCC2798 growth by approximately 58% on SDA and 95% on TSA. Furthermore, anti-aflatoxigenic experiments demonstrated complete suppression of aflatoxin production above 1 µg/l in all tested Aspergillus flavus strains after 14 days in TSB at 25 °C, along with a 77% reduction in aflatoxin levels in chilli pepper when co-cultivated with Bacillus amyloliquefaciens. These findings underscore the strong biocontrol potential of Bacillus amyloliquefaciens SBCHGBA1, naturally sourced from chilli pepper, for enhancing food safety in chilli-based products and related food systems.
ACKNOWLEDGMENTS
The authors thank Dr. AB Ramashree, Director Research (Spices Board, Govt. of India) and Ms. CM Srilatha, Senior Scientist (Spices Board, Govt. of India) for providing laboratory facilities and support for conducting these studies.
CONFLICT OF INTEREST
The authors declare that there is no conflict of interest.
AUTHORS’ CONTRIBUTION
All authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
FUNDING
None.
DATA AVAILABILITY
All datasets generated or analyzed during this study are included in the manuscript.
ETHICS STATEMENT
Not applicable.
- Payne GA, Brown MP. Genetics and physiology of aflatoxin biosynthesis. Annu Rev Phytopathol. 1998;36(1):329-362.
Crossref - Bedard LL, Massey TE. Aflatoxin B1-induced DNA damage and its repair. Cancer Lett. 2006;241(2):174-183.
Crossref - Nayak S, Dhua U, Samanta S. Morphological and toxigenic study of Aspergillus flavus from indoor air. J Environ Biol. 2019;40(4):648-654.
Crossref - Marchese S, Polo A, Ariano A, Velotto S, Costantini S, Severino L. Aflatoxin B1 and M1: Biological Properties and Their Involvement in Cancer Development. Toxins (Basel). 2018;10(6):214.
Crossref - Kumar P, Mahato DK, Kamle M, Mohanta TK, Kang SG. Aflatoxins: A Global Concern for Food Safety, Human Health and Their Management. Front Microbiol. 2017;17:07.
Crossref - Yin Y ni, Yan L yan, Jiang J hua, Ma Z hua. Biological control of aflatoxin contamination of crops. J Zhejiang Univ Sci B. 2008;9(10):787-792.
Crossref - Nayak S, Dhua U, Samanta S. Antagonistic activity of cowshed Bacillus sp. bacteria against aflatoxigenic and sclerotic Aspergillus flavus. J Biol Control. 202030;34(1):52-58.
Crossref - Costa J, Rodriguez R, Garcia-Cela E, et al. Overview of Fungi and Mycotoxin Contamination in Capsicum Pepper and in Its Derivatives. Toxins (Basel). 2019;11(1):27.
Crossref - Ezekiel CN, Ortega-Beltran A, Oyedeji EO, et al. Aflatoxin in Chili Peppers in Nigeria: Extent of Contamination and Control Using Atoxigenic Aspergillus flavus Genotypes as Biocontrol Agents. Toxins (Basel). 2019;11(7):429.
Crossref - Hajare SN, Gautam S, Sharma A. A novel strain of Bacillus amyloliquefaciens displaying broad spectrum antifungal activity and its underlying mechanism. Ann Microbiol. 2016;66(1):407-416.
Crossref - Chen XH, Koumoutsi A, Scholz R, et al. Comparative analysis of the complete genome sequence of the plant growth-promoting bacterium Bacillus amyloliquefaciens FZB42. Nat Biotechnol. 2007;25(9):1007-1014.
Crossref - Ngalimat MS, Yahaya RSR, Baharudin MMA Adil, et al. A Review on the Biotechnological Applications of the Operational Group Bacillus amyloliquefaciens. Microorganisms. 2021;9(3):614.
Crossref - Arrebola E, Jacobs R, Korsten L. Iturin A is the principal inhibitor in the biocontrol activity of Bacillus amyloliquefaciens PPCB004 against postharvest fungal pathogens. J Appl Microbiol. 2010;108(2):386-95.
Crossref - Zhao P, Quan C, Wang Y, Wang J, Fan S. Bacillus amyloliquefaciens Q 426 as a potential biocontrol agent against Fusarium oxysporum f. sp. spinaciae. J Basic Microbiol. 2014;54(5):448-456.
Crossref - Halket G, Dinsdale AE, Logan NA. Evaluation of the VITEK2 BCL card for identification of Bacillus species and other aerobic endosporeformers. Lett Appl Microbiol. 2010;50(1):120-126.
Crossref - Schulthess B, Ledermann R, Mouttet F, et al. Use of the Bruker MALDI Biotyper for Identification of Molds in the Clinical Mycology Laboratory. J Clin Microbiol. 2014;52(8):2797-2803.
Crossref - Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution (N Y). 1985;39(4):783-791.
Crossref - Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16(2):111-120.
Crossref - Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016;33(7):1870-1874.
Crossref - Gibson AM, Baranyi J, Pitt JI, Eyles MJ, Roberts TA. Predicting fungal growth: the effect of water activity on Aspergillus flavus and related species. Int J Food Microbiol. 1994;23(3-4):419-431.
Crossref - Wilson DM, Bell DK. Aflatoxin Production by Aspergillus flavus and A. parasiticus on Visibly Sound Rehydrated Peanut, Corn and Soybean Seed1. Peanut Science. 1984;11(1):43-45.
Crossref - Akiyama H, Goda Y, Tanaka T, Toyoda M. Determination of aflatoxins B1, B2, G1 and G2 in spices using a multifunctional column clean-up. J Chromatogr A. 2001;932(1-2):153-157.
Crossref - Lee HA, Kim JH. Isolation of Bacillus amyloliquefaciens Strains with Antifungal Activities from Meju. Prev Nutr Food Sci. 2012;17(1):64-70.
Crossref - Feng RY, Chen YH, Lin C, Tsai CH, Yang YL, Chen YL. Surfactin secreted by Bacillus amyloliquefaciens Ba01 is required to combat Streptomyces scabies causing potato common scab. Front Plant Sci. 2022;13:998707.
Crossref - Lin LZ, Zheng QW, Wei T, et al. Isolation and Characterization of Fengycins Produced by Bacillus amyloliquefaciens JFL21 and Its Broad-Spectrum Antimicrobial Potential Against Multidrug-Resistant Foodborne Pathogens. Front Microbiol. 2020;11:579621.
Crossref - Wang J, Qiu J, Yang X, et al. Identification of Lipopeptide Iturin A Produced by Bacillus amyloliquefaciens NCPSJ7 and Its Antifungal Activities against Fusarium oxysporum f. sp. niveum. Foods. 2022;11(19):2996.
Crossref - Yu GY, Sinclair JB, Hartman GL, Bertagnolli BL. Production of iturin A by Bacillus amyloliquefaciens suppressing Rhizoctonia solani. Soil Biol Biochem. 2002;34(7):955-963.
Crossref - Shi C, Zhang Y, Yin Y, et al. Amino acid and phosphorus digestibility of fermented corn-soybean meal mixed feed with Bacillus subtilis and Enterococcus faecium fed to pigs1. J Anim Sci. 2017;95(9):3996-4004.
Crossref - Zhang Y, Shi C, Wang C, et al. Effect of soybean meal fermented with Bacillus subtilis BS12 on growth performance and small intestinal immune status of piglets. Food Agric Immunol. 2018;29(1):133-146.
Crossref - WoldemariamYohannes K, Wan Z, Yu Q, et al. Prebiotic, Probiotic, Antimicrobial, and Functional Food Applications of Bacillus amyloliquefaciens. J Agric Food Chem. 2020;68(50):14709-14727.
Crossref - Wu Y, Wang Y, Zou H, et al. Probiotic Bacillus amyloliquefaciens SC06 Induces Autophagy to Protect against Pathogens in Macrophages. Front Microbiol. 2017;8.
Crossref - Du R, Jiao S, Dai Y, et al. Probiotic Bacillus amyloliquefaciens C-1 Improves Growth Performance, Stimulates GH/IGF-1, and Regulates the Gut Microbiota of Growth-Retarded Beef Calves. Front Microbiol. 2018;9:02006.
Crossref - Du H, Yao W, Kulyar MFEA, et al. Effects of Bacillus amyloliquefaciens TL106 Isolated from Tibetan Pigs on Probiotic Potential and Intestinal Microbes in Weaned Piglets. Microbiol Spectr. 2022;10(1):e01205-21.
Crossref - Melo Gonzalez MG, Romero SM, Arjona M, Larumbe AG, Vaamonde G. Microbiological quality of Argentinian paprika. Rev Argent Microbiol. 2017;49(4):339-346.
Crossref - Bata-Vidacs I, Baka E, Toth A, et al. Investigation of regional differences of the dominant microflora of spice paprika by molecular methods. Food Control. 2018;83:109-117.
Crossref - Gyorgy E, Laslo E, Antal M, Andras CD. Antibiotic resistance pattern of the allochthonous bacteria isolated from commercially available spices. Food Sci Nutr. 2021;9(8):4550-4560.
Crossref - Banerjee M, Sarkar PK. Microbiological quality of some retail spices in India. Food Res Int. 2003;36(5):469-474.
Crossref - Raman J, Noh JS, Kim JS, et al. Role of Bacillus Species in Food Industry: Advantages and Limitations. J Microbiol Biotechnol. 2025;35:e2507043.
Crossref - Lu H, Yang P, Zhong M, et al. Isolation of a potential probiotic strain Bacillus amyloliquefaciens LPB-18 and identification of antimicrobial compounds responsible for inhibition of food borne pathogens. Food Sci Nutr. 2023;11(5):2186-2196.
Crossref
© The Author(s) 2026. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.