ISSN: 0973-7510
E-ISSN: 2581-690X
This research sought to determine optimal conditions to maximize prodigiosin production by an indigenous Egyptian marine bacterial strain Serratia rubidaea RAM_Alex. Serratia rubidaea RAM_Alex isolated from bivalve samples of Temsah Lake, Ismailia, Egypt was used to investigate the production of the natural red pigment prodigiosin. Pigment production was assayed in different growth conditions using Nutrient broth as production medium. The water insoluble red pigment was extracted using ethanol and further purified by organic solvents. The pigment extract showed absorbance with a UV-Vis spectrophotometer at 535 nm and further characterized using TLC, FTIR and 1H-NMR. A statistical screening procedure was adopted to select the main factors affecting production. Analyses of Plackett- Burman design results demonstrated that peptone, NaCl, and culture volume were the most important independent variables. The near optimum medium contained (g/L): peptone 7, beef extract 5, yeast extract 1, NaCl 10, pH 6, using 25 ml culture volume, 100 ¼l inoculum size and incubation statically for 48 h at 30oC. When this condition was employed, a two fold increase in pigment yield was achieved reaching ~1600.511 mg/l.
Biopigments, nutrient broth, Plackett-Burman design, prodigiosin, Serratia rubidaea.
Prodigiosin is a characteristic member of a group of compounds with a common pyrrolyl pyrromethene (PPM) skeleton that belongs to a family of pyrrole red pigments1. Prodiginines are secondary metabolites produced by different bacterial species including Serratia marcescens, Pseudomonas magneslorubra, Vibrio psychroerythrus, Vibrio gazogenes, Alteromonas rubra, Streptomyces lividans and Streptomyces coelicolor2.
In light of its potential commercial values, there is a demand to optimize culture conditions to maximize prodigiosin production3. The use of statistical models to optimize culture medium components and conditions has increased in present-day biotechnology, due to its propensity and relevance. Production of prodigiosin is greatly influenced by physical factors such as temperature, pH, incubation time, inorganic phosphate, inoculum, substrate concentration and media components that include carbon and nitrogen sources, so it is important to find out an inexpensive and optimized media for the production of prodigiosin4,3,5. Identification of prodigiosin is carried out by means of Thin Layer Chromatography (TLC)6,7,1. Also instrumentation and analytical methods like Nuclear Magnetic Resonance (NMR), liquid-chromatography mass spectrometry (LC-MS), and Fourier-Transform Infrared Spectroscopy (FT-IR Spectroscopy) were applied to characterize and identify the purified compound4.
The present investigation focuses on the production and characterization of prodigiosin from a novel strain S. rubidaea RAM_Alex. Optimization of cultural parameters to achieve the enhanced production of the pigment was carried using Plackett-Burman design.
Bacterial isolation and identification
Nutrient broth (g/l): peptone 5, beef extract 3, yeast extract 2, NaCl 55 was used for the isolation of bacteria from clam samples collected from Temsah Lake, Egypt. For molecular identification, genomic DNA was isolated and 16S rDNA was amplified by polymerase chain reaction (PCR) using a forward primer (5’ AGAGTTTGATCMTGGCTCAG 3’) and a reverse primer (5’ TACGGYTACCTTGTTACGACTT 3’)8. 16S DNA was sequenced and sequence analysis was used to construct the phylogenetic tree. Biochemical characterization was carried out using VITEK 2 Compact, a fully automated microbial identification system (bioMérieux VITEK ®).
Extraction, purification and identification of the pigment
Cells of a 48 h old culture were separated by centrifugation at 10,000 x g for 10 min at 4°C. Pigment was extracted from cell pellets by ethanol9 and purified10. For pigment identification, absorption pattern of the purified pigment at different pHs was examined using a UV-Visible spectrophotometer (Unico, Shanghai) to determine maximum absorbance. The chemical structure of the purified product was characterized4 by Thin-Layer Chromatography (TLC) (TLC cards, Sigma, Germany) using solvent system consisting of hexane: ethyl acetate (3:1; v/v) and Fourier Transform Infrared Spectroscopy (FT-IR) (Bruker, Germany). The structure of the pigment was identified by Proton Nuclear Magnetic Resonance Spectroscopy (1H NMR) (JEOL, Tokyo) using D-chloroform (CDCl3) as solvent. Pigment concentration was measured using Beer-Lambert Law11.
A = µ l c
Where: A is an absorbance, µ is the molar absorptivity of the solution, l is the length of solution the light passes through (cm), c is the concentration of solution in mol/l.
Growth condition and pigment production
A loop of cells grown on NA plates for two days was used to inoculate a 250 ml flask containing 50 ml of Nutrient Broth. For prodigiosin production, the bacteria were grown in NB at 30°C and 120 rpm for 18 h. A standard inoculum (1%) of culture was added to 50 ml of production media in a 250 ml flask and incubated either static or in a shaker at 120 rpm and 30°C. Growth was measured at 600 nm in a UV-Visible spectrophotometer.
Optimization of factors affecting prodigiosin production by Plackett-Burman design
The variables chosen for the present study and their levels are given in Table 1. All variables were denoted as numerical factors and investigated at two widely spaced intervals designated as -1 (low level) and +1 (high level)12.
Table (1):
Factors examined as independent variables affecting prodigiosin production by Serratia rubidaea RAM_Alex and their levels in the Placket-Burman experimental design.
| Variable | Symbol | Level | ||
|---|---|---|---|---|
| -1 | 0 | +1 | ||
| Peptone (g/l) | P | 3 | 5 | 7 |
| Beef extract (g/l) | B | 1 | 3 | 5 |
| Yeast extract (g/l) | Y | 1 | 2 | 3 |
| NaCl (g/l) | Na | 3 | 5 | 10 |
| Inoculum size(µl) | IS | 100 | 500 | 1000 |
| Culture volume(ml) | CV | 25 | 50 | 75 |
| pH | pH | 6 | 7 | 8 |
Characterization and identification of isolated bacterium
The bacterium formed round, smooth, opaque, convex and red pigmented colonies on nutrient agar. Cells were Gram-negative and non-sporulating short rods. Data of VITEK revealed 99% similarity to Serratia rubidaea. Analysis of 16S rRNA showed 99% similarity to Serratia rubidaea strain SP25 and the isolate was thus designated as Serratia rubidaea RAM_Alex. The sequence was submitted to Genbank with Accession number KM411440 at NCBI. Figure 1 illustrates the phylogenetic tree showing the most related species to the strain.

Fig. 1: 16S rDNA-based dendogram showing the phylogenetic position of Serratia rubidaea RAM_Alex among representatives of related bacterial species
Extraction, purification and characterization of pigment
TLC analysis showed a single band with an Rf value of 0.62 (Fig. 2). The FTIR spectrum (Fig. 3) showed bands at 2926cm-1 attributed to the C-H group. Peaks at 3445cm-1 are due to aliphatic alcohols, primary amines and amide. Peaks at 1733 indicate the C=O, whereas, the peak at 1461cm-1 (C-H) refers to the bending vibration ethylene diamine, and 1380 refers to C-O in prodigiosin. The visible peak at 1279cm-1 corresponds to C-N. From the spectrum, the main functional group that resulted in red pigments is methylene. Pure pigment was analyzed for 1H-NMR (500 mHz) resulting in peaks correspond to chemical shifts at 7.25 ppm (C2), 6.95ppm (C12), 4.20 ppm (C11), 3.078 ppm (C18), 1.28 ppm (C21) and C22 at 0.8732 ppm. Data in figure 4 is in agreement to the structure of prodigiosin.
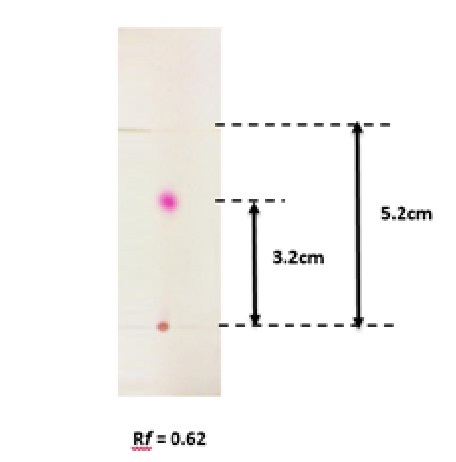

Fig. 2: Thin layer chromatography of Serratia rubidaea RAM_Alex prodigiosin. Solvent system consisted of hexane: ethyl acetate (3:1, v/v)
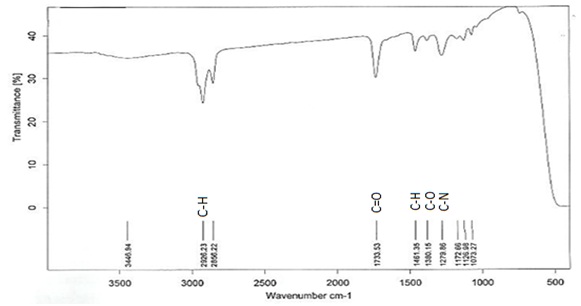
Fig. 3: FT-IR spectrum of Serratia rubidaea RAM_Alex red pigment
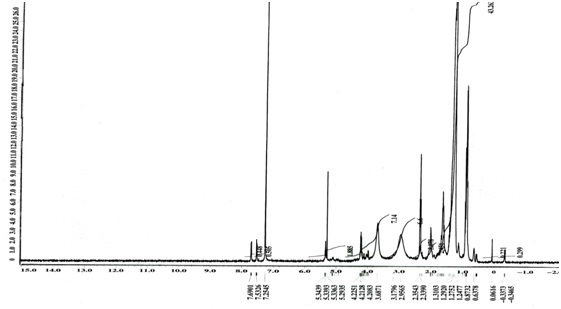
Fig. 4. 1H- NMR spectroscopy of Serratia rubidaea RAM_Alex red pigment
Growth condition and prodigiosin production by Serratia rubidaea RAM_Alex
Prodigiosin is a known secondary metabolite that does not have a role in growth, development and reproduction, typically formed during the end or near the stationary phase of growth. In this study, the production of prodigiosin increases linearly between 12 and 46 h under static condition, with maximum production (1707.39 mg/l) after 48h (Fig. 5). It is worth to mention that no pigmentation was observed when shacked cultures were used.
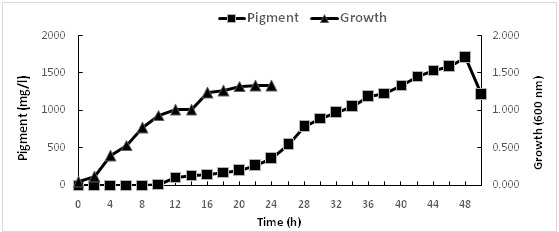 Fig. 5: Growth and prodigiosin production by Serratia rubidaea RAM_Alex grown in nutrient broth , pH 7under static condition for 2 days at 30oC. Pigment (%), Growth (²%)
Fig. 5: Growth and prodigiosin production by Serratia rubidaea RAM_Alex grown in nutrient broth , pH 7under static condition for 2 days at 30oC. Pigment (%), Growth (²%)Elucidation of factors affecting prodigiosin production using Plackett-Burman design
The design was applied with 7 different factors and all experiments were performed in duplicates and the average of results was presented as prodigiosin yield that was measured at 535nm after 48h (Table 2). The main effect was calculated as the difference between the average of measurements made at the high level setting (+1) and the average of measurements made at the low level setting (-1) for each factor. From the main effect analysis (Fig. 6) it was found that peptone, beef extract and NaCl in their high concentration positively affected prodigiosin production, while yeast extract, culture volume and pH negatively affected the process.
 Fig. 6: Positive and negative influence of different variables on prodigiosin production by Serratia rubidaea RAM_Alex based on the result of Plackett_Burman design
Fig. 6: Positive and negative influence of different variables on prodigiosin production by Serratia rubidaea RAM_Alex based on the result of Plackett_Burman designTable (2):
The Placket-Burman experimental design(in coded levels) with prodigiosin production as response.
| Independent variable | ||||||||
|---|---|---|---|---|---|---|---|---|
| Run | P | B | Y | NaCl | IS | CV | pH | mg/l |
| 1 | -1 | -1 | -1 | 1 | 1 | 1 | -1 | 586.48 |
| 2 | 1 | -1 | -1 | -1 | -1 | 1 | 1 | 367.85 |
| 3 | -1 | 1 | -1 | -1 | 1 | -1 | 1 | 669.77 |
| 4 | 1 | 1 | -1 | 1 | -1 | -1 | -1 | 1634.51 |
| 5 | -1 | -1 | 1 | 1 | -1 | -1 | 1 | 621.18 |
| 6 | 1 | -1 | 1 | -1 | 1 | -1 | -1 | 780.82 |
| 7 | -1 | 1 | 1 | -1 | -1 | 1 | -1 | 253.33 |
| 8 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 756.53 |
| 9 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 697.53 |
In order to evaluate the accuracy of the applied Plackett-Burman statistical design, a verification experiment was applied to compare between the predicted optimum levels of independent variables and the basal condition settings (Fig. 7). It was found that the yield of prodigiosin increased by 2 fold (~1600.511 mg/l) after validation under optimized conditions as compared with a basal medium (~765.02 mg/l).
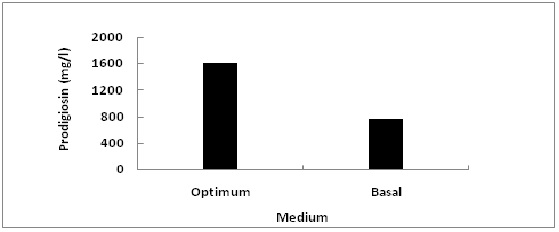 Fig. 7: Comparison between the predicted optimum levels of independent variables and the basal condition settings
Fig. 7: Comparison between the predicted optimum levels of independent variables and the basal condition settings During the last few decades, increasing attention has been paid to natural dye applications. This is due to increasing popularity of more natural lifestyle based on naturally sustainable goods13. Prodigiosin is one of the most popular natural dyes that possesses antibacterial, antifungal, antiprotozoal, cytotoxic, antitumor, antimalarial, antidiabetic, nonsteroidal and anti-inflammatory properties. There is a demand to develop high throughput and cost effective bioprocesses for pigment production3. Nutrient broth, a cost effective medium was used in this study for pigment production by a novel strain Serratia rubidaea RAM_Alex in batch fermentation. The observed high pigment production under statistic condition can be attributed to the effect of shear stress (from the shaking condition), which has been reported to be higher when the cells are either in aggregates or clusters compared to single cells14. In agreement with other researchers15,4, the present study shows that prodigiosin from Serratia rubidaea RAM_Alex is a secondary metabolite generally produced after the log phase of cell growth was highest in the stationary. Additionally, the characteristics of the production curve are similar to the production pattern of secondary metabolites16.
The red pigment was analyzed by TLC using mixture of n-hexane: ethyl acetate at (3:1; v/v). A single red band with Rf value of 0.62 was obtained is similar to that reported for Serratia marcescens UTM114. The FTIR spectrum was similar to those reported for prodigiosin17,6,18. The position of each proton in1H-NMR analysis clearly confirmed that the pigment isolated from Serratia rubidaea RAM_Alex is prodigiosin.
Several factors such as inorganic phosphate availability, medium composition, temperature and pH appear to affect the production of prodigiosin19. For improvement of pigment production by Serratia rubidaea RAM_Alex, Plackett–Burman design was applied. The positive effect of peptone and beef extract on pigment production is attributed to the contents of amino acids, vitamins and coenzymes, growth factors of natural components used 20,5. After optimization, a yield of ~1600.511 mg/l was achieved. It was reported that Serratia marcescens produced 184.32 mg/l and 277.74 mg/l of prodigiosin in glycerol and mannitol containing medium, respectively16, while mannitol was found to be suitable for growth and prodigiosin production instead of glycerol21. Other studies have reported different results associated with prodigiosin production. Wei and Chen observed 56-790 mg/l of prodigiosin cultured in oil supplemented Luria-Bertani broth medium22. The production of prodigiosin was reported in mutant Serratia marcescens 02 at a concentration of 96.5-583 mg/l23. Additionally, Gutiérrez-Román et al. reported a production of prodigiosin of 60 mg/l by Serratia marcescens CFFSUR-B2 cultivated in peanut medium24. Maximum optimal composition of the cultivated medium for prodigiosin production by Serratia marcescens by adding sucrose and glycine as the carbohydrate and energy source to cultivate medium resulted in prodigiosin yield increasing 2.12-fold (~579.02 mg/l) and 2.15-fold (~587.64 mg/l), respectively25. It was reported that the addition of maltose and glucose to nutrient broth gave a two-fold (~0.79 mg/ml and ~0.29 mg/ml, respectively) increase in yield over nutrient broth (~0.354 mg/ml) and peptone glycerol broth alone (~0.569 mg/ml)14.
From our results we can conclude the composition of the pre-optimized medium and condition for high prodigiosin production as follows (g/l): peptone 7, beef extract 5, yeast extract 1, NaCl 10, inoculum size 100 µl, pH 6, culture volume 25 ml at 30°C under static condition after 48 h that yielded ~1600.511 mg/l.
Although several recent studies disclosed a number of interesting biological properties of prodigiosin, this paper reports a higher value of prodigiosin production by a newly isolated indigenous marine bacteria Serratia rubidaea rather than Serratia marcescens from a relatively cheap medium. The work adds new information for microbial pigment production and optimization. Biopigments produced by bacteria have important biological activities, so in this study, the statistically based experimental designs proved to be an effective tool in optimizing the medium for prodigiosin production by Serratia rubidaea RAM_Alex and indicated that nutritional status of the growth medium plays an important role in the biosynthesis of prodigiosin. The present study demonstrated that peptone, NaCl, and culture volume significantly increased the production of prodigiosin from Serratia rubidaea RAM_Alex using a statistical screening procedure via a Plackett- Burman design.
ACKNOWLEDGMENTS
The authors are grateful to Botany & Microbiology Department, Faculty of Science, Alexandria University, Egypt and Institute of Oceanography and Fisheries, Alexandria, Egypt for providing facilities and expertise that greatly assisted this research.
- Lapenda Lins, J.C., Maciel, C.C.S, Xavier, H.S., Alves da Silva, C.A., Campos-Takaki, G.M. Production and toxicological evaluation of prodigiosin from Serratia marcescens UCP/WFCC1549 on mannitol solid medium. Int. J. Appl. Res. Nat. Prod., 2014; 7: 32-38.
- Venil, C.K., Lakshmanaperumalsamy, P. An insightful overview on microbial pigment, prodigiosin. Electronic Journal of Biology., 2009; 5: 49-61.
- Bhattacharya, S., Gulani, C., Das, A. Assessment of process parameters influencing the enhanced production of prodigiosin from Serratia marcescens and evaluation of its antimicrobial, antioxidant and dyeing potentials. Mal. J. Microbiol., 2012; 8: 116-122.
- Wang, S.L., Wang, C.Y., Yen, Y.H., Liang, T.W., Chen, S.Y., Chen C.H. Enhanced production of insecticidal prodigiosin from Serratia marcescens TKU011 in media containing squid pen. Process Biochem., 2012; 47: 1684-1690.
- Pradeep, B.V., Pradeep, F.S., Angayarkanni, J., Palaniswamy, M. Optimization and production of prodigiosin from Serratia marcescens MBB05 using various natural substrates. Asian J. Pharm. Clin. Res., 2013; 6: 34-41.
- Song, M.J., Bae, J., Lee, D.S., Kim, C.H., Kim, J.S., Kim, S.W., Hong, S.I. Purification and characterization of prodigiosin produced by integrated bioreactor from Serratia sp. KH-95. J. Biosci. Bioeng., 2006; 101: 157-161.
- Samrot, A.V., Senthilkumar, P., Chandana, K., Narendrakumar, G. Optimization of prodigiosin production by Serratia marcescens SU-10 and evaluation of its bioactivity. Int. Res. J. Biotechnol., 2011; 2: 128-133.
- Choudhury, J.D., Pramanik, A., Webster, N.S., Llewellyn, L.E., Gachhui, R., Mukherjeea, J. Draft genome sequence of Pseudoalteromonas sp. strain NW 4327 (MTCC 11073, DSM 25418), a pathogen of the Great Barrier Reef sponge Rhopaloeides odorabile. Genome Announc., 2014; 2: 1-14.
- Darshan, N., Manonmani, H.K. Prodigiosin and its potential applications. J. Food Sci. Technol., 2015; 52: 5393-5407.
- Shahitha S., Poornima, K. Enhanced production of prodigiosin production in Serratia marcescens. J. Appl. Pharm. Sci., 2012; 2: 138-140.
- Swinehart, D.F. The Beer-Lambert Law. J Chem Educ., 1962; 39: 333.
- Plackett, R.L., Burman, J.P. The design of optimum multifactor experiments. Biometrika., 1946; 33: 305-325.
- Shahid, M., Ul-Islam, S., Mohammad, F. Recent advancements in natural dye applications: a review. J. Cleaner Prod., 2013; 53: 310-331.
- Aruldass, C.A., Venil, C.K., Zakaria, Z., Ahmad, W.A. Brown sugar as a low-cost medium for the production of prodigiosin by locally isolated Serratia marcescens UTM1. Int. Biodeterior. Biodegrad., 2014; 95: 19-24.
- Harris K.P., Williamson, R., Slater, H., Cox, A., Abbasi, S., Foulds, I., Simonsen, T., Leeper, J., Salmond, P.C. The Serratia gene cluster encoding biosynthesis of the red antibiotic, Prodigiosin, shows species and strain dependent genome context variation. Microbiol., 2004; 150: 3547–3560.
- Kurbanoglu E.B., Ozdal, M., Ozdal, O.G., Algur, O.F. Enhanced production of prodigiosin by Serratia marcescens MO1 using ram horn peptone. Braz. J. Microbiol., 2015; 46: 631-637.
- Montaner, B., Castillo-Avila, W., Martinell, M., Ollinger, R., Aymami, J., Giralt, E., Pérez-Tomás, R. DNA interaction and dual topomerase I and II inhibition properties of the anti-tumor drug prodigiosin. Toxicol. Sci., 2005; 85: 870-879.
- Sumathi, C., MohanaPriya, D., Swarnalatha, S., Dinesh, M.G., Sekaran, G. Production of prodigiosin using tannery fleshing and evaluating its pharmacological effects. Sci. World J., 2014; doi: 10.1155/2014/290327.
- Williamson, N.R., Simonsen, H.T., Ahmed, R.A., Goldet, G., Slater, H., Woodley, L., Leeper, F.J., Salmond. G.P. Biosynthesis of the red antibiotic, prodigiosin, in Serratia: identification of a novel 2-methyl-3-n-amylpyrrole (MAP) assembly pathway, definition of the terminal condensing enzyme, and implications for undecylprodigiosin biosynthesis in Streptomyces. Mol. Microbiol., 2005; 56: 971-989.
- Bharmal, M.H., Jahagirdar, N., Aruna, K. Study on optimization of prodigiosin production by Serratia marcescens MSk1 isolated from air. Int. J. Adv. Biol. Res., 2012; 2: 671-680.
- Araújo, H. W., Fukushima, K., Takaki, G.M. Prodigiosin production by Serratia marcescens UCP 1549 using renewable-resources as a low cost substrate. Molecules, 2010; 15: 6931-6940.
- Wei, Y., Chen W.C. Enhanced production of prodigiosin-like pigment from Serratia marcescens SM”R by medium improvement and oil-supplementation strategies. J. Biosci. Bioeng., 2005; 99: 616-622.
- Tao, J.L., Wang, X.D., Shen, Y.L., Wei, D.Z. Strategy for the improvement of prodigiosin production by a Serratia marcescens mutant through fed-batch fermentation. World J. Microbiol. Biotechnol., 2005; 21: 969-972.
- Gutiérrez-Román, M.I., Holguín-Meléndez, F., Bello-Mendoza, R., Guillén-Navarro, K., Dunn, M.F., Huerta-Palacios, G. Production of prodigiosin and chitinases by tropical Serratia marcescens strains with potential to control plant pathogens. World J. Microbiol. Biotechnol., 2012; 28: 145-153.
- Venil, C.K., Zakaria, Z.A., Ahmad, W.A. Bacterial pigments and their applications. Process Biochem., 2013; 48: 1065-1079.
© The Author(s) 2017. Open Access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License which permits unrestricted use, sharing, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.


